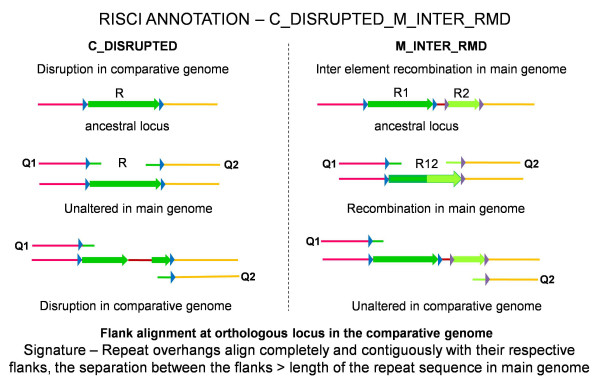
Figure 3.
Alignment signatures for M_INTER_RMD (Inter element recombination in main genome) or C_DISRUPTED (Disruption in comparative genome). In both cases, the repeat overhangs align completely and contiguously with their respective flanks and the separation between Q1 and Q2 is greater than the length of the transposon in the main genome. Left panel - Lone transposon in main genome (R1 - green arrow) disrupted by insertion of exogenous sequence (brown line) in the comparative genome. Q1 - upstream flank with 50 base repeat overhang. Q2 - downstream flank with 50 base repeat overhang. Right panel-From left to right - Pink line - 5' flank of R1, blue triangles - target site duplications of R1, green arrow - first repeat copy (R1), brown line - intervening sequence, grey triangles - target site duplications of R2, light green arrow - second repeat copy (R2 - same color indicating homology), orange line - 3' flank of R2. R12 - recombined repeat with a consequential loss of one copy of homologous region (R2) and the intervening sequence. Note that the 5' TSD of R12 comes from R1 (blue triangle) and the 3' TSD comes from R2 (grey triangle).