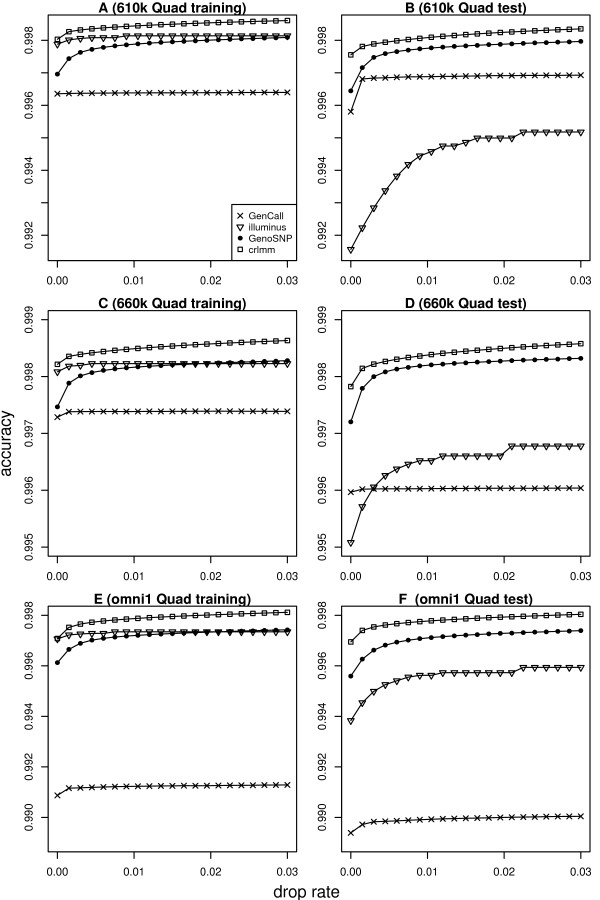
Figure 1.
Accuracy versus drop rate plots for the four methods tested. Figures on the left-hand side show results for the training data sets from 610 k Quad (A), 660 k Quad (C) and omni1 Quad (E) BeadChips. Figures on the right-hand side show results for the test data sets from the 610 k Quad (B), 660 k Quad (D) and omni1 Quad (F) BeadChips. Results are shown for autosomal SNPs only. CRLMM gives slightly more correct calls than the other methods for these high density chip types. GenCall is almost always slightly worse than the other methods. GenoSNP performs very consistently between data sets, achieving accuracy slightly below CRLMM. The accuracy of Illuminus seems to improve as the number of samples available increases (accuracy starts off at around 0.992 in B with 27 samples, and increases to 0.995 in D with 47 samples and 0.998 in A and C where in excess of 200 samples are available).