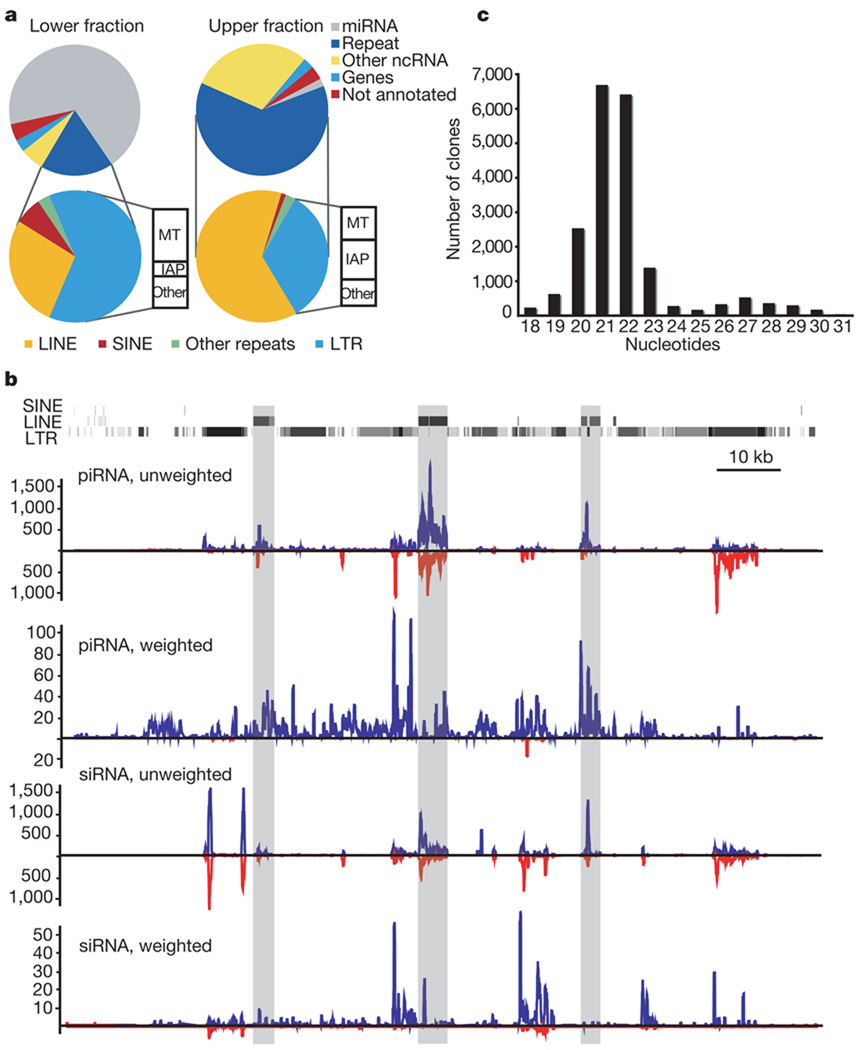
Figure 1. Both piRNA and siRNA systems control transposons in mouse oocytes.
a, Small RNA libraries from 19–24-nucleotide (lower fraction, left) and 24–30-nucleotide RNAs (upper fraction, right) were deeply sequenced. Reads were assigned an annotation as previously described12. The fraction of reads in each category is depicted. The repeat-annotated small RNAs were designated as LINE, SINE (short interspersed element), LTR and other, with the LTR category further divided between MT (mouse transcript), IAP and other. b, A representative piRNA locus on chromosome 10 is shown with the content of LINE, SINE and LTR fragments. Shading indicates the degree of match to the consensus element. Frequency plots for piRNAs and siRNAs from this locus are shown below. ‘Unweighted’ plots each match to the cluster; ‘weighted’ normalizes each match, dividing by its genomic frequency4. Blue and red lines indicate small RNAs mapping to the upper and lower genomic strand, respectively. c, RNAs from 19–30 nucleotides were deeply sequenced. Reads matching the piRNA cluster shown in b were used to construct a frequency plot by length.