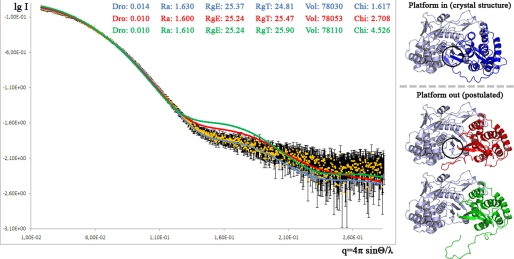
FIGURE 7.
Conformation of J4_NS5b in solution compared with the crystal structure and more open models. SAXS spectra measured in solution (yellow), calculated from the crystal structure J4*_O2 (blue, Platform in) or from opened models (red and green, Platform out) are shown. In a Δ55 structure (PDB code 1GX5), the thumb domain is slightly more open than in Δ21 structures (9). The red thumb model was obtained by applying this rotation to the thumb of J4*_O2 three times and moving the linker out of the catalytic core, which is the minimal deformation for the enzyme to accommodate a double-stranded RNA. The green thumb model was obtained by moving the linker further out starting from the red thumb model, which is more consistent with the elongation process occurring. The approximate expected position of the initiation platform buttressing the base of the priming nucleotide is circled in the blue thumb crystal structure and the red thumb minimally opened model.