Abstract
Background
The molecular mechanisms that modify genome structures to give birth and death to alleles are still not well understood. To investigate the causative chromosomal rearrangements, we took advantage of the allelic diversity of the duplicated p1 and p2 genes in maize. Both genes encode a transcription factor involved in maysin synthesis, which confers resistance to corn earworm. However, p1 also controls accumulation of reddish pigments in floral tissues and has therefore acquired a new function after gene duplication. p1 alleles vary in their tissue-specific expression, which is indicated in their allele designation: the first suffix refers to red or white pericarp pigmentation and the second to red or white glume pigmentation.
Results
Comparing chromosomal regions comprising p1-ww[4Co63], P1-rw1077 and P1-rr4B2 alleles with that of the reference genome, P1-wr[B73], enabled us to reconstruct additive events of transposition, chromosome breaks and repairs, and recombination that resulted in phenotypic variation and chimeric regulatory signals. The p1-ww[4Co63] null allele is probably derived from P1-wr[B73] by unequal crossover between large flanking sequences. A transposon insertion in a P1-wr-like allele and NHEJ (non-homologous end-joining) could have resulted in the formation of the P1-rw1077 allele. A second NHEJ event, followed by unequal crossover, probably led to the duplication of an enhancer region, creating the P1-rr4B2 allele. Moreover, a rather dynamic picture emerged in the use of polyadenylation signals by different p1 alleles. Interestingly, p1 alleles can be placed on both sides of a large retrotransposon cluster through recombination, while functional p2 alleles have only been found proximal to the cluster.
Conclusions
Allelic diversity of the p locus exemplifies how gene duplications promote phenotypic variability through composite regulatory signals. Transposition events increase the level of genomic complexity based not only on insertions but also on excisions that cause DNA double-strand breaks and trigger illegitimate recombination.
Background
An exciting challenge of biological research has been to understand phenotypic diversity within a species, which affects virtually every organ and cell type. In plants, this intraspecific diversity is often readily visible in the size, shape, color and number of flowers, fruits and seeds. Diversity can occur in every region of the gene, in coding regions or in regulatory sequences including upstream promoter and enhancer sequences, 5' and 3' UTRs and regulatory introns [1,2]. Changes in regulatory regions affecting allele expression and transcript amount can be simple, such as small and large indels, or more complex, such as transposon insertions and structural rearrangements. Molecular mechanisms responsible for the sequence modifications are replication errors, recombination and transposition. Although the majority of allelic variation is due to nucleotide polymorphisms, phenotypic differences can be caused by epigenetic modifications such as DNA methylation [3,4].
Sequence comparisons among inbred lines revealed that maize is a highly polymorphic species regarding genes and intergenic space [5]. Consequently, maize lends itself to studying the molecular basis of phenotypic variation. As an example for a detailed allelic analysis, we chose the p1 locus, which maps to the short arm of chromosome 1, for several reasons: 1) p1 produces a visible, quantitative phenotype in different tissues (Figure 1A). It encodes a R2R3 Myb-like transcription factor that activates the structural genes c2, chi1 and a1 of the phlobaphene biosynthesis pathway (Figure 1B) [6]. Phlobaphenes, which are reddish flavonoid pigments, accumulate in male and female floral organs. 2) The p1 gene is dispensable for the organism. Loss-of-function or change-of-function alleles will not be eliminated from the gene pool. 3) The p1 gene is characterized by its tremendous allelic diversity. More than hundred p1 alleles with distinct spatial and temporal expression pattern are reported although only few are molecularly defined [7]. 4) Approximately 2.75 million years ago, the p1 gene arose as a tandem duplication of the p2 gene [8]. If the p2 gene is the older of the two, it probably is the orthologous gene copy to the p3 gene on chromosome 9 because maize arose by allotetraploidization about 5 mya [9,10]. Therefore, we also refer to p2 as ortholog and p1 as paralog. The p2 gene is not involved in phlobaphene pigmentation, but like p1, is a QTL for maysin production (Figure 1B) [11].
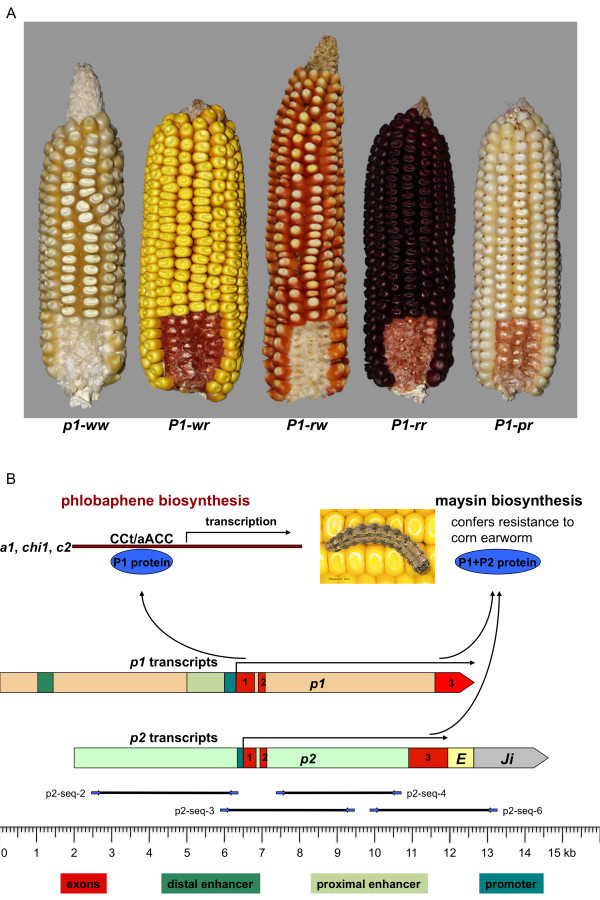
Figure 1.
Phenotype, structure and function of p1 and p2 alleles. (A) p1 alleles. p1 alleles are phenotypically classified according to their pericarp (i.e. the outer layer of the kernel) and cob glume pigmentation. The first suffix of the allele designation refers to pericarp color (r for red, w for white or colorless, p for patterned), while the second suffix stands for glume color. (B) Genomic structure of p1 and p2 alleles. p1 alleles with the exception of P1-rr and p2 alleles contain three almost identical exons depicted in red. They also share a basal promoter sequence. However, enhancer regions that were identified in P1-rr are absent from p2. p regulatory elements are drawn in shades of green. The p2 coding sequence is flanked by fragments of Eninu (E) and Ji (J) retrotransposons. The P1 protein is a Myb-like transcriptional regulator of the phlobaphene biosynthesis pathway that activates transcription of the target genes a1, chi1 and c2 by binding to a CCt/aACC site. p1 is also a QTL for maysin accumulation. Maysin is a C-glycosyl flavone found in silk that confers resistance to corn. earworm (Helicoverpa zea, Boddie). While p2 does not control phlobaphene pigmentation, it is involved in maysin synthesis. PCR primers designed for p2 allele cloning and sequencing are indicated. Corn earworm image courtesy of Marlin E. Rice, Iowa State University Department of Entomology http://www.ent.iastate.edu/.
Phlobaphene pigmentation is most readily visible in the pericarp, i.e. the outer layer of the kernel, and the cob glumes. Traditionally, p1 alleles are phenotypically categorized and named based on expression in these tissues. The p1 gene designation is followed by a two-letter suffix that refers to pericarp and cob color, respectively. For instance, the P1-rr allele exhibits red pericarp and red cob glume pigmentation while the P1-rw allele has red pericarp and white or colorless cob glumes (Figure 1A). Each phenotypic p1 group may consist of structurally very different alleles. Only few p1 alleles have been structurally determined of which only a small number has been completely or partially sequenced. P1-rr4B2 [12] and P1-rw1077 [13] are single copy genes that both were introgressed into the inbred line 4Co63. This inbred line contains a loss-of-function p1-ww allele. P1-wr in inbred line B73 is a multi-copy allele, consisting of 11 P1-wr tandem repeats that are flanked by p2/p1 and p1/p2 hybrid genes upstream and downstream of the cluster, respectively [10]. A large retroelement cluster is inserted in the 3' UTR of the p1/p2 hybrid gene.
The p1-ww alleles do not encode a functional P1 transcription factor; therefore pericarp as well as cob glumes are colorless (Figure 1A). While loss-of-function alleles often result in deleterious or even lethal conditions for the organism, non-functional p1 alleles do not cause any impairment that would reduce the fitness of the mutant plant. The p1-ww alleles can vary in origin and structure. Most of the structurally known p1-ww alleles are derived from P1-rr by transposon insertions and/or excisions. The p1-ww1112 null allele, for example, arose from a transposon-induced recombination event between the 5.2-kb direct repeats, which led to the deletion of the entire coding sequence [14]. However, the origin of p1-ww allele in the inbred line 4Co63 is not known, but p1-ww[4Co63] is often used in genetic crosses. Brink, for instance, introgressed more than 100 p1 alleles in the inbred line 4Co63 [7]. Knowledge of the p1-ww[4Co63] sequence could help clarify whether p1-ww[4Co63] is derived from P1-rr [12], P1-wr [10,15], P1-rw [13] or even a different p1 allele and provide further insights into other intermediates of chromosomal rearrangements.
To shed light on the origin of p1 allelic variability, we analyzed here three p1 alleles in their chromosomal context, namely p1-ww[4Co63], P1-rr4B2 and P1-rw1077. First we resolve the structural organization of these p1 alleles and their corresponding p2 alleles on the single-nucleotide sequence level. Subsequently we compare their sequences also to the recently sequenced P1-wr[B73] cluster [10] to find large and small scale nucleotide polymorphisms that enable us to infer mechanisms for genome rearrangements. In particular, we focus on evolutionary changes in p1 alleles that occurred in the putative distal enhancer region and in the 3' UTRs.
Results
The structural organization of p1-ww[4Co63] and linked p2 gene
A partial genomic lambda library was constructed using EcoRI-digested 4Co63 DNA. Filters were screened with the probe p15 (Figure 2), which hybridizes to a distal enhancer region thought to be present in all p1 alleles at the time of p1-ww[4Co63] cloning. A lambda clone containing 11,073-bp genomic DNA was isolated and sequenced [GenBank:HM454274]. Interestingly, this sequence is 99.7% identical to the displaced p1/p2[B73] 3' UTR and its 3' intergenic region, starting from an EcoRI recognition sequence in the retrotransposon Opie, and ending in an EcoRI site in the retroelement Shadowspawn (Figure 2).
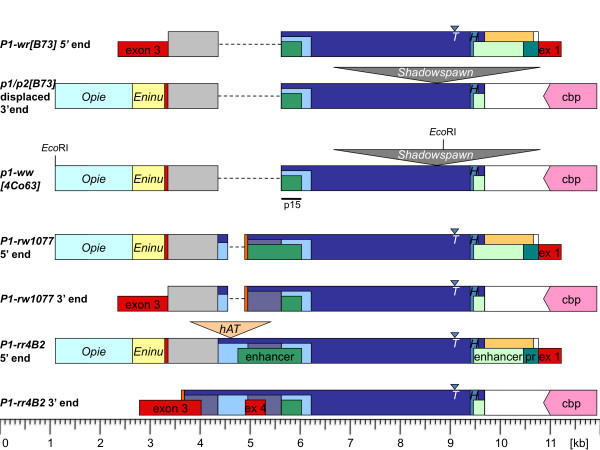
Figure 2.
Schematic alignment of p non-coding sequences. p non-coding regions, which include regulatory elements (distal and proximal enhancer and promoter), can be located upstream, downstream or on both sides of p coding sequences. However, only cis-elements in the upstream non-coding regions have been shown to have regulatory function. P1-wr[B73] is the only p1 allele shown with a multi-gene structure, as indicated by exon 3 stemming from a neighboring P1-wr[B73] repeat. Due to the tandem array, each P1-wr[B73] coding region is flanked by this non-coding sequence. The p1/p2[B73] 5' end (not shown) is identical to the P1-wr[B73] 5' end. The p1/p2[B73] 3' end is detached from the p1/p2[B73] coding region due to retrotransposon insertions. Opie (light turquoise rectangle), Eninu (light yellow rectangle) and the displaced former 3' UTR of p1/p2[B73] (red bar) are displayed. A shadowspawn retroelement (not drawn to scale) is inserted further downstream. p1-ww[4Co63] is virtually identical to the displaced p1/p2[B73] 3' end. P1-rw1077 and P1-rr coding regions are bordered by large direct repeats (blue rectangles). Both 5' and 3' repeats are depicted with their adjacent exons indicating the omitted coding region. p1-ww[4Co63] and the p1/p2[B73] 3' end are flanked by the same 5' sequences as P1-rr and P1-rw1077 (namely Opie, Eninu and the displaced 3' UTR). In addition, p1-ww[4Co63] and p1/p2[B73] share the 3' flanking sequences with P1-rr and P1-rw1077 as indicated by the downstream gene coding for a calmodulin-binding protein (cbp, pink pentagon). Notice that P1-wr[B73], P1-rw1077 and P1-rr vary mainly in a region that contains a fragmented MULE insertion (purple rectangle) and a sequence specified by light blue rectangles, which is partially and completely duplicated in P1-rw1077 and P1-rr, respectively. The orange bar stands for the MULE TIR that is missing in the 5' P1-rr repeat. Regulatory elements, i.e. distal and proximal enhancer and basal promoter, depicted in shades of green, were only determined for P1-rr. In other p genes or alleles, green rectangles merely refer to sequence similarity with P1-rr. Functional homology has not been investigated. The dark yellow rectangle represents a further Mu-like transposon that overlaps with the proximal enhancer and promoter region. A Heartbreaker MITE (blue bar) is part of the proximal enhancer region. Transposable elements of various families are shown as triangles above the schematic sequences. Notice the missing Tourist MITE in p1-ww[4Co63] and the displaced 3' end of p1/p2[B73]. Exons (red rectangles) are added with the intention to facilitate orientation. The EcoRI cloning sites and the hybridization site of probe 15 used to screen the lambda library for p1-ww[4Co63] sequencing are marked.
The extensive similarity with the p1/p2[B73] 3' end and intergenic region together with the identical Shadowspawn insertion suggests that both sequences continue to be similar past the end of the lambda clone. To confirm this assumption, we extended the sequence by genomic PCR from the Shadowspawn element to neighboring genes that are unrelated to p and therefore do not participate in potential p recombination events. PCR primer pairs were designed based on the equivalent P1-wr[B73] cluster, and PCR products were cloned and sequenced [GenBank:HM454275]. The analysis of 6,587 bp revealed that 4Co63 and B73 are virtually identical in this sequence; they consist of the 3' end of the Shadowspawn retroelement, a gene encoding a calmodulin-binding protein, part of a gene encoding a protein of unknown function, and intergenic regions (Figure 2). The calmodulin-binding protein, which in 4Co63 measures 361 aa, is 7 aa larger than in B73 and also contains two amino acid substitutions due to 3 indels and 10 SNPs. Based on maize and other EST data, this gene is transcribed and is very conserved in grass species such as rice, sorghum and barley.
Although we could not find any remnants of p1 in the 3' flanking regions, we also needed to extend the analysis to the 5' flanking regions. We knew that a functional p2 gene, as visualized in silk browning reactions, still had to be present. The browning reaction of freshly cut back silk correlates with silk maysin concentration and is therefore induced by both p genes [16]. To extend the sequences of p1-ww[4Co63] up to the p2 gene, we used a genomic PCR approach by taking advantage of existing p2, p2/p1[B73] and p1/p2[B73] sequences for the primer design. We sequenced a total of 10,753 bp that include the complete p2 and flanking sequences [GenBank:HM454271] (Figure 3). No p1 fragments or traces were detected. The p2[4Co63] and p2[p1ww1112] alleles with their flanking sequences are 94.56% identical. They differ in 57 SNPs and multiple indels, of which the largest consists of a Heartbreaker MITE insertion of 317 bp in the second intron. Because the 1,008 bp coding sequences of the two duplicated genes only vary by one synonymous substitution each in exon 2 and 3, their deduced P2 protein sequences of 335 aa are identical (Additional file 1: Supplemental Figure S1). RT-PCR experiments confirm that p2[4Co63] is expressed in silk tissue as predicted by the silk browning reaction (data not shown).
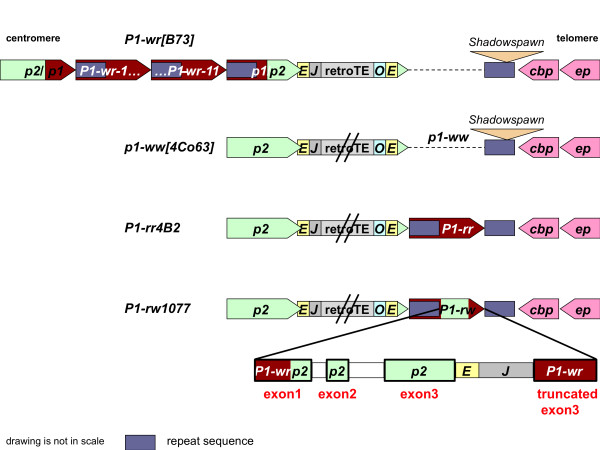
Figure 3.
Schematic alignment of p1 and p2 alleles. P1-wr[B73], p1-ww[4Co63], P1-rw1077 and P1-rr4B2 alleles are represented as dark red and their corresponding p2 alleles as light green pentagons with their apex pointing in the direction of transcription. The retrotransposon cluster downstream of p2 that was only entirely sequenced in P1-wr[B73] is 68 kb in size. In all remaining lines only the end sequences of the cluster that consist of Eninu (E), Ji (J) and Opie (O) retroelement fragments were determined. Downstream flanking genes, which encode a calmodulin-binding protein (here labeled as cbp) and an expressed protein (ep), are illustrated as pink pentagons. Both genes are conserved in grasses and arranged in opposite transcriptional orientation to p alleles. Purple rectangles refer to a sequence that most likely originated as the 3' intergenic region of an ancestral p gene. Due to the duplication event that gave rise to p1 and p2, this region also became present upstream of p1 alleles (see Figure 2 for details). The triangles on top of some purple rectangles stand for retrotransposon (Shadowspawn) insertions. The coding regions of p2/p1[B73], p1/p2[B73] and P1-rw1077 consists of p2 and p1 sequences. In P1-rw1077, the 5' end of exon 1 can be attributed to p1 while the 3' end of exon 1, exon 2 and 3 and flanking retrotransposon sequences are derived from p2 [13]. Interestingly, the retroelements are followed by a truncated P1-wr[B73] exon 3. Note that the drawing is not in scale.
The sequence 246 bp downstream of the p2 stop codon is composed of a partial Eninu retroelement of 540 bp followed by a Ji retrotransposon, which covers the remaining 1,322 bp of the available sequence (Figure 3 and 4B). The 4Co63 p2 allele differs from the B73 p2 sequences extracted from the p2/p1[B73] and p1/p2[B73] alleles in many SNPs and indels including transposon insertions, suggesting that p1-ww[4Co63] may not have arisen from P1-wr[B73] by recombination events in a direct lineage.
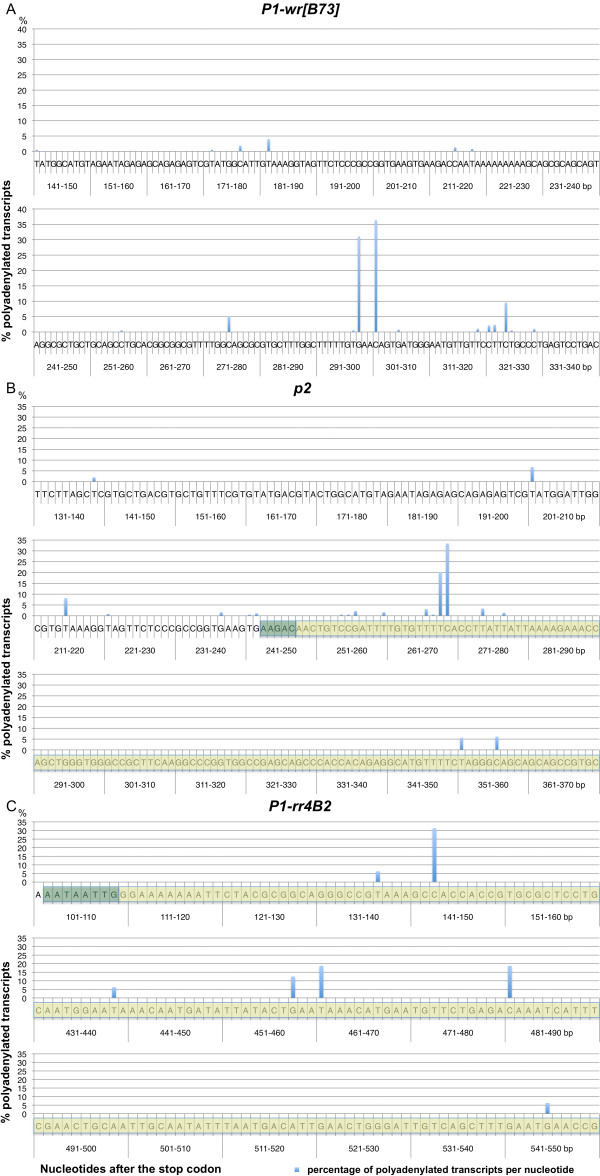
Figure 4.
Polyadenylation sites of P1-wr[B73], P1-rr4B2 and p2. Figures A to C display the partial 3' UTR sequences of P1-wr, p2 and P1-rr, respectively. Numbers on the x-axis refer to the nucleotide position after the stop codon. The y-axis reflects the percentage of transcripts that were polyadenylated at each nucleotide. (A) P1-wr[B73] polyadenylation sites. 18 polyadenylation sites are shown for P1-wr transcripts isolated from pericarp tissue. The most frequently used polyadenylation sites are 298 nt and 301 nt downstream of the stop codon. (B) Polyadenylation sites of p2 are located in a flanking LTR region. The sequence highlighted in yellow belongs to an Eninu retrotransposon that inserted 1.38 mya in p2. The target site duplication (TSD) AAGAC upon insertion is highlighted in green. The first and second bp of the LTR are mutated from TGCTGT to AACTGT. The majority of transcripts are polyadenylated within the LTR sequence. (C) Polyadenylation sites of P1-rr4B2 are located in a flanking MULE sequence. The MULE sequence is highlighted in yellow and the potential TSD in green. Note the sequence gap 161-430 bp after the stop codon, which does not contain any polyadenylation sites. All P1-rr4B2 transcripts are polyadenylated within the transposon.
Interestingly, the retrotransposons at the 3' end of p2, namely Eninu and Ji, and at the 5' end of the above described "p1-ww" lambda clone, namely Opie and Eninu, are identical to the retroelement cluster of P1-wr[B73] in sequence, insertion site and consequently target site duplications. Although we did not clone and sequence the complete retroelement cluster in p1-ww[4Co63] it is most likely that both clusters in 4Co63 and B73 are identical, at least in their initial transposition of Eninu and their nested insertions of Ji and Opie (Figure 3).
In brief, whereas p2 is present and functional in the 4Co63 inbred line, p1 coding and regulatory sequences are missing with the exception of the distal enhancer region. The structure of the p1-ww[4Co63] allele does not unambiguously point to a single known p1 allele where p1-ww[4Co63] is derived from, although, mechanistically, unequal crossing over between flanking sequences of the p1 gene could have been involved as discussed below.
The structural organization of P1-rr4B2 and linked p2 gene
How does the sequence arrangement of P1-wr[B73] and p1-ww[4Co63] including their flanking genes compare to P1-rr4B2, a p1 single-copy allele that produces red pericarp and red glumes? P1-rr4B2 contains two large repeats flanking the coding sequence, which are about 5.2 kb in size [6,12,17]. Interestingly, the sequence upstream of the 5' large repeat contains fragments of Opie and Eninu retroelements inserted in the same position as in p1-ww[4Co63] and p1/p2[B73] as described above (Figures 2 and 3). Likewise, Eninu is bordered by the detached p 3' UTR sequence of 78 bp. Subsequently, P1-rr is highly similar to a single P1-wr[B73] copy with few exceptions: the upstream regulatory region is more complex in P1-rr than in P1-wr[B73] (Figure 2) and both sequences diverge shortly after the stop codon (see below). By sequencing two plasmids, SA206 and PA103, which contain the 3' large repeat and are derived from lambda clones used for the isolation of P1-rr 5' and coding sequences [18] (see Methods), we extended our P1-rr sequence analysis by 8,923 bp past the 3' large repeat and into flanking genes [GenBank:HM454276]. By aligning both large repeats we found 14 polymorphisms including the insertion of a transposable element of 1,616 bp in the 5' repeat (Figure 2). This element is flanked by 8-bp direct repeats (CCAGTGAG), which is typical for transposons of the hAT superfamily. The 3' large repeat following the upstream regulatory sequence resembles p1-ww[4Co63] but does not contain the Shadowspawn retrotransposon insertion (Figures 2 and 3). Furthermore, the final 4,341 bp of the plasmid insert, not related to P1-rr, are highly similar to the equivalent p1-ww[4Co63] and P1-wr[B73] sequences (Figures 2 and 3). The 3' flanking sequence contains one complete gene and one partial gene in opposite transcriptional orientation compared to P1-rr. The first gene, which is separated from P1-rr by 1,175 bp (measured from the end of the 3' P1-rr repeat to the stop codon), encodes the 4Co63-type calmodulin-binding protein consisting of 361 amino acids. No more than 609 bp of intergenic sequence divide the first from the second gene, of which only the final two exons are present in the plasmid clone.
The P1-rr sequence analysis revealed that P1-rr is located between the retroelement cluster and the gene encoding a calmodulin-binding protein. Most interestingly, the corresponding site in p1-ww[4Co63] and P1-wr[B73] is empty, i.e. this region does not contain a p1 gene copy (Figure 3). Based on the first maize p2 allele that was isolated from a line which contains the p1-ww1112 allele [8] we assume that a functional p2 allele of P1-rr4B2 is present upstream of the retroelement cluster because p1-ww1112 and P1-rr4B2 are both derived from the same allele. Furthermore, the p2[p1-ww1112] allele ends in Eninu and Ji retroelement fragments exactly like p2[4Co63] and p1/p2[B73] suggesting structural similarity among these alleles. Therefore, we decided to extend our sequence analysis to the p2 allele that is linked to P1-rr4B2. We used the same genomic PCR strategy as described above to clone and sequence 10,423 bp of p2[P1-rr4B2] [GenBank:HM454272]. Indeed, the alignment of p2[P1-rr4B2] with p2[p1-ww1112] showed no SNPs but only four 1-bp indels that are not part of exons or putative regulatory sequences. Hence both p2 alleles are coding for an identical P2 protein (Additional file 1: Supplemental Figure S1). As expected, p2[P1-rr4B2] is also flanked by Eninu and Ji retroelement sequences. Introgression of P1-rr4B2 in 4Co63 probably included p2 as well because p2[P1-rr4B2] differs from p2[4Co63].
The structural organization of P1-rw1077 and linked p2 gene
The P1-rw allele specifies red pericarp and colorless cob glumes (Figure 1A). In general, the structure of P1-rw1077 resembles P1-rr4B2 [13]. P1-rw1077 is a single-copy gene, which consists of a coding region flanked by two 6.3-kb direct repeats (Figure 2). The coding sequence of P1-rw1077 is chimeric in nature. While the 5' UTR is similar to p1, the remaining coding region and adjacent Eninu and Ji retroelements (spanning about 6.9 kb) are p2-like (Figure 3) [13]. Sequence alignments establish that the p2 fragment is more closely related to p2[P1-rr4B2]/[p1-ww1112] than to p2[4Co63]. Interestingly, the Ji retrotransposon is followed by a truncated P1-wr-like exon, which is not included in the P1-rw1077 transcript. This organization of sequences suggests that P1-rw1077 originated from a gene conversion event between p1 and p2 [13]. The P1-rw1077 sequence upstream of the 5' large repeat is very similar to the corresponding P1-rr4B2 region, suggesting that both alleles occupy the same chromosomal location. We confirmed this by PCR-amplification and sequencing of a 1,651-bp fragment that connects the 3' large repeat of P1-rw1077 with the gene encoding the calmodulin-binding protein. P1-rw1077 was introgressed in 4Co63, and indeed the 3' end of the intergenic region between the 3' large repeat and the neighboring gene is indistinguishable from 4Co63. Since P1-rw1077 and P1-rr4B2 occupy the same chromosomal position we wanted to find out whether the similarity extends to the region upstream of the retrotransposon cluster (Figure 3). We performed genomic PCR as described above to amplify and subsequently sequence 11,313 bp [GenBank:HM454273] that are 99.8% identical to the p2[4Co63] sequence. The 18 SNPs and 3 short indels, which are distributed over a consensus sequence of 10,703 bp, are not included in the p2[P1-rw1077] coding sequence and consequently do not alter the P2 protein sequence (Additional file 1: Supplemental Figure S1). The polymorphisms between p2[P1-rw1077] and other p2 alleles suggest that this p2 sequence was introgressed together with P1-rw1077 into the 4Co63 background. This implies that the p2 part of P1-rw1077, which is p2[P1-rr4B2]-like, is derived from a p2 source other than p2[P1-rw1077]. The p2[P1-rw1077] 3' sequence is also flanked by Eninu and Ji retroelement fragments, linking p2[P1-rw1077] to P1-rw1077 across the retrotransposon cluster (Figure 3).
In summary, while p1 alleles can be located on both sides of the retroelement cluster (Figure 3), complete p2 alleles have so far only been found upstream of the retroelement cluster.
Evolution of a putative distal enhancer by non-homologous end-joining and transposition
Because all known p1 alleles produce almost identical P1 proteins (Additional file 1: Supplemental Figure S1), differential expression of p1 alleles could have evolved through changes in regulatory sequences, which control time-and tissue-specific p1 expression [17]. Sequences containing regulatory elements are only determined for P1-rr [19,20], but based on sequence similarities have likely the same function in other p1 alleles as well. While all known p1 alleles share the P1-rr promoter and proximal enhancer sequences, they differ in the sequence arrangement that contains the distal P1-rr enhancer. Comparing putative distal enhancer regions of p1 alleles reveals that the single P1-wr[B73] gene carries the simplest and therefore possibly the most ancestral form, which is confirmed by the presence of an almost identical enhancer region at the 3' intergenic region of the p2 gene in a wild relative of maize (Teosinte accession Zea mays ssp. parviglumis) [8]. Complexity of this chromosomal region increased with P1-rw1077 and then P1-rr4B2. Therefore, we can use the changes in sequence organization to explain the origin of the P1-rw1077 and P1-rr4B2 enhancer region within the P1-wr repeat context, where the 3' end of one copy equals the 5' end of the downstream copy.
P1-rw1077 is a complex allele that must have been shaped by multiple recombination events (Figure 3) [13]. Interestingly, the sequence following the p2 portion resembles the junction of two P1-wr[B73] copies in a head-to-tail arrangement, suggesting that P1-rw1077 arose from P1-wr-like tandem repeats. The sequence similarity between P1-rw1077 and P1-wr[B73] starts with the truncated exon 3. P1-rw1077 and P1-wr[B73] are identical until they diverge 1,001 bp after the truncated P1-rw1077 exon 3. The next 734 bp of P1-rw are of mixed origin and mostly unrelated to P1-wr. P1-rw continues its homology with P1-wr 3' of the 734-bp insertion, but not at the sequence where both alleles deviate from each other. Instead, the P1-rw sequence downstream of the insertion is identical to the region of P1-wr upstream of the insertion, i.e. the insertion is flanked by a 203-bp repeat sequence. The sequence after the point of divergence originated from an unknown Mu-like transposable element (MULE) in reverse orientation (Additional file 2: Supplemental Figure S2). Based on BLASTN searches, the sequence consists of two MULE fragments that in a putative autonomous MULE would be separated by approximately 444 bp (for additional information on this new MULE family in the maize B73 genome see Additional file 3). While the initial 279 bp, starting with the TIR (GGAAAAAATT...), are derived from the MULE 3' end, the remaining 446-bp fragment stems from a sequence partially encoding the C-terminus of the MULE transposase. The final 9 bp (AACCTATGT) of the 734-bp insertion may represent filler DNA (see bottom panel of Figure 5). The 9-bp fragment is identical to a P1-wr[B73] sequence, which is located 27 bp downstream of the point of P1-rw1077 and P1-wr[B73] alignment. Filler DNA, which is usually found at repair sites of DNA double-strand breaks can be simple as described here or complex, consisting of a patchwork of multiple sequences. Filler DNA is associated with non-homologous end-joining and is usually derived from nearby sequences of either end of the break [21-23] (for additional information on the mechanism of NHEJ see Additional file 4).
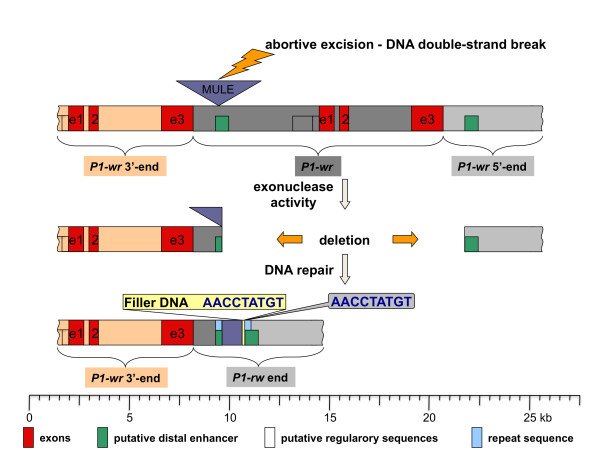
Figure 5.
Model for the origin of the P1-rw1077 enhancer region by non-homologous end-joining. The bottom bar illustrates the P1-rw1077 end adjacent to a P1-wr repeat. The top bar represents schematically a full length P1-wr repeat flanked by partial P1-wr copies on both sides (drawn as dark grey, tan and light grey rectangles, respectively). The green rectangle indicates part of a sequence that was proven to have enhancer function in P1-rr. A model demonstrating the progression from a P1-wr sequence to a unique P1-rw1077 enhancer structure is briefly outlined. A DNA double-strand break was initiated by the excision of a Mu-like transposable element (purple triangle) and the resulting gap was expanded by exonuclease activity. DNA repair was accomplished by non-homologous end-joining as implied by the presence of filler DNA (yellow rectangle). The filler DNA AACCTATGT is derived from a sequence close to the deletion end point (see light grey balloon). The neighboring 725 bp (purple rectangle) originated from the excised MULE. The light blue rectangles specify sequences that are duplicated due to the deletion of less than a full-length P1-wr repeat. Note that the terminal P1-wr copy only contains the 5' end and is similar to the 3' large repeat of P1-rr and P1-rw, which does not have any gene function and therefore should be considered as intergenic region. Due to the unknown origin of P1-rw1077 and P1-rr, we use the designation P1-wr also for P1-wr-like alleles that share regulatory and coding regions with P1-wr[B73].
P1-rr is structurally more complex than P1-rw1077 and a single P1-wr[B73] gene. P1-rw1077 and P1-rr contain the same MULE fragments and filler DNA inserted in exactly the same sequence position. However, a 1.2-kb duplication in P1-rr that partially includes the fragmented MULE suggests that P1-rr is derived from P1-rw1077. This duplication results in the addition of a fourth exon, which is unique to P1-rr. A closer look at the P1-rr 3' UTR may help to shed light on the evolution of the P1-rr enhancer region (see bottom panel of Figure 6).
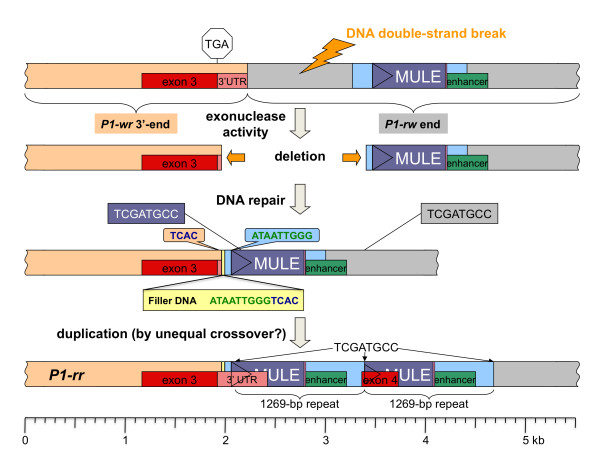
Figure 6.
Model for the origin of the unique P1-rr 3' end and enhancer region by non-homologous end-joining and unequal crossover. The bottom bar represents the unique P1-rr 3' end with tandem direct repeats. The top bar shows schematically the junction sequence of two P1-wr repeats drawn as tan and grey rectangles. The P1-wr copy at the right side was modified into a P1-rw-like sequence as outlined in Figure 5 (see MULE, filler DNA and repeats depicted as purple, pink and light blue rectangles, respectively). The green rectangle stands for part of a sequence, which was shown to have enhancer function in P1-rr. A model explaining the conversion from a P1-wr and P1-rw1077 sequence to a unique P1-rr structure is briefly described. A DNA double-strand break of unknown cause was expanded by exonuclease activity. DNA repair occurred by non-homologous end-joining as evidenced by filler DNA (yellow rectangle). While the initial 9 bp (ATAATTGGG) of the filler DNA stem from a sequence 55 bp downstream of the deletion end point (see light blue balloon), the adjacent 4 bp TCAC correspond to a sequence 21 bp upstream of the insertion site (tan balloon). The MULE fragment contains an 8-bp sequence TCGATGCC also found 1269 bp further downstream (shown on top of the grey rectangle). Unequal crossover at the 8-bp site resulted in the duplication of the 1269-bp sequence in tandem fashion and addition of a fourth exon.
Whereas P1-rw1077 and P1-rr are identical in the initial 3' UTR, they diverge 35 bp following the stop codon. The next 13 bp (ATAATTGGGTCAC) in P1-rr originated from two separated P1-rw1077 sequences, 1,410 bp apart, implying a deletion event in P1-rr compared to P1-rw1077. The 13-bp (ATAATTGGGTCAC) insertion in P1-rr can be assigned to P1-rw1077 sequences upstream and downstream of the deletion site. ATAATTGGG is duplicated 59 bp downstream and includes the first two bp of the MULE TIR. Obviously, the adjacent TCAC occurs frequently within the P1-rw1077 sequence. However, the closest TCAC can be located 21 bp upstream of the insertion site. The 13-bp P1-rr sequence subsequent of the point of divergence with P1-rw1077 is suggestive of filler DNA, indicating that a previous DNA double-strand break in P1-rw1077 was restored by the NHEJ pathway. A tandem duplication of 1,269 bp that comprises the majority of both MULE fragments and 3' flanking enhancer sequences generated the current P1-rr 3' end and enhancer region.
In summary, DNA double-strand breaks in a P1-wr-like tandem array were probably repaired by NHEJ events that could have resulted in the rearrangements and duplications of enhancer-carrying sequences and consequently in novel p1 alleles as discussed below.
The p alleles differ in their 3' UTRs and polyadenylation sites
Although p1 and p2 share nearly the same coding sequences, their downstream sequences vary remarkably (Additional file 5: Supplemental Figure S3). Sequence alignments of p1 and p2 alleles revealed that the P1-rr4B2 and p2 divergence from P1-wr[B73] is caused by transposon insertions. In P1-rr4B2, a Mu-like element was placed 109 bp downstream of the stop codon probably due to a deletion event (see above), and p2 alleles are followed by an Eninu retroelement 248 bp after the stop codon. The insertion sites close to the stop codon raise the question whether these transposable elements eliminated the transcription termination signals and the polyadenylation sites in the P1-rr4B2 and p2 3' UTRs. In general, the 3' UTR is also important for post-transcriptional regulation such as microRNAs and translational control, and gain or loss of cis elements within the 3' UTR could contribute to allelic diversity. Therefore, we decided to map the polyadenylation sites of these alleles.
The P1-wr[B73] coding sequence is not flanked by transposons, and its 3' UTR probably represents the original 3' UTR structure of all p alleles before transposon modifications. We performed 3' RACE experiments to identify the P1-wr[B73] 3' UTR using three different gene-specific primers (p1 race 5'-1 to 3, see Table 1) and three independent pericarp tissue sources. RNA was extracted 20 days after pollination (DAP) from plants that contain the P1-wr[B73] cluster. Since the combination of different primers and tissues gave the same result we merged the data sets as shown in Figure 4A. Interestingly, we detected 18 polyadenylation sites in P1-wr[B73] that are spanning 189 nt from 141 to 329 nt after the stop codon. However, 36% of transcripts are polyadenylated 301 nt after the stop codon. The P1-wr[B73] cluster consists of 11 P1-wr tandem repeats and a p2/p1[B73] hybrid gene that differ by few polymorphisms in their transcribed regions. As these 18 polyadenylation sites are not specific for a particular repeat, the polymorphisms apparently do not affect the polyadenylation signals.
Table 1.
PCR primers used in this report to amplify p2, p1 and flanking sequences and for 3' RACE.
| p2-amplification | |
|---|---|
| p2-seq-2-for | CGCGTGATTGGCTCCTCGGATTACC |
| p2-seq-2-rev | TTTTCGGGACTGCGTGCATTGACTC |
| p2-seq-3-for | GGACGGCGGAGGAGGACCAGTTA |
| p2-seq-3-rev | TGATAGCTCGCCAGTTTTGTTAGAGGAT |
| p2-seq-4-for | ATGGCTGGCCCGATCGGTTGAGAGTTA |
| p2-seq-4-rev | CCGCTGCTGCTGTTGGGCTGGTTCG |
| p2-seq-6-for | CGCGCATTGGCTAGCTTCCCTGTT |
| p2-seq-6-rev | GCTTGTCGCCGGTCTCCATCTCCT |
| p2 3' RACE | |
| p2 race 5'-1 | CTCCCGCCGGTGAAGTGAAGACAA |
| p2 race 5'-2 | CGGACCGATCAGACAGACAGACAGACCA |
| p2 race 5'-3 | GCCGTGGGTGCTGGAGCCGATAGA |
| p1 3' race | |
| p1 race 5'-1 | GAGGAGGGGCCCAGCAGCGAGGAC |
| p1 race 5'-2 | GCCGCCGAGCCGCTGGAAGTTGC |
| p1 race 5'-3 | TCACCGGACCGATCAGACAGACCAACCA |
| 3' flanking sequences | |
| shad-gene1-2 f | AGGGCAGCGTCTCCACCATCTA |
| shad-gene1-2 r | CAAAACCCTCAACCCCGTATTCTC |
| shad-gene1-3 f | CGTTGCTTCACTCCCCCGTTAGA |
| shad-gene1-3 r | GCTGATCAATGCGCTCGTCGTTC |
| r2-gene1-2a f | CGATGCATGCACTGTCCGATTTA |
| r2-gene1-2 r | CGGCGGTGGCGGCTACTTCT |
| r2-gene1-1 f | GCTACCCTCAATGCATGCACTGTCCT |
| r2-gene1-1 r | CGCGCTTCACGGGCTCACCAA |
| gene 1-2 f | GGACGAGCGGGACGAGGCGGTTAC |
| gene 1-2 r | GTCTGGCACTTCTTCCCCTGTCCT |
RT-PCR results indicated that p2[4Co 63], p2[P1-rr4B2] and p1/p2[B73] are expressed in silk tissue. Accordingly, we carried out 3' RACE experiments using total RNA from silk and three different primers (p2 race 5'-1 to 3, see Table 1), which hybridize to exon 3 of p2. The RNAs extracted from p2[4Co63], p2[P1-rr4B2] and p1/p2[B73] lines produced almost identical results with all PCR primers, which allows us to combine the data for ease of presentation (Figure 4B). We found 19 polyadenylation sites in a 218-bp interval that is located between 139 and 356 nt past the p2 stop codon. Whereas seven minor polyadenylation sites (adding up to 21% of the total events) are upstream of the retrotransposon, 12 sites lie within the LTR, including the major site (33% of polyadenylated p2 mRNAs), which is 269 nt from the stop codon and 22 nt into the LTR. The sequence alignment between p2 and P1-wr[B73] shows that the main polyadenylation site of P1-wr[B73] is 87 bp past the point of p2 and P1-wr[B73] divergence. The equivalent p2 fragment was displaced by retroelement insertions, and therefore cannot serve its original function. Nevertheless, p2 was able to recruit alternative polyadenylation signals and sites located mostly in the Eninu LTR.
Subsequently, we performed 3' RACE experiments on P1-rr4B2 total RNA extracted from silk and one primer binding (p2 race 5'-3, see Table 1) to exon 3. This exon contains the 3' UTR of the alternatively spliced P1-rr4B2 transcript, which encodes the functional P protein. We sequenced significantly fewer clones compared to P1-wr[B73] and p2 and obtained fewer polyadenylation sites. While polyadenylation sites are distributed over 403 nt from 143 to 545 nt measured from stop codon, the first site is used most often (31%) (Figure 4C). All seven polyadenylation sites are located in the MULE fragments, two within the TIR, the remainder in the transcribed part. Due to the partial deletion of the former 3' UTR alternative polyadenylation signals and sites had to be employed from adjacent sequences as described above. Note that the MULE borders P1-rr4B2 in opposite transcriptional orientation. A transcript from an intact member of the same MULE family could therefore produce antisense RNA that is complementary to P1-rr4B2 mRNA.
Discussion
Models for the evolution of p1 alleles
A distinguishing feature of the p locus is its tremendous allelic diversity, which makes it a preferable locus to study evolutionary changes and chromosomal dynamics on a larger and smaller scale. Although the grass family arose by an ancient whole genome duplication (WGD) event [24], the p gene has only a single ortholog in rice and sorghum, indicating that one copy was lost from the paleoploid ancestral genome. However, the more recent allotetrapolidization event, which formed the ancestor of maize about 5 mya [9], gave rise to two p copies located in the homoeologous regions of chromosomes 9 and 1. The copy on chromosome 1 was then duplicated in tandem 2.75 mya, thereby evolving into the current p2 (ortholog) and p1 (paralog) genes [8,10]. The bulk of retrotranspositions in most grasses occurred more recently. A series of nested insertions that split approximately 80 bp of the p 3' UTR occurred between 1.4 to 0.2 mya [10]. Although retroelements are highly repetitive in the genome, insertions of retroelements in a nested fashion create unique sequence junctions and become chromosomal markers [25]. However, we do not know whether the retroelements transposed into the paralog or ortholog repeat, or maybe even into a later-generated copy. A model proposed for the evolution of single-copy alleles states that the retroelement insertion occurred in the 3' UTR of p2, thereby separating p1, which turned into P1-rr and P1-rw [8,17] (Figure 3). In contrast, transposition into the 3' UTR of p1 retains the repeat structure and allows the amplification of additional copies by unequal crossover as suggested for the evolution of the multi-copy P1-wr[B73] allele [10] (Figure 3). Theoretically, only few recombination events are needed to transfer p1 and p2 sequences across the retroelement cluster. Therefore multi-copy alleles in a tandem array could have been created upstream, downstream or on both sides of the cluster simultaneously. These intermediate structures that enable us to discover the step-by-step evolution of all p alleles might still exist in the maize germplasm. Our current analysis allows us to present new and refined models for the evolution of p1-ww[4Co63], P1-rr4B2 and P1-rw1077.
Models for the evolution of p1-ww[4Co63]
p1-ww is a null allele because p1 specific sequences such as coding-, promoter-and proximal enhancer sequences are absent in 4Co63 (Figure 3). Is it possible that p1-ww[4Co63] represents a haplotype where the tandem duplication of the ancestral p gene never took place? According to this hypothesis, the nested retroelement insertions in 4Co63 that are identical to alleles containing p1 and p2 sequences must have happened before the p duplication event. However, since the p duplication occurred 2.75 mya, 1.37 million years before the first retroelement insertion, we can disregard this possibility [8,10]. Thus, the alternative explanation that a functional p1 allele was deleted to give rise to p1-ww[Co63] is more likely. The p1-ww[4Co63] structure does not reveal the functional p1 allele(s) and their deletion or recombination events that resulted in the current null allele. Considering that p1 alleles are located on both sides of the retroelement cluster, multiple recombination events could have occurred to create the p1-ww[4Co63] allele.
One possible scenario for the origin of p1-ww in inbred line 4Co63 is that the null allele, which carries a functional p2, is derived from P1-wr[B73]. While unequal crossover among repeat sequences can lead to an increase of copy numbers, the alternative outcome is a reduction of repeats. During the evolution of the P1-wr[B73] allele, unequal crossover between the flanking genes of the cluster, namely p2/p1[B73] and p1/p2[B73], could have caused the deletion of all P1-wr[B73] repeats, and would still have generated a functional p2 gene (Figure 7A). However, p2 in 4Co63 differs by various SNPs and indels from the corresponding p2/p1[B73] and p1/p2[B73] sequences of P1-wr[B73], indicating that P1-wr[B73] might not be the immediate progenitor for p1-ww[4Co63].
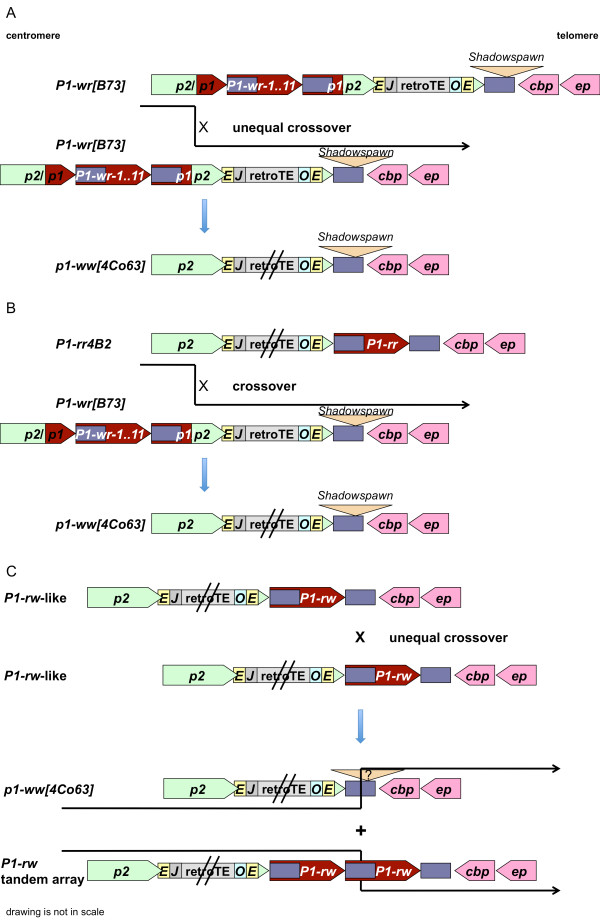
Figure 7.
Models for the origin of p1-ww[4Co63]. Various models may help explain the evolution of p1-ww[4Co63]. Notice that the same color code is used here as in Figure 3, which illustrates schematic alignment of p1 and p2 alleles. (A) Expansion and contraction of the p locus generates P1-wr[B73] and p1-ww[4Co63], respectively. A simplified representation of the P1-wr[B73] cluster is shown. Unequal crossing over between p2/p1[B73] (or p2) and p1/p2[B73] could have caused the deletion of the internal P1-wr[B73] repeats. Such a recombination event could have occurred any time during the evolution of the cluster, independent of the P1-wr copy number, retroelement insertions at the 3' end and the deletion at the p2/P1-wr junction sequence. (B) Similarly, alignment along the retrotransposon cluster and crossover between P1-rr and P1-wr[B73] could have generated the p1-ww[4Co63] structure. (C) A misalignment at the repeat sequences of P1-rw-like alleles, which contained the simple P1-wr[B73] distal enhancer region, and unequal crossover could have led to a duplication or deletion event giving rise to p1-ww[4Co63]. However, this model does not account for the Shadowspawn retroelement in p1-ww[4Co63].
p1-ww[4Co63] also could have evolved by a recombination event that involved two different p1 alleles. Unequal crossing over between p2 of P1-rr4B2 and p1/p2[B73] of P1-wr[B73] could have generated the current p1-ww[4Co63] structure and could have restored the p2 copy (Figure 7B). Even then, the deletion of the original paralog would have been derived from the P1-wr[B73] allele. Nevertheless, this could not have happened recently (on an evolutionary time scale) because of sequence polymorphisms in the participating alleles. Interestingly, both p1-ww[4Co63] and P1-wr[B73] carry as a signature the Shadowspawn retroelement in the same position, indicating that p1-ww[4Co63] most likely derived from P1-wr[B73] in multiple steps.
In addition, we can envision a P1-rw-like allele, which is similar to P1-wr[B73] in the distal enhancer structure. Such a P1-rw allele has been described [17]. An unequal crossover between the large repeats flanking the coding regions duplicates the p1 gene or deletes the coding sequences, resulting in the p1-ww[4Co63] structure (Figure 7C). This scenario resembles the origin of p1-ww1112 [14]. However, this model does not directly account for the Shadowspawn retroelement in p1-ww[4Co63]. All models demonstrate the complexity of the p locus and reveal the countless possibilities for recombination to occur whenever paralogous sequences are present.
Model for the evolution of P1-rw1077 and P1-rr4B2, with focus on regulatory sequences
Despite the repeat structure of the P1-wr[B73] cluster, a single P1-wr[B73] copy has the least complicated p1 allele composition, followed by P1-rw1077 and then P1-rr4B2. We hypothesize that P1-rw1077 originated from a P1-wr-like tandem array (Figure 5) because P1-rw1077 comprises a sequence fragment downstream of the p2 section that is virtually identical with the junction sequence of two P1-wr[B73] repeats in a head-to-tail assembly. This P1-wr tandem array could have been located on either side of the retroelement cluster.
A plausible sequence of events is as follows. A Mu-like element inserted into one of the P1-wr repeats 1,204 bp after exon 3 of the previous copy. Then an aberrant transposition event (abortive excision event) of this MULE caused a DNA double-strand break that enabled exonucleases to digest the unprotected DNA ends (Figure 5) thereby extending the gap into the adjacent P1-wr repeat. The deletion would have included MULE sequences (about 3.5 kb compared to a putative autonomous element) and almost the entire length of a P1-wr repeat (more than 12 kb). Non-homologous end-joining [21-23,26], copying a 9-bp sequence (AACCTATGT) that is located 27 bp downstream from the deletion endpoint, must have repaired the break. Due to the nature of tandem repeats, the large deletion described above results in small repeats of 203 bp that are flanking the MULE fragments. Interestingly, this duplication is part of a 1.2-kb sequence that contains the enhancer element of P1-rr.
A single P1-wr-like allele downstream of the retroelement cluster that is flanked by large repeats due to the retroelement insertion in p2 could have been converted into a tandem array by unequal crossover between the large repeats (Figure 7C). Gene conversion events then could have transferred the altered region that originated at the 3' large repeat to the 5' large repeat where the distal enhancer sequence functions [17]. Alternatively, P1-rw1077 arose from P1-wr repeats upstream of the retrotransposon cluster. The sequence 3' of this cluster, which corresponds to the 3' intergenic region of p2 as found in the P1-wr[B73] cluster and p1-ww[4Co63], is nearly identical with the 5' end of a P1-wr[B73] repeat over a stretch of 5.2 kb (Figure 2 and Additional file 6: Supplemental Figure S4A). Due to this sequence similarity, a recombination event between p1-ww[4Co63] and the proposed P1-rw1077 precursor could have occurred that positioned P1-rw1077 downstream of the retrotransposon cluster (Additional file 6: Supplemental Figure S4A). This arrangement assumes that the P1-rw1077 allele resembles p1-ww[4Co63] at the 5' end. After the recombination break point, P1-rw1077 has to be closer to P1-wr[B73] because, based on our model, P1-rw1077 is derived from P1-wr. Indeed, a sufficient amount of polymorphisms between the p1-ww[4Co63] and P1-wr[B73] alleles enables us to verify the predicted structure and to place the possible recombination site between 567 and 713 bp after the point of p1-ww[4Co63] and P1-wr[B73] alignment. Further recombination/gene conversion events contributed to the evolution of the present P1-rw1077 allele.
The presence of the MULE fragments and filler DNA in P1-rr in exactly the same sequence context as in P1-rw1077 agrees with our model that P1-rr continued to evolve from P1-rw1077. In our model for the origin of P1-rr, we propose a second DNA double-strand break (DSB) that occurred in P1-rw1077 in between the stop codon and the MULE insertion (Figure 6). In contrast to the first DSB, there is no evidence for the participation of a TE, leaving the cause for the DSB unknown. Exonuclease activities expanded the gap until both ends were joined in a NHEJ fashion by synthesizing two short DNA pieces (filler DNAs) from sites close to the deletion end points into the repair site. The DSB repair caused a deletion of 1,410 bp across the repeat junction that spanned almost the entire sequence from the stop codon to the MULE fragments. Interestingly, this intermediate P1-rr structure can be found at the 3' end of P1-rr1088, P1-rrCFS36 and P1-rwCFS342 [17].
The 5' transposon fragment happened to contain an 8-bp sequence close to the TIR (55-62 bp) that is present 1,269 bp further downstream as well. Unequal crossover between those 8 bp resulted in a tandem direct duplication of this 1,269 bp sequence. Accordingly, the final 318 bp of exon three, being part of the repeat, were replicated, too. A sequence at the 3' end of the first repeat was adopted as a splice acceptor site thereby generating a fourth exon. Although alternative splicing of exon 1, 2 and 4 has been reported, the protein product is of unknown function or may not have any function at all [27].
This putative evolutionary pathway explains how the P1-wr 3' UTR was almost entirely replaced by a MULE, how the fourth exon unique to P1-rr was generated and how the 1,269 bp SalI fragment containing the P1-rr distal enhancer was nearly completely duplicated (the initial 175 bp of the enhancer region are missing from the first repeat). Subsequently, gene conversion events could have placed part of the modified enhancer sequence of the downstream copy to the upstream large repeat [17]. Alternatively, if this P1-rr module arose on the P1-wr[B73] side of the retroelement cluster as we also discussed for P1-rw1077, then a recombination event between p1-ww[4Co63] and the P1-rr ancestor could have transferred the P1-rr end to a position downstream of the retrotransposon cluster (Additional file 6: Supplemental Figure S4B). The crossing over took place in the 595 bp sequence between the duplicated MULEs, which is why the repeat structure of P1-rr at the 5' end differs from the 3' end whereas they are identical in P1-rw1077. Lastly, a 1.6 kb hAT-like transposable element inserted 340 bp upstream of the MULE or 159 bp 5' of the enhancer region. This transposition did not occur in P1-rr1088 [17]. Taken together, the novel distal enhancer structure of P1-rr could be the result of a MULE insertion and excision, deletion and repair by NHEJ, and duplication and deletion by recombination. This series of events from P1-wr to P1-rr confirms the sequential model of P1-rw and P1-rr evolution based on phylogentic analysis [17].
Function of the enhancer region rearrangements on p1 expression
When the p1 paralog was formed, it probably included the complete p coding sequence and the basal promoter that controls p expression in silk tissue. Then the paralog acquired two additional regulatory sequences adding equally to the basal expression in pericarp and glume. The enhancer sequences were identified and tested in transient and transgenic plants using P1-rr fragments fused to a GUS reporter gene [19,20]. A 1-kb sequence adjacent to the promoter contains a regulatory sequence termed proximal enhancer while a 1.2-kb fragment further upstream includes a distal enhancer (Figure 2). The proximal enhancer region corresponds mostly to a truncated MULE that captured part of a host gene in between the TIR [10]. The proximal enhancer region and the basal promoter sequence are virtually identical in all sequenced p1 alleles to date (Figure 2). In contrast, the distal enhancer region varies in all p1 alleles as described above. Therefore, we hypothesize that the different spatial and temporal expression patterns of p1 alleles are caused by distinct distal enhancer regions [17]. The distal enhancer as defined in P1-rr is located within a 1,269-bp SalI fragment [19,20], out of which 671 bp are derived form the Mu-like transposon (Figure 2). Although this MULE fragment is missing in P1-wr[B73], transgenes constructed from P1-wr upstream regulatory sequences linked to P1-rr cDNA produced red pericarp and cob glumes in transgenic plants [28], indicating that the enhancer sequence is included in the 589-bp region downstream of the MULE. Since this 589-bp region is duplicated in P1-rr, P1-rr has two enhancer sites that are separated by the MULE fragment. Additional P1-rr alleles, namely P1-rr1088 and P1-rrCFS36, were shown to have the same enhancer structure as P1-rr4B2 with exception of the missing hAT insertion in P1-rr1088 [17]. Therefore, the hAT transposable elements inserted in the upstream copy of the enhancer region of P1-rr4B2 and P1-rrCFS36 obviously do not disrupt the enhancer sequence and function. Compared to P1-rr, P1-rw1077 has a deletion of 381 bp in the upstream repeat, which causes the loss of cob glume pigmentation [13]. Interestingly, two additional P1-rw alleles, P1-rwCFS302 and P1-rwCFS342, lack the entire upstream repeat and the MULE fragment, thus having the identical enhancer arrangement as a single P1-wr[B73] copy [17]. Taken together, the analysis of three P1-rr and three P1-rw alleles revealed that P1-rr alleles contain two copies of the specific enhancer sequence while P1-rw alleles only have one [17]. Interestingly, this region coincides with a tissue-specific DNase I-hypersensitive site that remains closed in pericarp tissue of P1-pr, a silenced epiallele of P1-rr4B [29]; the P1-pr phenotype is shown in Figure 1A. It was reasoned that the upstream enhancer repeat that is missing in P1-rw1077 controls the glume-specific expression in a position-dependent manner, since the identical enhancer region is located 671 bp further downstream [13]. An alternative explanation was prompted by the fact that p1 expression in pericarp is weaker and delayed in P1-rw1077 compared to P1-rr. We hypothesize that the transcriptional strength of p1 alleles is correlated with the enhancer copy number, which is supported by similar findings in human upstream enhancers [30]. Consequently, P1-rw1077 produces less P1 protein than P1-rr in all expressing tissues. Also, each p1 allele is not expressed uniformly in female and male floral tissues within a plant. For example in P1-rr, p1 transcription is usually higher in pericarp than in cob glumes [15].
Therefore, we propose that the presence of only one distal enhancer site in the P1-rw1077 allele results in weak expression in pericarp tissue but no expression in cob glumes. Due to the duplication of the enhancer sequence as outlined in our model, p1 transcription in pericarp and glume tissue was equally elevated such that p1 is strongly expressed in pericarp and weakly expressed in glumes, thereby giving rise to P1-rr alleles. Note that comparisons with P1-wr alleles are not appropriate due to their post-transcriptional silencing, which potentially is repeat induced [31]. This model is supported by an analysis of the spatial expression pattern in transgenic plants where various p1 constructs were expressed only in few p expressing tissues, resembling P1-rr or P1-rw phenotypes. It was shown in these transgenic plants that p1 expression follows a spatial hierarchy that begins with pericarp and continues with cob glumes, husk, silk, and tassel glumes in decreasing order [32,33]. For instance, if the transgenes had been expressed in only one tissue, then it would have had to be in pericarp, in the case of two tissues then in pericarp and glumes, and so on.
The p alleles differ in their 3' UTR
Polyadenylation is involved in many facets of mRNA metabolism including enhancement of mRNA stability, transport of mRNA from the nucleus into the cytoplasm, and regulation of mRNA translation. Although polyadenylation signals in plants are less conserved than in mammals [34], three signals were identified in maize, rice, and Arabidopsis: the far upstream element (FUE, located -150 to -35 nt upstream of the cleavage site), the near upstream element (NUE, situated -35 to -10 nt upstream of the cleavage site) and the cleavage element (CE, positioned -10 to +15 nt upstream and downstreams of the cleavage site) [35,36]. As we have shown above, a fragmented MULE was placed adjacent to the P1-rr4B2 stop codon possibly due to a NHEJ event. All mapped polyadenlation sites of the P1-rr4B2 transcript are located within the MULE sequence, indicating that P1-rr4B2 successfully recruited alternative polyadenylation signals in the transposon. Similarly, a Mu insertion in the 3' UTR of the rf2a locus also resulted in the adoption of new polyadenylation signals and sites [37]. Retroelements, the most common transposons in maize, also insert in 3' UTRs without disrupting polyadenylation as demonstrated above for the p2 alleles. Our results suggest that polyadenylation in maize is a highly dynamic process which despite its importance for the cell is not tightly regulated. The large amount of polyadenylation sites found in our analysis of P1-wr[B73] transcripts that do not contain a transposon insertion supports this conclusion. A genome-wide analysis of genomic and transcript data could shed light on the mechanism of polyadenylation in maize and could establish the proportion of genes that terminate in transposable elements. Interestingly, it has been shown that many polyadenlylation signals in human and mouse genes have been derived from transposable elements [38]. Besides polyadenylation signals, transcriptional as well as translational regulators have been identified in the 3' UTR of plant and animal genes, and their gain or loss could cause allelic diversity. For example, targets of microRNAs are often located in 3' UTRs [38,39].
Gene copying events promote allelic diversity
Recombination is crucial for the evolution of genomes [40,41]. In particular, the non-homologous recombination pathway is frequently used to repair DNA double-strand breaks in somatic plant cells [26]. Previously, we reported a probable NHEJ event involved in the formation of the P1-wr[B73] cluster [10] that produced a hybrid gene due to the ligation of deletion end points located within two genes. Similarly, deletions and repair by NHEJ in the above mentioned alleles could have resulted in the restructuring of an enhancer region and formation of a novel 3' UTR.
The exceptional allelic variation at the p locus prompts the question about its similarities and differences to genes that exhibit less variation. We propose that the main cause for the diversity might lie in tandem gene amplification [8,17,42,43]. Once a gene underwent an initial tandem duplication, multiple unequal recombination events can follow as seen in the P1-wr[B73] multi-gene cluster [10]. A single crossing over or gene conversion event between misaligned paralogous gene copies can generate many new alleles including deletion and amplification derivatives. Interestingly, in plants such events can occur mitotically and can be transmitted into the next generation, thereby increasing allelic variation [44]. This explanation then implies that other loci exhibiting an increased allelic variation are multi-copy genes as well. Indeed, the complex r1 locus in maize is analogous to p1 in many aspects. The r1 locus, which also encodes a transcription factor, confers bluish anthocyanin pigmentation to various vegetative and floral plant tissues. Two r1 alleles, R-st and R-r, are molecularly well characterized. R-st contains various r1 genes, four of which are in tandem orientation [45]. R-r consists of one complete and three truncated r1 genes that originated from tandem duplication [46,47]. Comparable to p1 in complexity, both alleles undergo recombination and transposition events creating numerous derivative alleles. Paralogous gene copies in maize were also found at the pl1 [48] and a1 loci [49]. Especially the prolamine gene family with nearly 50 copies distributed over several chromosomes exemplifies the outcome of gene duplications [50]. Actually, a large proportion of genes are tandemly duplicated in Arabidopsis, rice, and maize [51-53]. Considering the amount of paralogous sequences and their possibilities to recombine, a single reference genome providing just one allele can obviously not reflect this allelic potential of the maize genome. Not surprisingly, a recent genomic comparison between the B73 and Mo17 inbred lines [54] revealed a large quantity of copy number variations and presence/absence variations confirming previous results [55]. Nonetheless, epialleles remain invisible in a traditional sequence comparison. Allelic diversity studies as presented here are essential for our understanding of the remarkably dynamic maize genome.
Conclusion
Allelic diversity is the source for evolution and domestication. While allelic variation in wild species ensures the best possible adaption to changing environmental conditions, humans have profited from allelic pools in crop plants by selecting phenotypic variations that best meet their needs. Alleles differ most often in small-scale nucleotide polymorphisms but also in large-scale sequence rearrangements. Maize has been shown to be a highly polymorphic species well suited to study genome dynamics and the underlying molecular mechanisms. In particular, the maize p locus with its well-established genetic history offers a tremendous amount of ancient allelic variations, some representing intermediate steps in large-scale sequence rearrangements. The tandemly duplicated p1 and p2 genes encode virtually identical Myb-like transcriptional activators, but only p1 controls the accumulation of reddish flavonoid pigments in maize female and male floral organs. Because all P1 proteins are almost identical, the phenotypic variation must be due to p1 regulation. Therefore, this locus represents an ideal example of how genomic rearrangements can contribute to novel regulatory elements.
Here, we used targeted genome sequencing to apply comparative genomics to the maize genome. Sequence alignments of orthologs and paralogs of different genotypes of a single genomic region allow us to reconstruct the repair of double strand breaks from transposition events within gene copies and their flanking regions. Such drastic invasions of new sequence elements in flanking regions result in the de novo creation of regulatory elements involved in the transcriptional and post-transcriptional regulation of gene expression that differentiate gene copies in their function. Interestingly, sequence chimerism in the 3' untranslated portion of the mRNA gave rise to multiple poly-A addition signals with similar strength, indicating a more relaxed sequence restriction of the 3' processing machinery than previously believed.
Methods
Plant material
Seeds containing P1-rr4B2 and P1-rw1077 alleles, which were introgressed in a 4Co63 background, were thankfully provided by Tom Peterson, Iowa State University. The inbred lines B73 and 4Co63 carrying P1-wr and p1-ww alleles, respectively, were obtained from the Maize Genetics Cooperation Stock Center (maizecoop.cropsci.uiuc.edu) collection. Traditionally, p1 alleles are classified and named according to their pericarp and cob glume pigmentation, implicating that phenotypically similar but structurally different alleles share the same name. In this report, we use the inbred line where the p1 allele was originally described in as additional allelic designation such as P1-wr[B73] and p1-ww[4Co63]. Similarly, the inbred line will be used as allele description for p2, for example p2[4Co63]. Whenever the p2 source is unknown, the name of the linked p1 allele will be added to p2, such as p2[P1-rr4B2] and p2[P1-rw1077].
p1-ww[4Co63] isolation and sequencing
The inbred line 4Co63 contains a p1-ww allele, according to the colorless pericarp and cob phenotype of 4Co63 ears. We constructed a size-restricted lambda library using a lambda DASH II/EcoRI vector kit (Agilent Technologies) and EcoRI-digested 4Co63 genomic DNA. The lambda library was screened by hybridizing filters with probe 15 [12], which is derived from a distal enhancer fragment of P1-rr and is unique to p1 alleles. Two positively hybridizing lambda clones were isolated and subcloned into pBluescript II SK+ vectors (Agilent Technologies). Insert size and both end sequences of each clone were determined and found identical. A transposon minilibrary (Finnzymes) of one clone was constructed according to the manufacturer's instructions. Sequencing was performed with the ABI PRISM BigDye Terminator Cycle Sequencing Ready Reaction kit and an ABI 3730 capillary sequencer (Applied BioSystems). Sequence assembly and analysis were carried out using Lasergene (DNAstar) programs. Sequence gaps were closed by primer walking.
p2 amplification and sequencing
Genomic PCR was performed to amplify p2 alleles. PCR primers (see Table 1) (Figure 1B) were designed based on p2 sequences from p2[p1-ww1112] [8], p2/p1[B73] and p1/p2[B73] [10]. The PCR-amplified products were cloned into pGEM-T Easy vector (Promega). The individual clones were completely sequenced using primers that are spanning the entire repeat length (approximately one primer every 300 bp, primer sequences available upon request). The sequencing reactions were carried out with the ABI PRISM BigDye Terminator Cycle Sequencing Ready Reaction kit and analyzed on an ABI 3730 capillary sequencer (Applied BioSystems). The sequences were assembled and evaluated with the Lasergene software (DNAstar).
Sequencing of P1-rr 3' noncoding region and flanking genes
The majority of P1-rr sequence was determined in P1-ovov1114 (orange variegated pericarp and cob) that is derived from P1-vv. The Ac element of P1-vv located in the second intron excised and reinserted 161 bp further upstream in the opposite direction [56], still allowing a considerable amount of phlobaphene accumulation in pericarp and cob. Similarly, P1-rr4B2 is a P1-rr revertant that also originated from P1-vv by Ac excision. When not otherwise specified, we use P1-rr (without additional allele designation) to refer to functional P1-rr alleles that are derived from the same P1-vv. Two EcoRI fragments, isolated from P1-ovov1114, were cloned in lambda using two EcoRI recognition sites outside of P1-ovov1114. The third site was provided by the Ac transposon [18]. The 3' fragment of 14.5 kb was further divided in two plasmids, SA206 and PA103, which we gratefully received from Thomas Peterson. A transposon minilibrary of both plasmids (Finnzymes) was constructed as per the manufacturer's protocol. Clones were sequenced using transposon primers, ABI 3730 capillary sequencers, and the ABI PRISM BigDye Terminator Cycle Sequencing Ready Reaction kit (Applied BioSystems). Both plasmids contain 12418 bp non-overlapping P1-rr and 3' flanking sequences.
Amplification and sequencing of p1 intergenic region and flanking genes
Genomic PCR was performed to amplify p1 intergenic region and flanking genes. PCR primers (see Table 1) were designed based on corresponding sequences from B73 [10] and P1-ovov1114 (this study). The PCR products were cloned, sequenced and analyzed as described above for p2.
Amplification of 3' cDNA ends (3' RACE)
Total RNA was extracted from pericarp tissue 20 days after pollination and emerging silk with the RNeasy Plant Mini Kit (Qiagen). RNA was reverse-transcribed to cDNA using the GeneRacer Kit (Invitrogen) with the GeneRacer oligo(dT) primers. cDNA was PCR amplified with the GeneRacer 3' primer and a gene-specific primer (see Table 1). In general, 96 RT-PCR products per primer pair (but only 18 for P1-rr4B2 samples) were cloned into pGEM-T Easy vector (Promega) and sequenced with universal primers. DNA sequences were analyzed with Lasergene (DNAstar) software. Polyadenylation sites were only plotted in Figure 6A to 6C, when they occurred more than once.
Sequence annotation and GenBank accession numbers
The maize sequences were manually annotated using homology searches in various GenBank databases with multiple BLAST programs [57]. The sequences were submitted to GenBank and were assigned following accession numbers: p2[4Co63]: HM454271, p2[P1-rr4B2]: HM454272, p2[P1-rw1077]: HM454273, p1-ww[4Co63]: HM454274, p1-ww[4Co63] 3' flanking region: HM454275, P1-ovov1114 3' end: HM454276
Authors' contributions
WG and JM conceived of the study. WG designed and carried out the experiments. WG analyzed the data. WG and JM wrote the paper. All authors read and approved the final manuscript.
Supplementary Material
P protein alignment. Supplemental Figure S1 and figure legend.
MULE structure and location. Supplemental Figure S2 and figure legend.
Characterization of a Mutator-like transposable element. Supplemental description and Supplemental Tables S1-S3.
Mechanism of NHEJ. Supplemental information.
p 3' UTR sequence alignment. Supplemental Figure S3 and figure legend.
Recombination between modified P1-wr repeats and p1-ww[4Co63] can place p1 sequences across the retrotransposon cluster. Supplemental Figure S4 and figure legend.
Contributor Information
Wolfgang Goettel, Email: goettel@waksman.rutgers.edu.
Joachim Messing, Email: messing@waksman.rutgers.edu.
Acknowledgements
We thank Hugo Dooner for critical reading of the manuscript. We are grateful to Thomas Peterson for kindly providing SA206 and PA103 plasmids and P1-rw1077 seeds. This work was supported by the Selman A. Waksman Chair in Molecular Genetics to JM.
References
- Alonso-Blanco C, Mendez-Vigo B, Koornneef M. From phenotypic to molecular polymorphisms involved in naturally occurring variation of plant development. The International journal of developmental biology. 2005;49(5-6):717–732. doi: 10.1387/ijdb.051994ca. [DOI] [PubMed] [Google Scholar]
- Alonso-Blanco C, Aarts MG, Bentsink L, Keurentjes JJ, Reymond M, Vreugdenhil D, Koornneef M. What has natural variation taught us about plant development, physiology, and adaptation? The Plant cell. 2009;21(7):1877–1896. doi: 10.1105/tpc.109.068114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das OP, Messing J. Variegated phenotype and developmental methylation changes of a maize allele originating from epimutation. Genetics. 1994;136(3):1121–1141. doi: 10.1093/genetics/136.3.1121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robbins ML, Wang P, Sekhon RS, Chopra S. Gene structure induced epigenetic modifications of pericarp color1 alleles of maize result in tissue-specific mosaicism. PloS one. 2009;4(12):e8231. doi: 10.1371/journal.pone.0008231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gore MA, Chia JM, Elshire RJ, Sun Q, Ersoz ES, Hurwitz BL, Peiffer JA, McMullen MD, Grills GS, Ross-Ibarra J. et al. A first-generation haplotype map of maize. Science (New York, NY. 2009;326(5956):1115–1117. doi: 10.1126/science.1177837. [DOI] [PubMed] [Google Scholar]
- Grotewold E, Drummond BJ, Bowen B, Peterson T. The myb-homologous P gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset. Cell. 1994;76(3):543–553. doi: 10.1016/0092-8674(94)90117-1. [DOI] [PubMed] [Google Scholar]
- Brink RA, Styles ED. A collection of pericarp factors. Maize Genet Cooperation Newsletter. 1966;40:149–160. [Google Scholar]
- Zhang P, Chopra S, Peterson T. A segmental gene duplication generated differentially expressed myb-homologous genes in maize. The Plant cell. 2000;12(12):2311–2322. doi: 10.2307/3871231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swigonova Z, Lai J, Ma J, Ramakrishna W, Llaca V, Bennetzen JL, Messing J. Close split of sorghum and maize genome progenitors. Genome research. 2004;14(10A):1916–1923. doi: 10.1101/gr.2332504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goettel W, Messing J. Change of gene structure and function by non-homologous end-joining, homologous recombination, and transposition of DNA. PLoS genetics. 2009;5(6):e1000516. doi: 10.1371/journal.pgen.1000516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang P, Wang Y, Zhang J, Maddock S, Snook M, Peterson T. A maize QTL for silk maysin levels contains duplicated Myb-homologous genes which jointly regulate flavone biosynthesis. Plant Mol Biol. 2003;52(1):1–15. doi: 10.1023/A:1023942819106. [DOI] [PubMed] [Google Scholar]
- Lechelt C, Peterson T, Laird A, Chen J, Dellaporta SL, Dennis E, Peacock WJ, Starlinger P. Isolation and molecular analysis of the maize P locus. Mol Gen Genet. 1989;219(1-2):225–234. doi: 10.1007/BF00261181. [DOI] [PubMed] [Google Scholar]
- Zhang F, Peterson T. Comparisons of Maize pericarp color1 Alleles Reveal Paralogous Gene Recombination and an Organ-Specific Enhancer Region. Plant Cell. 2005;17(3):903–914. doi: 10.1105/tpc.104.029660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Athma P, Peterson T. Ac induces homologous recombination at the maize P locus. Genetics. 1991;128(1):163–173. doi: 10.1093/genetics/128.1.163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chopra S, Athma P, Peterson T. Alleles of the maize P gene with distinct tissue specificities encode Myb-homologous proteins with C-terminal replacements. The Plant cell. 1996;8(7):1149–1158. doi: 10.2307/3870358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levings CS, Stuber CW. A maize gene controlling silk browning in response to wounding. Genetics. 1971;69:491–498. doi: 10.1093/genetics/69.4.491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang F, Peterson T. Gene conversion between direct noncoding repeats promotes genetic and phenotypic diversity at a regulatory locus of Zea mays (L.) Genetics. 2006;174(2):753–762. doi: 10.1534/genetics.105.053942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Athma P, Grotewold E, Peterson T. Insertional mutagenesis of the maize P gene by intragenic transposition of Ac. Genetics. 1992;131(1):199–209. doi: 10.1093/genetics/131.1.199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sidorenko LV, Li X, Cocciolone SM, Chopra S, Tagliani L, Bowen B, Daniels M, Peterson T. Complex structure of a maize Myb gene promoter: functional analysis in transgenic plants. Plant J. 2000;22(6):471–482. doi: 10.1046/j.1365-313x.2000.00750.x. [DOI] [PubMed] [Google Scholar]
- Sidorenko L, Li X, Tagliani L, Bowen B, Peterson T. Characterization of the regulatory elements of the maize P-rr gene by transient expression assays. Plant Mol Biol. 1999;39(1):11–19. doi: 10.1023/A:1006172815663. [DOI] [PubMed] [Google Scholar]
- Wessler S, Tarpley A, Purugganan M, Spell M, Okagaki R. Filler DNA is associated with spontaneous deletions in maize. Proc Natl Acad Sci USA. 1990;87(22):8731–8735. doi: 10.1073/pnas.87.22.8731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorbunova VV, Levy AA. How plants make ends meet: DNA double-strand break repair. Trends Plant Sci. 1999;4(7):263–269. doi: 10.1016/S1360-1385(99)01430-2. [DOI] [PubMed] [Google Scholar]
- Yan X, Martinez-Ferez IM, Kavchok S, Dooner HK. Origination of Ds elements from Ac elements in maize: evidence for rare repair synthesis at the site of Ac excision. Genetics. 1999;152(4):1733–1740. doi: 10.1093/genetics/152.4.1733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salse J, Abrouk M, Bolot S, Guilhot N, Courcelle E, Faraut T, Waugh R, Close TJ, Messing J, Feuillet C. Reconstruction of monocotelydoneous proto-chromosomes reveals faster evolution in plants than in animals. Proc Natl Acad Sci USA. 2009;106(35):14908–14913. doi: 10.1073/pnas.0902350106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devos KM, Ma J, Pontaroli AC, Pratt LH, Bennetzen JL. Analysis and mapping of randomly chosen bacterial artificial chromosome clones from hexaploid bread wheat. Proceedings of the National Academy of Sciences of the United States of America. 2005;102(52):19243–19248. doi: 10.1073/pnas.0509473102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puchta H. The repair of double-strand breaks in plants: mechanisms and consequences for genome evolution. J Exp Bot. 2005;56(409):1–14. doi: 10.1093/jxb/eri025. [DOI] [PubMed] [Google Scholar]
- Grotewold E, Athma P, Peterson T. Alternatively spliced products of the maize P gene encode proteins with homology to the DNA-binding domain of myb-like transcription factors. Proc Natl Acad Sci USA. 1991;88(11):4587–4591. doi: 10.1073/pnas.88.11.4587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cocciolone SM, Chopra S, Flint-Garcia SA, McMullen MD, Peterson T. Tissue-specific patterns of a maize Myb transcription factor are epigenetically regulated. Plant J. 2001;27(5):467–478. doi: 10.1046/j.1365-313X.2001.01124.x. [DOI] [PubMed] [Google Scholar]
- Lund G, Das OP, Messing J. Tissue-specific DNase-I-sensitive sites of the maize p-gene and their changes upon epimutation. Plant J. 1995;7:797–807. doi: 10.1046/j.1365-313X.1995.07050797.x. [DOI] [PubMed] [Google Scholar]
- Blain M, Zeng Y, Bendjelloul M, Hallauer PL, Kumar A, Hastings KE, Karpati G, Massie B, Gilbert R. Strong muscle-specific regulatory cassettes based on multiple copies of the human slow troponin I gene upstream enhancer. Human gene therapy. 2010;21(1):127–134. doi: 10.1089/hum.2008.208. [DOI] [PubMed] [Google Scholar]
- Chopra S, Cocciolone SM, Bushman S, Sangar V, McMullen MD, Peterson T. The maize unstable factor for orange1 is a dominant epigenetic modifier of a tissue specifically silent allele of pericarp color1. Genetics. 2003;163(3):1135–1146. doi: 10.1093/genetics/163.3.1135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cocciolone SM, Nettleton D, Snook ME, Peterson T. Transformation of maize with the p1 transcription factor directs production of silk maysin, a corn earworm resistance factor, in concordance with a hierarchy of floral organ pigmentation. Plant Biotechnol J. 2005;3(2):225–235. doi: 10.1111/j.1467-7652.2005.00120.x. [DOI] [PubMed] [Google Scholar]
- Cocciolone SM, Sidorenko LV, Chopra S, Dixon PM, Peterson T. Hierarchical patterns of transgene expression indicate involvement of developmental mechanisms in the regulation of the maize P1-rr promoter. Genetics. 2000;156(2):839–846. doi: 10.1093/genetics/156.2.839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J, Geraghty D, Heidecker G, Hu N-T, Kridl JC, Rubenstein I. In: Genetic Engineering of Plants. Kosuge T, Meredith CP, Hollaender A, editor. New York: Plenum Press; 1983. Plant Gene Structure; pp. 211–227. [Google Scholar]
- Shen Y, Ji G, Haas BJ, Wu X, Zheng J, Reese GJ, Li QQ. Genome level analysis of rice mRNA 3'-end processing signals and alternative polyadenylation. Nucleic acids research. 2008;36(9):3150–3161. doi: 10.1093/nar/gkn158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu L, Ueda T, Messing J. The formation of mRNA 3'-ends in plants. Plant J. 1995;8(3):323–329. doi: 10.1046/j.1365-313X.1995.08030323.x. [DOI] [PubMed] [Google Scholar]
- Cui X, Hsia AP, Liu F, Ashlock DA, Wise RP, Schnable PS. Alternative transcription initiation sites and polyadenylation sites are recruited during Mu suppression at the rf2a locus of maize. Genetics. 2003;163(2):685–698. doi: 10.1093/genetics/163.2.685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feschotte C. Transposable elements and the evolution of regulatory networks. Nat Rev Genet. 2008;9(5):397–405. doi: 10.1038/nrg2337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghosh T, Soni K, Scaria V, Halimani M, Bhattacharjee C, Pillai B. MicroRNA-mediated up-regulation of an alternatively polyadenylated variant of the mouse cytoplasmic {beta}-actin gene. Nucleic acids research. 2008;36(19):6318–6332. doi: 10.1093/nar/gkn624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaut BS, Wright SI, Rizzon C, Dvorak J, Anderson LK. Recombination: an underappreciated factor in the evolution of plant genomes. Nat Rev Genet. 2007;8(1):77–84. doi: 10.1038/nrg1970. [DOI] [PubMed] [Google Scholar]
- Li J, Hsia AP, Schnable PS. Recent advances in plant recombination. Curr Opin Plant Biol. 2007;10(2):131–135. doi: 10.1016/j.pbi.2007.01.007. [DOI] [PubMed] [Google Scholar]
- Xu JH, Messing J. Amplification of prolamin storage protein genes in different subfamilies of the Poaceae. TAG Theoretical and applied genetics. 2009;119(8):1397–1412. doi: 10.1007/s00122-009-1143-x. [DOI] [PubMed] [Google Scholar]
- Das OP, Messing JW. Allelic variation and differential expression at the 27-kilodalton zein locus in maize. Molecular and cellular biology. 1987;7(12):4490–4497. doi: 10.1128/mcb.7.12.4490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das OP, Levi-Minzi S, Koury M, Benner M, Messing J. A somatic gene rearrangement contributing to genetic diversity in maize. Proceedings of the National Academy of Sciences of the United States of America. 1990;87(20):7809–7813. doi: 10.1073/pnas.87.20.7809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eggleston WB, Alleman M, Kermicle JL. Molecular organization and germinal instability of R-stippled maize. Genetics. 1995;141(1):347–360. doi: 10.1093/genetics/141.1.347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker EL, Robbins TP, Bureau TE, Kermicle J, Dellaporta SL. Transposon-mediated chromosomal rearrangements and gene duplications in the formation of the maize R-r complex. EMBO J. 1995;14(10):2350–2363. doi: 10.1002/j.1460-2075.1995.tb07230.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dooner HK, Kermicle JL. Structure of the R r tandem duplication in maize. Genetics. 1971;67(3):437–454. doi: 10.1093/genetics/67.3.437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pilu R, Piazza P, Petroni K, Ronchi A, Martin C, Tonelli C. pl-bol3, a complex allele of the anthocyanin regulatory pl1 locus that arose in a naturally occurring maize population. Plant J. 2003;36(4):510–521. doi: 10.1046/j.1365-313X.2003.01898.x. [DOI] [PubMed] [Google Scholar]
- Yandeau-Nelson MD, Xia Y, Li J, Neuffer MG, Schnable PS. Unequal sister chromatid and homolog recombination at a tandem duplication of the A1 locus in maize. Genetics. 2006;173(4):2211–2226. doi: 10.1534/genetics.105.052712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu JH, Messing J. Organization of the prolamin gene family provides insight into the evolution of the maize genome and gene duplications in grass species. Proceedings of the National Academy of Sciences of the United States of America. 2008;105(38):14330–14335. doi: 10.1073/pnas.0807026105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arabidopsis Genome I. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408(6814):796–815. doi: 10.1038/35048692. [DOI] [PubMed] [Google Scholar]
- International-Rice-Genome-Sequencing-Project. The map-based sequence of the rice genome. Nature. 2005;436(7052):793–800. doi: 10.1038/nature03895. [DOI] [PubMed] [Google Scholar]
- Messing J, Bharti AK, Karlowski WM, Gundlach H, Kim HR, Yu Y, Wei F, Fuks G, Soderlund CA, Mayer KF. et al. Sequence composition and genome organization of maize. Proc Natl Acad Sci USA. 2004;101(40):14349–14354. doi: 10.1073/pnas.0406163101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Springer NM, Ying K, Fu Y, Ji T, Yeh CT, Jia Y, Wu W, Richmond T, Kitzman J, Rosenbaum H. et al. Maize inbreds exhibit high levels of copy number variation (CNV) and presence/absence variation (PAV) in genome content. PLoS genetics. 2009;5(11):e1000734. doi: 10.1371/journal.pgen.1000734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai J, Li Y, Messing J, Dooner HK. Gene movement by Helitron transposons contributes to the haplotype variability of maize. Proc Natl Acad Sci USA. 2005;102(25):9068–9073. doi: 10.1073/pnas.0502923102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson T. Intragenic transposition of Ac generates a new allele of the maize P gene. Genetics. 1990;126(2):469–476. doi: 10.1093/genetics/126.2.469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic acids research. 1997;25(17):3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
P protein alignment. Supplemental Figure S1 and figure legend.
MULE structure and location. Supplemental Figure S2 and figure legend.
Characterization of a Mutator-like transposable element. Supplemental description and Supplemental Tables S1-S3.
Mechanism of NHEJ. Supplemental information.
p 3' UTR sequence alignment. Supplemental Figure S3 and figure legend.
Recombination between modified P1-wr repeats and p1-ww[4Co63] can place p1 sequences across the retrotransposon cluster. Supplemental Figure S4 and figure legend.