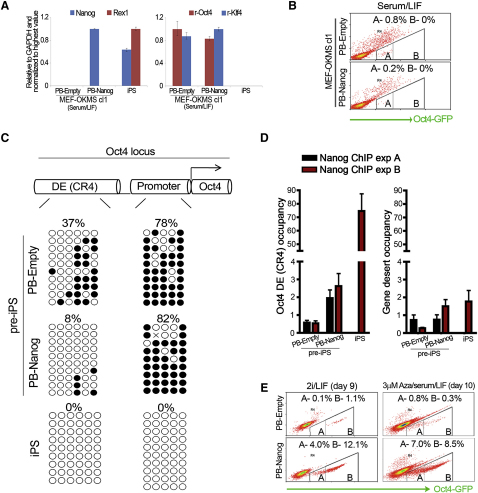
Figure 2.
Nanog Enhances Reprogramming in Synergy with 2i or Inhibition of DNA Methylation
(A) A piggyBac (PB) transgene was used to generate stable Nanog expressing cells in a clonal line of pre-iPS cells (MEF-OKMS clone 1). qRT-PCR analysis comparing expression of Nanog, Rex1, retroviral (r) Oct4 and r-Klf4 in PB-Nanog and PB-Empty pre-iPS cells expanded in serum/LIF. Error bars indicate the range of fold change relative to the sample with highest expression.
(B) Flow cytometry analysis indicates the proportion of cells with Oct4-GFP reporter activity in both transgenic backgrounds in serum/LIF.
(C) Bisulfite sequencing analysis of DNA methylation in the Oct4 distal enhancer and promoter in PB-Empty and PB-Nanog pre-iPS cells and 2i-iPS cells. The percentage of methylated CpG sites is indicated above each methylation panel.
(D) Chromatin immunoprecipitation analysis with a native Nanog antibody to the CR4 element in the Oct4 distal enhancer and a gene desert control region. Results of two independent experiments are shown. iPS cell occupancy was not measured in experiment A. Occupancy is plotted as fold enrichment over IgG after normalization to the input, and error bars represent standard deviation of the technical replicates of the qPCR for each experiment.
(E) Flow cytometry analysis comparing Oct4-GFP reporter activity after treatment of PB-Nanog and PB-Empty pre-iPS cells with 2i/LIF in serum-free medium or AZA (3 μM) in presence of serum/LIF. See also Figure S2.