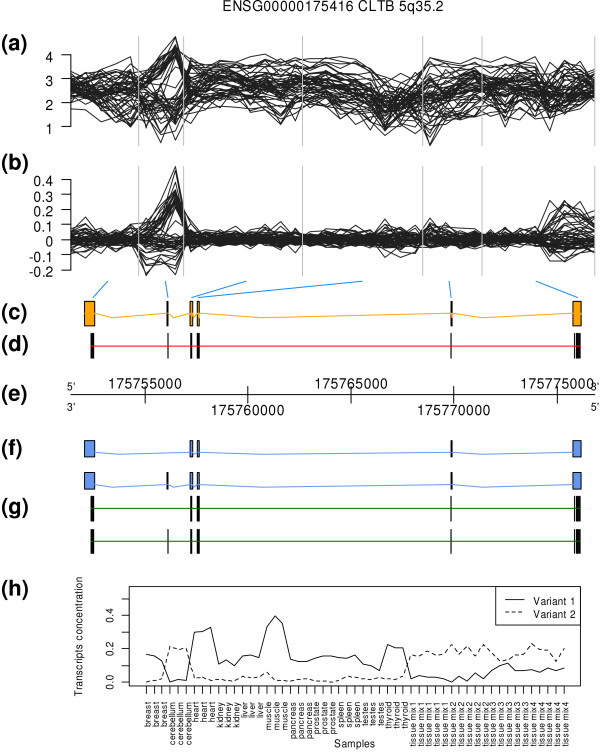
Figure 4.
Results for CLTB gene (Affymetrix sample dataset of human tissues, reverse strand). (a) Log of the intensities for all the probes within the gene. Each line corresponds to a different sample. (b) Residuals of the RMA normalization model. The residuals for some exons are much larger than for others. One of these exons is predicted as a cassette alternative splicing event (skipped exon). The vertical bars detach the different exons (or piece of exons) that correspond to each group of probes. (c) Structure of the CLTB gene, i.e. the exons that appear in any of its transcripts. (d) Locations of the Exon probes. It can be observed that each exon has one or several probes mapped to them. (e) Genomic positions of the probes. (f) Structure of the different transcripts of CLTB gene in Ensembl release 51. Two different transcripts are represented. (g) Predicted structure using SPACE. The predicted number of transcripts is two and their predicted structure is shown. Predicted transcripts match exactly the Ensembl annotated ones. (h) Estimated concentration of each of the two transcripts (variant 1 and 2) in each sample. Variant 2 is predominant in cerebellum tissue as shown by RT-PCR in [8]. These graphs were generated using GenomeGraphs [30].