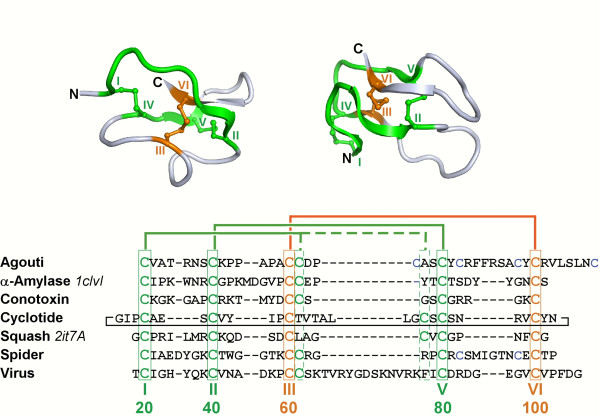
Figure 1.
Top, cartoon representations: 3D structures of the squash inhibitor EETI-II (left, PDB ID 2it7A) and of the α-amylase inhibitor AAI (right, PDB ID 1clvI). The two-disulfide macrocycle is shown in green and the penetrating disulfide is shown in orange. The structures are in similar but different orientations. Bottom, sequences: selected knottins from 7 families. Families, Swiss-Prot IDs (PDB IDs for the above structures), from top to bottom: Agouti-related, ASIP_HUMAN; α-amylase inhibitor, IAAI_AMAHP (1clvI); Conotoxin1, CXO7C_CONMA; Cyclotide, CYO1_VIOOD, Serine protease inhibitor1, ITR2_ECBEL (2it7A); Spider, TOG4B_AGEAP; Virus1, Q89632_CVS. Knotted cysteines are boxed and numbered at the bottom. Roman numbers indicate the order of knotted cysteines while Arabic numbers indicate standard numbering used in the KNOTTIN database. Cysteine connectivities are shown as thick lines on top of sequences. Sequences were aligned using the Knoter1D tool [1].