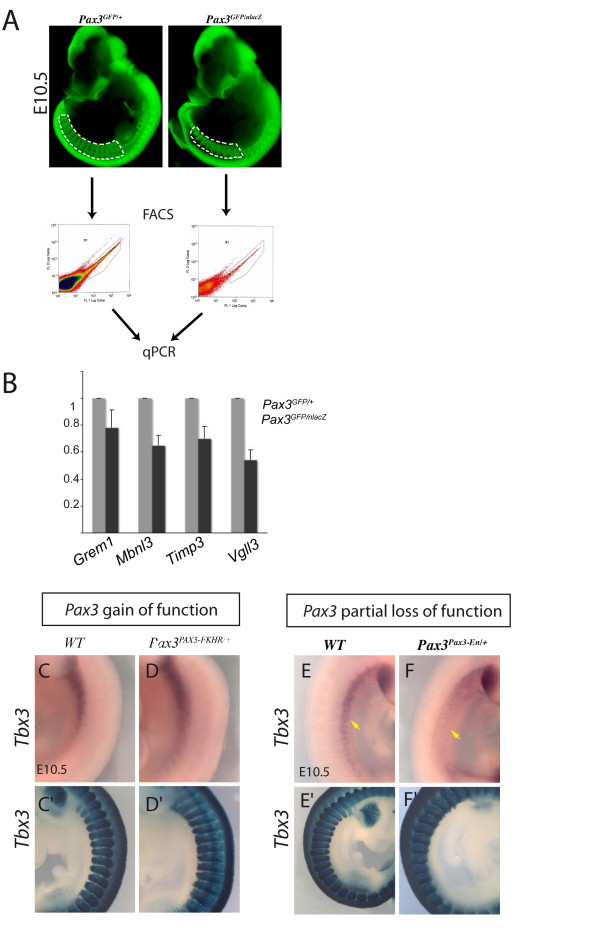
Figure 2.
Examples of validation on Pax3 loss of function genetic backgrounds. Genes that emerged from the microarray analyses as potential Pax3 targets from the gain of function screen were checked on Pax3 loss of function genetic backgrounds. (A-B) Quantitative PCR analysis of transcripts in Pax3-GFP cells separated by flow cytometry (FACS) from interlimb somites of Pax3GFP/+ and Pax3GFP/nlacZ embryos at E10.5 (A). The same number of cells were analysed for each genotype and the results for Gremlin1, Mbnl3, Timp3 and Vg113 transcripts are presented as histograms relative to the Pax3GFP/+ sample taken as 1 (B). In accordance with the microarray data, these genes are positively regulated by Pax3. (C-F) Whole mount in situ hybridization with a Tbx3 probe on control (C, E), Pax3PAX3-FKHR/+ gain of function (D) and Pax3Pax3-En/+ partial loss of function (F) embryos at E10.5, showing somites in the interlimb region. Tbx3 transcripts are high in the hypaxial somite domain, notably in more anterior somites, and their level depends positively on Pax3, as indicated by the microarray data. In the forelimb buds, there is extensive expression of Tbx3 in posterior mesenchyme which masks transcripts in Pax3 positive myogenic progenitors. In the lower panels, X-gal staining of β-galactosidase from nlacZ (C', E') (Pax3nlacZ/+) or IRES-nlacZ (Pax3PAX3-FKHR-IresnlacZ/+ in D', Pax3Pax3-En-IresnlacZ/+ in F') reporters shows the extent of the somites, notably the hypaxial domain which undergoes cell death in Pax3 mutants (F').