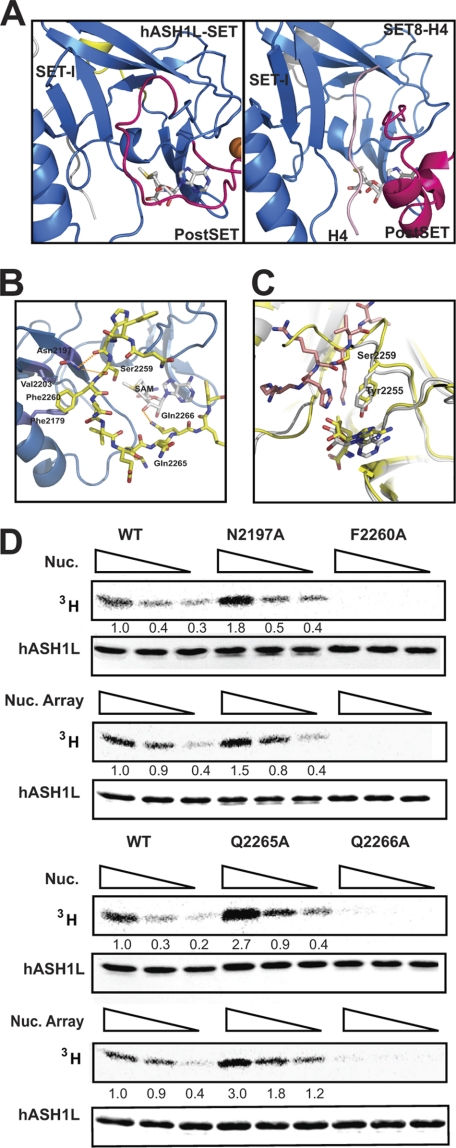
FIGURE 2.
The auto-inhibitory loop of hASH1L. A, comparison of the hASH1L-SET substrate binding pocket with that of the SET8-histone H4 complex. The auto-inhibitory loop in hASH1L occupies the substrate binding pocket where the histone substrate (colored in pink) binds in the SET8-histone complex structure. B, the auto-inhibitory loop interacts with the SET-I subdomain in the hASH1L SET domain. The carbonyl oxygens of His-2258 and Ser-2259 make hydrogen bonds with Asn-2197. Phe-2260 in the loop interacts with two hydrophobic residues, Val-2203 and Phe-2179. The auto-inhibitory loop also forms a hydrogen bond with AdoMet via Gln-2266. C, the hASH1L SET domain (shown in yellow) is superimposed on the SET8-histone H4 complex structure (SET8 shown in gray and H4 shown in pink). Ser-2259 of hASH1L occupies the substrate Lys position. D, disrupting the interactions between the auto-inhibitory loop and the SET-I subdomain or AdoMet affects the activity of hASH1L. Mutating Asn-2197 or Gln-2265 to alanine (N2197A, Q2265A) increases HMTase activity, whereas F2260A or Q2266A mutations abolish the activity. The HMTase activity was performed with tritium-labeled AdoMet with mononucleosome (Nuc.) or G5E4 nucleosomal array (Nuc. Array) as substrates.