Abstract
Adenylosuccinate lyase (ADSL, E. C. 4.3.2.2) carries out two non-sequential steps in de novo AMP synthesis, the conversion of succinylaminoimidazole carboxamide ribotide (SAICAR) to aminoimidazolecarboxamide ribotide (AICAR) and the conversion of succinyl AMP (AMPS) to AMP. In humans, mutations in ADSL lead to an inborn error of metabolism originally characterized by developmental delay, often with autistic features. There is no effective treatment for ADSL deficiency. Hypotheses regarding the pathogenesis include toxicity of high levels of SAICAR, AMPS, or their metabolites, deficiency of the de novo purine biosynthetic pathway, or lack of a completely functional purine cycle in muscle and brain. One important approach to understand ADSL deficiency is to develop cell culture models that allow investigation of the properties of ADSL mutants and the consequences of ADSL deficiency at the cellular level. We previously reported the isolation and initial characterization of mutants of Chinese hamster ovary (CHO-K1) cells (Ade I) that lack detectable ADSL activity, accumulate SAICAR and AMPS, and require adenine for growth. Here we report the cDNA sequences of ADSL from CHO-K1 and Ade I cells and describe a mutation resulting in an alanine to valine amino acid substitution at position 291 (A291V) in Ade I ADSL. This substitution lies in the “signature sequence” of ADSL, inactivates the enzyme, and validates Ade I as a cellular model of ADSL deficiency.
Keywords: adenylosuccinate lyase deficiency, Chinese hamster ovary cells, inborn errors, cell culture models
1. Introduction
The enzyme adenylosuccinate lyase (ADSL, E. C. 4.3.2.2) carries out two non-sequential steps in the de novo synthesis of AMP, namely the conversion of succinylaminoimidazole carboxamide ribotide (SAICAR) to aminoimidazolecarboxamide ribotide (AICAR) and the conversion of succinyl AMP (AMPS) to AMP. In humans, mutations in ADSL lead to an autosomal recessive inborn error of metabolism - ADSL deficiency (MIM# 103050) - originally characterized by accumulation of dephosphorylated substrates for ADSL, succinylaminoimidazole carboxamide riboside (SAICAr) and succinyl adenosine (S-ado) in body fluids of affected individuals, and developmental delay, often with autistic features [1]. Most often, the initial diagnosis is made by detection of SAICAr in urine using the Bratton-Marshall reaction for diazotizable amines in infants or children with developmental delay of unknown origin [2]. Since this assay can result in false positives, the diagnosis must be confirmed, usually by high performance liquid chromatography (HPLC) [1, 3, 4, 5] followed by genomic and/or cDNA sequencing and characterization of the recombinant mutant proteins [6]. So far, 68 patients with 49 mutations leading to ADSL deficiency have been found (http://udmp.lf1.cuni.cz/adsl) [7, 8, 9, 10, 11, 12]. A majority of the identified mutations represent missense mutations, although in some cases, promoter mutations or splice site mutations leading to a truncated protein have been reported [2, 3, 13, 14]. However, these were always present in compound heterozygotes with missense mutations in the second allele. In all the cases studied so far, the combination of the mutations produce ADSL enzymes that retain some residual enzyme activity [6, 9, 10, 15, 16].
As more cases of ADSL deficiency have been identified and characterized, the variability in phenotype has expanded considerably to include psychomotor delay, fatal neonatal encephalopathy with hypokinesia, severe mental retardation, dysmorphic features, behavioral abnormalities, stereotypic behaviors, autistic features, brain atrophy, hypotonia, seizures, abnormal brain glucose utilization, muscle wasting, and failure of muscle energy metabolism [7, 8, 9, 10].
ADSL deficiency can be grouped into two classes, type I and type II. Type I is a severe childhood form which presents within the first months of life, encompasses the whole spectrum of disease symptoms, and results in severe psychomotor retardation, developmental arrest, coma vigil and often early death. Type II is a moderate or mild form which presents within the first years of life either with psychomotor retardation or hypotonia. Autistic features are a common feature of ADSL deficiency. In a general sense, individuals in whom the S-ado/SAICAr ratio is less than two seem to be more severely affected, whereas if the ratio is greater than 2, the disorder tends to be less severe [15,17, 18,]. This classification is, however, not absolute, and ADSL deficiency presents with a broad spectrum of severity from relatively mild to severe.
A recent report demonstrates demyelination and brain atrophy in 7 of 7 individuals with ADSL deficiency [19]. In several cases the brain atrophy progressed with time. These investigators hypothesize that the observed brain atrophy is due to some aspect of ADSL deficiency and hypothesize that disorders of energy metabolism might contribute to the pathology. They further hypothesize that ADSL deficiency might affect multiple metabolic systems and that symptoms may reflect more than one of these [19]. This is a reasonable hypothesis since purines are essential components of a number of vital biomolecules such as RNA, DNA, essential coenzymes (NAD, NADH, FAD, and Coenzyme A for example) signaling molecules (cAMP, guanine nucleotides), and ATP. Neuronal cells, which have particularly high energy requirements, might be particularly sensitive to perturbations of purine metabolism.
Currently, there is no detailed knowledge about the pathogenesis of disease symptoms and hence no effective treatment for ADSL deficiency. Hypotheses regarding the pathogenesis include toxicity of high levels of SAICAR, AMPS, or their metabolites, inability to synthesize sufficient purines using the de novo purine biosynthetic pathway, or lack of a completely functional purine cycle in muscle and brain. Attempts have been made to correlate residual enzyme activity with severity of ADSL phenotype with some success [6, 16].
One important approach to understanding ADSL deficiency so that effective treatments can be devised is to develop cell culture models that allow investigation of the properties of ADSL mutants, and the consequences of ADSL deficiency, at the cellular level. Chinese hamster ovary (CHO-K1) cells, though not of neuronal origin, are among the most widely used mammalian cells for stable expression, production, and characterization of recombinant proteins. The produce accurately post-translationally modified proteins and allow various studies on recombinant protein biogenesis, intracellular trafficking, and cellular localization. ADSL is a ubiquitously expressed cytosolic protein. So far, there are no indications of cell-specific modification or intracellular localization of ADSL. In addition, primary neuronal cultures do not require exogenously added purines, implying that they possess ADSL activity. These observations provide an important rationale for the conclusion that many aspects of de novo purine synthesis would be similar in CHO-K1 cells and neuronal cells.
We previously reported the isolation of CHO-K1 mutants designated Adel that require adenine for growth, accumulate SAICAR and AMPS, and have little or no ADSL enzyme activity [20, 21]. We used these cells to map the gene for ADSL deficiency to human chromosome 22 [22]. Ade I cells provide a cell culture model for investigating the molecular, biochemical, and biophysical characteristics of ADSL, the cellular effects of various ADSL mutations associated with ADSL deficiency, and possible mechanisms for treating ADSL deficiency. Here we report the identification and characterization of the ADSL deficiency in the Ade I mutant. The deficiency is the result of the combination of a non-transcribed or deleted ADSL allele and an allele with a missense mutation (A291V) located in the “signature sequence” of ADSL [23]. This sequence has been hypothesized to participate in substrate binding, correct orientation of a catalytic amino acid, or providing a favorable environment for catalysis after substrate binding [23, 24].
2. Materials and Methods
2.1. Sequencing of ADSL from CHO-K1 and Ade I
Both 3′- and 5′-RACE [25, 26, 27] and PCR were used to determine the ADSL sequence from CHO-K1 and Ade I. The majority of the ADSL sequence was obtained using 3′-RACE. For cDNA sequencing, cells were homogenized using Qiashredder columns (Qiagen) and RNA was isolated using the RNeasy Mini Kit (Qiagen) following the manufacturer’s protocol. Initial ADSL cDNA sequence was obtained using a Roche 1st Strand cDNA Synthesis Kit (AMV). For some experiments reverse transcription was done using ThermoScript RT-PCR (Invitrogen).
Initially, the Anchor dT17 primer was used in conjunction with ADSL specific primers. The ADSL specific primers were R1, R2, R3 or R9 (see Supplementary Table 1 for all RACE primer sequences and Table 1 for all other primers). Primers F1, F2, R1 and R2 were designed using sequences highly homologous to mouse and human ADSL that were shown to recognize CHO ADSL sequence. Primer R9 was designed based on known CHO sequence determined as part of this study. Both conventional and touchdown PCR were used to determine the sequence of the ADSL gene from Ade I and CHO-K1 cDNA. Our default program for conventional PCR is: 95°C for 5 minutes, (95°C for 30 seconds, 55°C for 30 seconds, and 72°C for 30 seconds) for 35 cycles, and 72°C for 7 minut es. PCR products were treated with ExoSapIT (USB) to eliminate carryover of primers and unincorporated nucleotides prior to sequencing.
Table 1.
Primer Sequences
| Ala_Val_hADSL_F | 5′ CAGCAGATTGGCTCAAGTGTAATGCCATATAAGCGGAATC 3′ | SDM |
| Ala_Val_hADSL_R | 5′ GATTCCGCTTATATGGCATTACACTTGAGCCAATCTGCTG 3′ | SDM |
| T7 | 5′ TAATACGACTCACTATAGGG 3′ | Sequencing |
| BGH_rev | 5′ TAGAAGGCACAGTCGAGG 3′ | Sequencing |
| hADSL_1262R | 5′-TTTTCTCATGGCAATCCTGGCGGC-3′ | Sequencing |
| hADSL_1160F | 5′-ATTGAACGGCGCATTCGGCAAGAG-3′ | Sequencing |
| 4F_hADSL | 5′-CGTGTACCCCAAAGTAATTGAACG-3′ | RT-PCR; qPCR |
| 4R_hADSL | 5′-AATCTCTGCACCTGCTGGGA-3′ | RT-PCR; qPCR |
| CHObactin-F | 5′-GCTCTTTTCCAGCCTTCCTT-3′ | RT-PCR; qPCR |
| CHObactin-R | 5′-TGCTTGCTGATCCACATCTC-3′ | RT-PCR; qPCR |
After determining the distal portion of the CHO-K1 and Ade I ADSL genes using 3′ RACE, approximately the first 200 nucleotides of the ADSL sequence from CHO-K1 and Ade I, including a putative ATG start codon, were determined using the 5′-RACE System for Rapid Amplification of cDNA Ends, Version 2.0 (Invitrogen) according to the manufacturer's protocol. ADSL-specific primers for 5′-RACE were designed using the CHO- K1 ADSL sequence previously determined using 3′-RACE. Specificity for ADSL was monitored by electrophoresis of PCR products on 1% agarose gels. Sequencing of the final 5′-RACE PCR product was done using primer 298_ADSL_K1R by University of Colorado Cancer Center DNA Sequencing & Analysis Core.
After CHO-K1 and Ade I cDNAs were sequenced, genomic DNA was isolated and genomic regions spanning the detected sequence changes were sequenced in order to verify the presence of the mutation within the ADSL gene. Genomic DNA was isolated using the DNA Purification Kit for Cultured Cells (Puregene) using the manufacturer’s protocol. PCR conditions were the same for both K1 and Ade I DNA. PCR products were treated with ExoSapIT and sequenced using the same PCR primers that were used to amplify the ADSL gene in 3′-RACE protocols described above.
2.2. Introduction of mutations into human ADSL cDNA in pCR3.1
Plasmid hADSL-pCR3.1 [6] is a mammalian cell expression vector containing the wild-type human ADSL (hADSL) cDNA sequence. The Quick Change II Site-directed Mutagenesis kit (Stratagene) was used to introduce the A291V (gcg to gta) mutation (present in the Ade I ADSL gene), into hADSL-pCR3.1 following the manufacturer's protocol. (The g to a change in the third position was made so that the mutated codon accurately reflects the CHO sequence for that codon. It does not change the amino acid encoded). The Ala_Val_hADSL_F and Ala_Val_hADSL_R primers (Table 1) were designed using the Quick Change® Primer Design Program (www.stratagene.com). The resulting mutant plasmid was designated pLKV1. The presence of the A291V mutation was confirmed by DNA sequencing followed by alignment with the wild-type hADSL cDNA, NCBI accession number AF067853 [6] using Align2 BLAST [28]. The primers used to sequence hADSL were T7, BGH_rev, hADSL_1262R, and hADSL_1160F (Eurofins MWG Operon). Maxi-preps of pLKV1 and hADSL-pCR3.1 plasmid DNA were done using AX500 anion exchange columns and the Nucleobond Plasmid Purification Kit (Clontech). The purity and quantity of plasmid DNA was evaluated by UV absorbance measurements at λ 260 nm and 280 nm.
2.3. Transfection of cDNA clones into Ade I cells
Transfection of hADSL-pCR3.1 and pLKV1 into Ade I using Lipofectamine 2000 (Invitrogen) was done according to the manufacturer’s instructions, and as previously described [29]. The initial selection was done in F12FC10 plus 3×10−5M adenine plus Geneticin (400 µg/ml). Several independent Geneticin resistant clones were picked and expanded in this medium. After several passages, the clones were tested for growth in adenine free medium.
2.4. Assay for expression of hADSL cDNA in transfected clones
Total RNA was isolated from transfected Ade I clones, and untransfected K1 and Ade I cells using the standard TRI Reagent (Sigma-Aldrich) procedure; cDNA was reverse transcribed from total RNA using the High Capacity RNA-to-cDNA kit (Invitrogen) according to the manufacturer's protocols. The resulting cDNA was used as a template in 20 µl PCR reactions containing 50 mU of platinum taq DNA polymerase, 1.5 mM MgCl2, 200 µM dNTPs, 1 µM hADSL-specific primers (4F_hADSL and 4R_hADSL) under the following PCR conditions: initial denaturation at 95°C for 5 min followed by 30 cycle s of denaturation at 95°C for 30 s, primer annealing at 57°C for 30 s, and extension at 72°C f or 30 s. A final chain elongation was performed at 72°C for 7 min. The housekeeping gene, β-Actin, was amplified as a control using specific primers (CHObactin-F and CHObactin-R) following the conditions above. The PCR products were analyzed by gel electrophoresis on 2% agarose gels in 0.5 X TBE to confirm ADSL expression in transfected Ade I cells. As a positive control, cDNA was prepared on total RNA from HeLa cells (Clontech) and the human ADSL gene was amplified by RTPCR, as described above.
2.5. Expression levels of ADSL transcripts in CHO cells
ADSL expression levels in CHO cells (Ade I and K1) were determined by quantitative PCR (qPCR) and ΔΔCt analysis. Total RNA isolation and subsequent cDNA synthesis was carried out as described above. The cDNA was used as a template in 25µl reactions following QuantiFast SYBER Green PCR kit (Qiagen) with 1µM primers (Mm_Adsl_1_SG from Qiagen QuantiTech Primer Assay). Mouse primers were used to determine the expression of CHO ADSL transcripts because CHO specific primers were not available. The mouse primers were validated and it was determined that they were a viable alternative. The qPCR cycling conditions were as follows: 95°C for 5 min followe d by 35 cycles of denaturation at 95°C for 10 s, combined annealing/extension at 60°C for 30 s. Ribosomal (18s) RNA was amplified as a control using specific primers (Mm_Rn18s_2_SG from Qiagen QuantiTech Primer Assay) following the conditions above.
2.6. Western blot analysis
Cells were grown in Ham’s F12 supplemented with 10% fetal calf serum, 3×10−5 M adenine, and Normocin (100 µg/ml) (InvivoGen). Cells were washed with PBS, then harvested in cell lysis buffer (CLB) + protease inhibitors (PI) (Complete Mini, Roche Applied Science). Samples were centrifuged at 55,000 rpm for 10 minutes at ~4 to 15° C using a Beckman SW60 rotor. Supernatant was collected, frozen on dry ice, and stored at −70 C. Samples for SDS-PAGE were prepared by combining 2 volumes protein with one volume 3× Laemmli buffer/300 mM DTT. Prior to loading, samples were incubated for 5 minutes at ~90 C. Samples were run on Novex 8% tris/glycine gels (Invitrogen) following the manufacturer's protocol. Gels were “wet” blotted to nitrocellulose (Hybond, Amersham Biosciences) in 25 mM tris/190 mM glycine/20% methanol at 200 mA constant current, for one hour. Blots were allowed to dry overnight and transfer quality was evaluated by staining with 1× Ponceau S. After Ponceau S staining, blots were immersed in 1 × tris buffered saline (TBS) to destain, then were blocked with 1× TBS/5% BLOTTO. This was followed by 90 minute incubation in primary antibody (diluted in 1× TBS/1% BLOTTO/0.02% Triton X-100), three five minute washes, 60 minute incubation in secondary antibody (1:10,000 dilution, Goat Anti-Rabbit IgG (H+L)-AP Conjugate, Bio-Rad), two five minute washes, then two five minute washes in 1 × TBS. The primary antibodies are anti-ADSL (Sigma, 1:2500 dilution), and anti-β-actin (Cell Signalling, 1:1000 dilution). Blots were treated with Tropix CDP-Star Ready to Use with Nitro-Block II (Applied Biosystems), and visualized using a Kodak Image Station 4000R (Carestream Molecular Imaging). We used recombinant ADSL expressed from pMAL-c2 grown in DH5αF’IQ as previously described [6].
2.7. ADSL Enzyme Activity in Transfected CHO Ade I cells
ADSL activity assays were carried out as follows. Cells were harvested from three nearly confluent 10 cm plates of each cell line (CHO-K1, Ade I, and the wt-hADSL-pCR3.1 1.1a, wt-hADSL-pCR3.1 2.1c, A291V-pLKV1 1.1b, and A291V-pLKV1 2.1d transfectants). For collection, plates were rinsed with ice cold Saline D, then about 2 ml of 10 mM Tris (pH 7.4)/10 mM EDTA/300 mM sucrose was added to each plate. The cells were detached from the plate using a cell scraper, and then pelleted by centrifugation. Pellets were resuspended in the same buffer and sonicated in an ice bath using a Branson Sonifier 185. Cell lysates were centrifuged at 20,000 rpm (47,808 g) for 60 minutes at 4° C and su pernatants were assayed for total protein content using the Bio-Rad Protein Assay. ADSL enzyme activity was assayed in triplicate using 90 µg total protein from each tissue lysate in 40 mM Tris buffer (pH 7.4) with 90 µM AMPS (Sigma-Aldrich). A decrease in absorbance at 282 nm was measured over 20 minutes, as AMPS was converted to AMP. The assay buffer was incubated at 37°C just prior to addition of lysate as it increased the efficiency of the reaction. Specific activities (nmol of substrate converted/min/mg of total protein) of the ADSL enzyme in each cell lysate were calculated using the difference extinction coefficient of 10,000 M-1 cm-1 [30]. All assays were linear with time and amount of protein.
2.8. In-silico analysis of ADSL 3-D structure
Structural topology of the mutation was determined on protein database (PDB) deposited crystal structure of human ADSL with its substrate S-AMP and products AMP and fumarate (2VD6; available at http://www.pdb.org/pdb/cgi/explore.cgi?pdbId=2VD6). The region adjacent to the mutated residue was highlighted and visualized using PyMOL Viewer (http://www.pymol.org).
3. Results
3.1. The sequence of CHO wild-type and mutant ADSL cDNA
We amplified the wild-type CHO-K1 and Ade I ADSL cDNAs using 5’ and 3’ RACE and PCR and sequenced the products as described in Materials and Methods. The cDNA sequences have been submitted to GenBank (NCBI accession numbers GQ984234 and GQ984235, respectively).
The analysis of the CHO-K1 sequence revealed an A-G heterozygosity at nucleotide position 936 (with the A of the start codon = 1). The c.936A and c.936G genotypes encode isoleucine (p.Ile312) and methionine (p.Met312) residues in the CHO-K1 ADSL proteins. The CHO-K1 ADSL has 93% identity and 96% similarity to human ADSL (hADSL) and 95% identity and 97% similarity to mouse ADSL (mADSL). Both hADSL and mADSL have a methionine at position p.312. Isoleucine is a conservative substitution for methionine and ADSL from many organisms has an isoleucine residue at this position.
When compared to CHO-K1, the cDNA sequence from Ade I revealed a homozygous genotype c936G/c.936G encoding the p.Met312 variant, and a homozygous transition c.872C>T resulting in the non conservative amino acid exchange p.Ala291Val. Genomic sequencing confirmed the presence of the heterozygous genotype c.936A/c.936G and homozygosity for c.872C in CHO-K1. The Ade I genomic sequence confirmed the presence of the homozygous genotype c936G/c.936G but contrary to cDNA sequence it revealed a heterozygosity T-C at nucleotide c.872. These results mean that only the mutant allele c.872T encoding p.Val291 is expressed in AdeI. The second allele is either not transcribed or its transcript is lost in AdeI cells as a consequence of nonsense mediated decay resulting from partial deletion or splicing mutation.
3.2. The A291V mutation found in Ade I results in ADSL inactivation
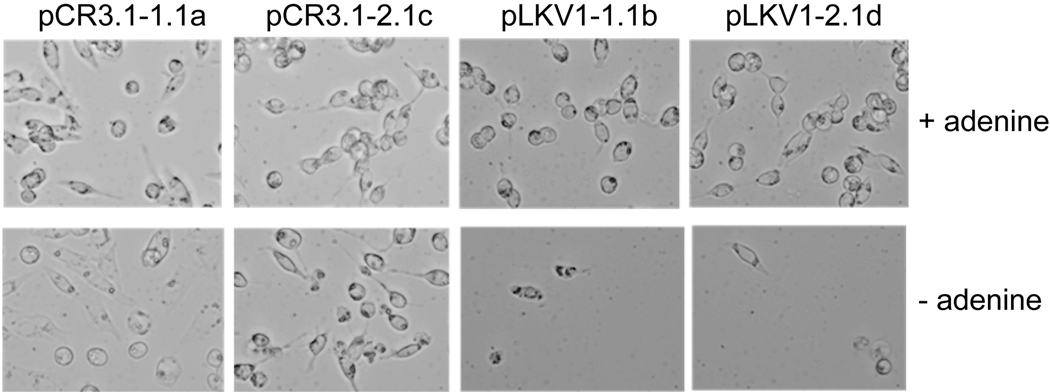
To assess the functionality of the mutant ADSL allele found in Ade I, we engineered the c.872C>T mutation into the hADSL expression clone, hADSL-pCR3.1 [6], which also contains the gene for Geneticin resistance. The resulting constructs, pLKV1, and hADSL-pCR3.1, were used separately to transfect Ade I cells and clones were selected for growth in the presence of Geneticin plus adenine. Transfection frequency (assessed by Geneticin resistance) was essentially equal for the two clones. We picked several independent clones from each transfection, grew them in the presence of Geneticin plus adenine, and checked each independent clone for growth in the absence of adenine. Clones transfected with hADSL-pCR3.1 were able to grow in the absence of adenine. Clones transfected with pLKV1 were unable to grow in the absence of adenine. Two independent clones of hADSL-pCR3.1 (hADSL-pCR3.1-1.1a and hADSL-pCR3.1-2.1c) and two of pLKV1 (pLKV1-1.1b and pLKV1-2.1d) were selected for further analysis. The growth response of these clones in medium plus and minus adenine is shown in Figure 1.
Figure 1. Complementation analysis of Ade I by wt- and A291V-hADSL plasmids.
Ade I cells were transfected with wt (hADSL-pCR3.1) or A291V (pLKV1) hADSL plasmids. After 24–48 hr, cells were split into fresh media containing Geneticin. Cells were split every 48–72 hr into media containing Geneticin for several days. Geneticin resistant clones were isolated and grown in the presence (+) or absence (−) of adenine. Cells were fixed and stained in 1% crystal violet (w/v) in 50:50 (v/v) dH2O: methanol.
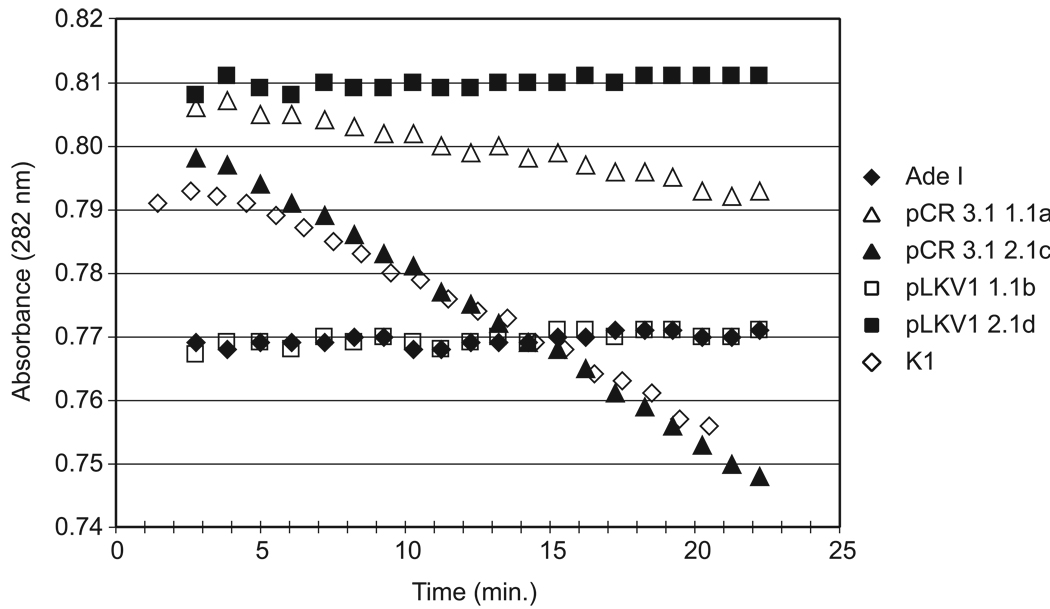
We measured ADSL enzyme activity in the transfectants and found that indeed the hADSL-pCR3.1 transfectants possessed significant ADSL enzyme activity, while the pLKV1 transfectants did not (Figure 2). We conclude from these experiments that hADSL can substitute for CHO-K1 ADSL in the de novo biosynthetic pathway. The A291V mutation found in AdeI, when introduced into hADSL, results in no detectable ADSL enzyme activity, and no substitution for native ADSL activity.
Figure 2. Enzymatic Assay of CHO Cell Lysates for ADSL Activity.
CHO cell lysates from untransfected K1 and Ade I (open and closed diamonds, respectively) and Ade I cells transfected with wild-type hADSL (hADSL-pCR3.1 clones 1.1a and 2.1c, open and closed triangles, respectively) or A291V hADSL (pLKV1 clones 1.1b and 2.1d, open and closed squares, respectively) were assayed for ADSL activity. Data represent one of three independent assays.
3.3. Levels of ADSL transcripts in Ade I
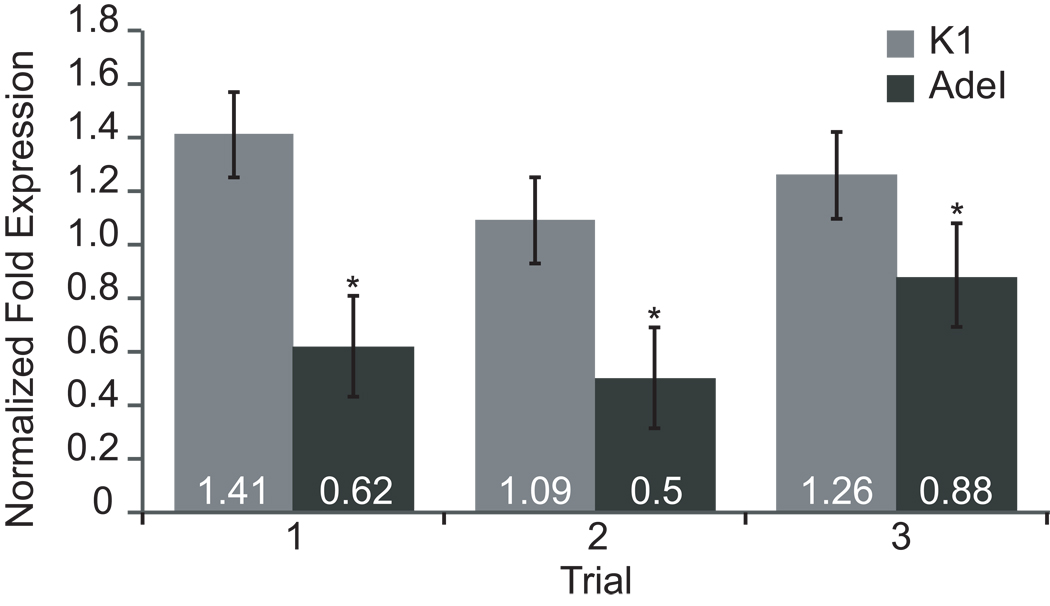
The lack of ADSL enzyme activity could be due to the production of an inactive form of ADSL, lack of production or instability of mutant ADSL mRNA, or production of an inactive ADSL protein. We quantified relative levels of ADSL mRNA in CHO-K1 and AdeI cells using qPCR. Figure 3 shows triplicate qPCR analyses of ADSL mRNA from CHO-K1 and AdeI cells. AdeI produces 53% as much ADSL mRNA as CHO-K1. This result is consistent with the possibility that one allele of ADSL is silenced in AdeI cells and that the other allele produces mutant ADSL mRNA.
Figure 3. qPCR analysis of ADSL mRNA levels in CHO-K1 and AdeI cells.
Total RNA was isolated and cDNA prepared from CHO cells (K1 and AdeI). Normalized fold expression levels of ADSL transcripts in K1 and AdeI was determined by qPCR, in triplicate, using ADSL gene specific primers and 18s ribosomal RNA reference gene primers. Relative expression levels were determined using the ΔΔCt method.
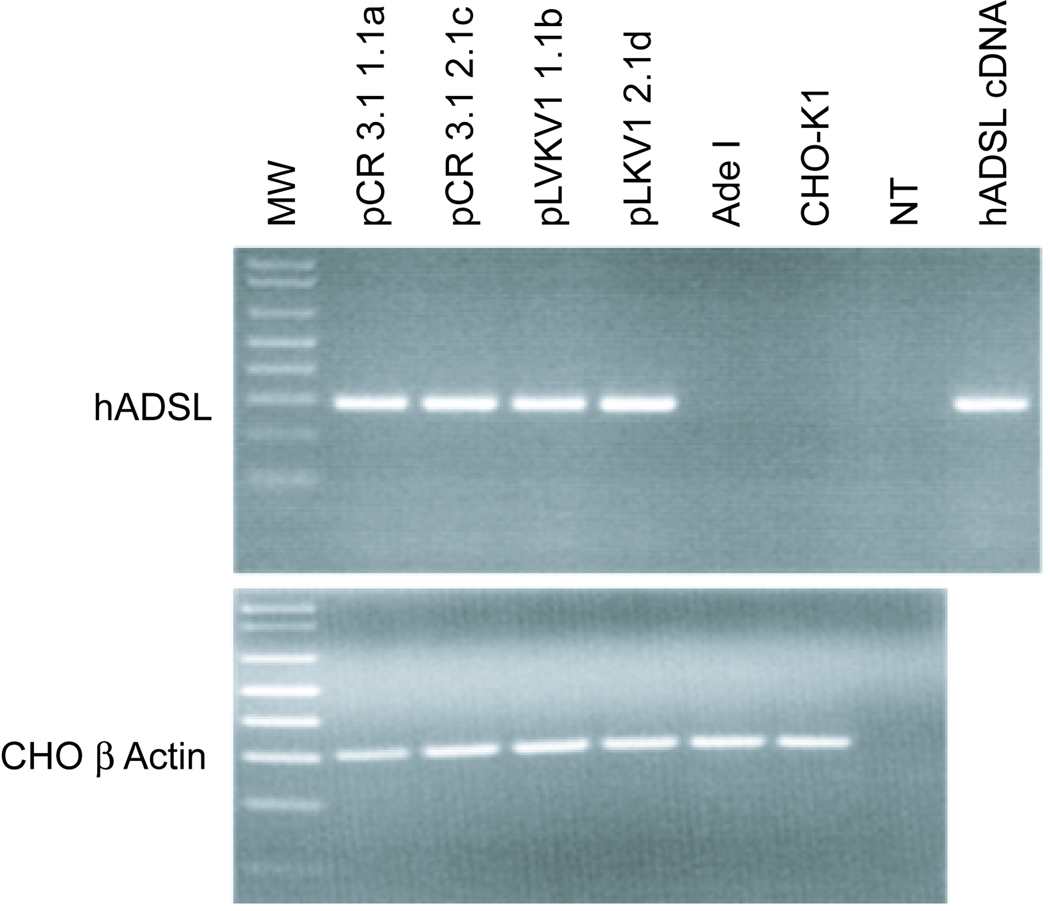
Agarose gel electrophoresis of the RT-PCR products confirms the presence of hADSL mRNA in transfected Ade I cells but not in the untransfected CHO-K1 and Ade I controls (Figure 4). Transcripts of the CHO β-actin gene are present in both transfected and untransfected controls. RT-PCR of hADSL from cDNA prepared from total RNA of HeLa cells (Clontech) results in a band of the same size as that in the transfected Ade I cells. The similar intensity of the bands indicates the hADSL gene is transcribed at approximately the same level in Ade I cells transfected with either wt (hADSL-pCR3.1-1.1a and -2.1c) or mutant (pLKV1-1.1b and -2.1d) hADSL clones. Any minor differences in hADSL transcript levels cannot account for the large difference observed in enzyme activity (Figure 2) between Ade I cells transfected with wt-hADSL versus A291V-hADSL plasmids.
Figure 4. Expression of hADSL mRNA in transfected Ade I cells.
(Upper panel) Total RNA was isolated and cDNA prepared from Ade I transfected with either wt (hADSL-pCR3.1) or A291V (pLKV1) hADSL plasmids. PCR was performed on cDNA from two independently-isolated clones for wt (1.1a and 2.1c) and A291V (1.1b and2.1d). Untransfected Ade I and CHO cDNA were used as negative controls. A no template control (NT) was also subjected to PCR. cDNA prepared from HeLa total RNA was used as a positive control. (Lower panel) The CHO β-actin gene was amplified by PCR to confirm RNA integrity.
3.4. ADSL protein expression in CHO-K1, AdeI, and transfectants hADSL-pCR3.1 2c.1 and pLKV1 1.1b
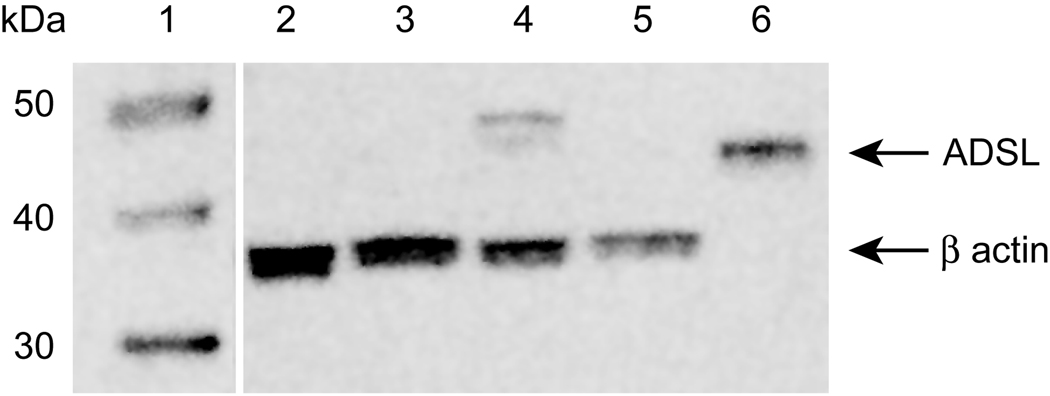
We examined CHO-K1, AdeI cells, and AdeI cells transfected with either wt or mutant hADSL for expression of ADSL protein by Western blotting using a commercially available anti-ADSL antibody (Sigma). This antibody does not recognize CHO ADSL under the conditions employed here. The results indicate that the hADSL-pCR3.1 transfected AdeI cells produce hADSL protein (Figure 5). However, no hADSL protein can be detected in AdeI cells transfected with pLKV1 1.1b. We consistently observe a slightly increased apparent molecular weight for hADSL expressed in AdeI cells compared to human recombinant ADSL protein. This may suggest some modification of hADSL in AdeI cells.
Figure 5. Western blot of AdeI transfectants.
Fifteen µg of protein from CHO cell lines was analyzed by western blotting using anti-ADSL and anti-B-actin polyclonal antibodies (see Materials and Methods). Each sample lane was loaded with with 15 ug protein. The lanes are: 1) MW (MagicMark XP, Invitrogen), 2) CHO-K1, 3) AdeI, 4) PCR 3.1 2.1c, 5) pLKV1 1.1b, and 6) Recombinant ADSL (Kmoch Lab).
4. Discussion
4.1. The identity of the amino acid substitution in Ade I
In this manuscript we demonstrate that the Ade I mutant of CHO-K1 cells contains a mutation in the ADSL gene causing an A291V amino acid substitution. We further show that this amino acid substitution inactivates hADSL. This is a highly conserved residue in ADSL, with rare substitutions to serine (S) (conservative) and threonine (T) (non-conservative) in a few organisms.
Interestingly, a polymorphism was found in the CHO-K1 cDNA and genomic sequence at position 936. This substitution results in either I or M at amino acid position 312. This is a common substitution in ADSL in many organisms and is a conservative amino acid substitution. The M at this site can be substituted by at least 13 other amino acids, so it seems unlikely that the M to I substitution at this site would have functional consequences for ADSL. This polymorphism was not seen in cDNA or genomic DNA from Ade I. This result suggests that the ADSL locus is present in two copies in CHO-K1, which makes both forms of ADSL. Ade I also appears to have two genomic copies of at least part of the ADSL gene. In this case, however, there is no polymorphism at position 312, but there is at position 291. Since there was no evidence for expression of wt ADSL at the mRNA level in Ade I, we hypothesize that one ADSL allele is inactive. Similar findings have been observed for some CHO-K1 AdeC mutants in the GART gene [31].
4.2. Molecular consequences of the AdeI mutation
By qPCR analysis, AdeI makes about 50% as much ADSL mRNA as CHO-K1 (Figure 3). This would be consistent with the interpretation that one ADSL allele is silent in AdeI and that the other encodes mutant ADSL protein. Currently, we do not have a satisfactory antibody that recognizes CHO ADSL protein. However, Western blotting of AdeI cells transfected with either wt or mutant hADSL demonstrate that wt hADSL is expressed in these cells, a finding consistent with direct enzyme assays and RT-PCR with human specific primers (Figures 2, 4, and 5). No hADSL protein was detectable in AdeI cells transfected with the A291V mutant form of hADSL even though hADSL mRNA is produced in these cells (Figures 4 and 5). Thus, either the A291V mutation makes hADSL protein unstable or the mutation reduces or eliminates translation of ADSL mRNA. We speculate that CHO mutant ADSL protein may be affected similarly. If so, this suggests that AdeI cells may not have a significant amount of ADSL protein. This point awaits development of antibodies that successfully recognize CHO ADSL. The slight difference in apparent molecular weight in hADSL expressed in AdeI cells compared to recombinant hADSL protein is consistently observed and suggests that hADSL may be post-translationally modified in AdeI cells.
4.3. Insights about ADSL function from the A291V mutation in Ade I
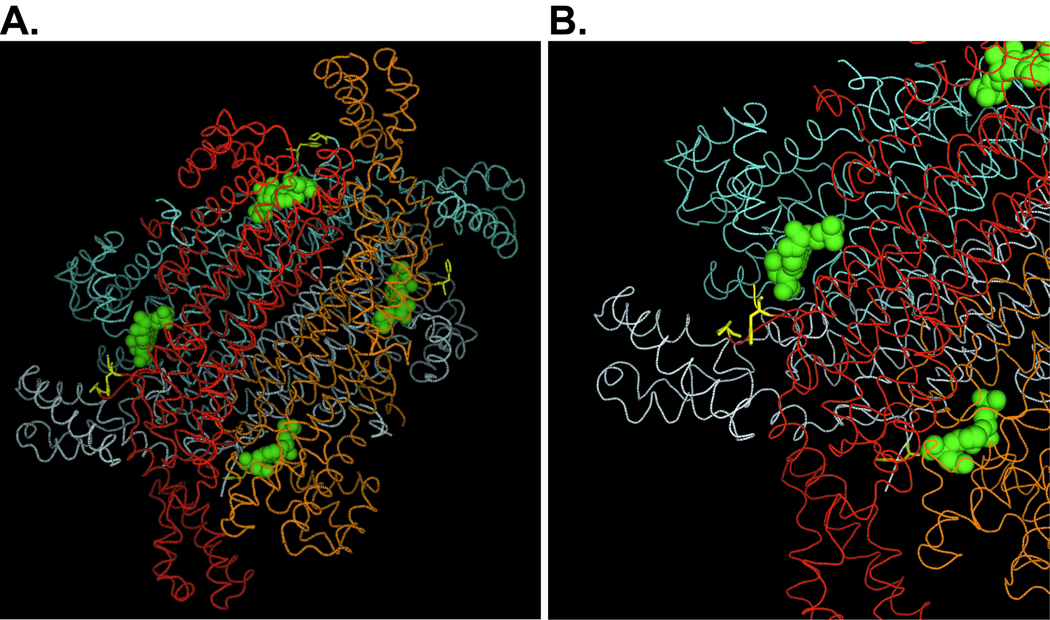
The alanine residue mutated in Ade I lies within the highly conserved signature sequence of ADSL. The signature sequence is thought to be part of a flexible loop encompassing residues 282 to 293 that folds over the active site of ADSL after substrate binding thus improving the environment for catalysis [23] (Figure 6). Sivendran and Colman [23] analyzed the loop region of B. subtilis and human ADSL and demonstrated that serine 289 is essential for enzyme activity while serine 290 is essential for maximal activity. Both B. subtilis and human ADSL have an A at position 291 (human numbering system). It may be that substitution of alanine with the more bulky valine at position 291 inhibits the mobility of the flexible loop, restricts the ability of the substrate to fit into the active site, or inhibits the contact of the active site serines with the substrate. Any of these alterations may also result in unstable ADSL protein.
Figure 6. Structural model of human ADSL.
(A) Four ADSL subunits forming native homotetramer are shown in different colors. All four of the enzyme active sites are highlighted by presence of one of the ADSL substrates, S-AMP (green spheres). A291V residues and adjacent flexible loops lining the substrate channels of the enzyme are present between Phe282 and Pro293 residues shown in yellow. (B) Detail of one of the active sites.
Recently, Bulusu et al [24] analyzed ADSL from the intracellular parasite Plasmodium falciparum, which causes malaria in humans. These investigators argue that the serine at position 298 in that organism (equivalent to 289 in hADSL) may indeed function in catalysis. P. falciparum and other organisms, including E. coli, have a threonine (T) at the amino acid position homologous to A291 in the mammalian enzyme. T has a bulkier side chain than A but not as large as V. Therefore, we hypothesize that T would be more restrictive than A but not as restrictive as V on the positioning of the serines required for optimal enzyme activity. This may help explain some of the differences in enzyme function observed between organisms with alanine or threonine at this position in ADSL.
4.4. The use of AdeI cells as a cell culture model of ADSL deficiency
Several features of the AdeI mutant described here make it potentially an excellent model for examining the in vivo characteristics of mutant forms of hADSL. First, and most important, AdeI cells have been shown to possess a missense mutation in ADSL rendering the enzyme inactive or unstable. While this was expected from previous work, it remained possible that the mutation in AdeI inactivated the normal ADSL gene by a mechanism other than a coding region mutation. Human ADSL is expressed and active when transfected into AdeI cells. Human-specific ADSL PCR primers and antibodies have been developed so that transcription of hADSL and the consequences of mutations in ADSL can be studied in transfected cells. This in vivo system is important because many features of ADSL synthesis and processing cannot be studied well in cell free extracts. For example, it would be difficult to characterize post-translational processing of ADSL in cell free extracts. Moreover, the intracellular (in vivo) environment is markedly different from the in vitro environment used for most analyses of ADSL thus far [32, 33]. The intracellular environment may have a total protein and RNA content approaching 300 to 400 gm/L and macromolecules may occupy over 30% of the cellular volume. This is a markedly different milieu, in which protein folding and aggregation takes place, than that found in most in vitro studies.
Methods are becoming available for studying these phenomena in whole cells [34, 35, 36]. Thus, Ade I cells can be used to assess the activities, stabilities, aggregation, multimer formation, and other structural consequences of ADSL mutants associated with the ADSL deficiency. As discussed above, although CHO cells are not derived from neuronal cells, it is highly likely that de many of these aspects of de novo purine synthesis are similar in all cell types. Similar approaches have been used to explore pathogenic mechanisms in other human genetic disorders, notably mutations in cytosolic superoxide dismutase (SOD1) [37, 38]. Further, AdeI cells can be used to assess the consequences of mutations introduced into the ADSL gene to test particular hypotheses regarding the function of specific amino acid substitutions. An interesting potential example would be to assess complementation of different ADSL mutations observed in cell free studies using purified ADSL [16].
Ade I cells accumulate SAICAR and AMPS [21]. One hypothesis of ADSL deficiency is that the accumulation of SAICAR or AMPS and their metabolites is toxic. This hypothesis can be tested on the cellular level by altering the level of accumulation of SAICAR or AMPS under conditions of no or limited exogenous adenine concentrations and monitoring cell survival and determining the mechanism of cell death.
Recently, the existence of a multienzyme complex containing all the enzymes of de novo purine synthesis, the purinosome, which includes ADSL, has been described in HeLa cells and human carcinoma liver cells, although it has not yet been described in neuronal cells [39, 40]. Such a complex has been hypothesized for decades, with little evidence. The purinosome is a transient structure existing only when the cells need to synthesize purines. The existence of the purinosome leads to an additional hypothesis for the ADSL deficiency phenotypes observed in humans. It is possible that some ADSL mutants may disrupt the formation of the purinosome at crucial times in development or in the cell cycle and that this may play a role in the pathogenesis of human ADSL deficiency. Some mutant ADSL enzymes may have a more profound effect on purinosome formation and function than others. This may explain why the severity of ADSL deficiency does not always correlate with residual enzyme activity as measured in cell free extracts or with purified enzyme. This hypothesis could be tested using Ade I cells.
ADSL deficiency commonly includes autistic features and brain atrophy. Thus, it may be possible that understanding this inborn error of metabolism will not only reveal pathogenic mechanisms for the syndrome itself but also some mechanisms relevant to development of autistic features and some aspects of aberrant brain development and neurodegeneration as well. In this regard, the creation of mouse models with altered ADSL would be a logical next step.
Supplementary Material
Acknowledgments
This work was supported by NIH grant MH 065431 and by grants from the Bonfils-Stanton Foundation, the Ludlow-Griffith Foundation, and the McCulloch McTavish Foundation. Support to MZ and SK was provided by grant 301/07/0600 from the Czech Science Foundation and institutional grants 1M6837805002 and MSM0021620806 provided by Ministry of Education of Czech Republic. The University of Colorado Cancer Center DNA Sequencing & Analysis Core is supported by the NIH-NCI grant P30 CA046934. The authors also wish to thank Miles Brennan for help designing PCR primers for 3’- RACE.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Jaeken J, Van den Berghe G. An infantile autistic syndrome characterised by the presence of succinylpurines in body fluids. Lancet. 1984;2:1058–1061. [PubMed] [Google Scholar]
- 2.Laikind PK, Seegmiller JE, Gruber HE. Detection of 5′-phosphoribosyl-4-(N-succinylcarboxamide)-5-aminoimidazole in urine by use of the Bratton-Marshall reaction: Identification of patients deficient in adenylosuccinate lyase activity. Anal. Biochem. 1986;156:81–90. doi: 10.1016/0003-2697(86)90158-2. [DOI] [PubMed] [Google Scholar]
- 3.Simmonds HA, Duley JA, Davies PM. Analysis of purines and pyrimidines in blood, urine, and other physiological fluids. In: Hommes FA, editor. Techniques in diagnostic human biochemical genetics. New York: Wiley-Liss; 1991. pp. 397–424. [Google Scholar]
- 4.Castro M, Pérez-Cerdá C, Merinero B, García MJ, Bernar J, Gil Nagel A, Torres J, Bermúdez M, Garavito P, Marie S, Vincent F, Van den Berghe G, Ugarte M. Screening for adenylosuccinate lyase deficiency: clinical, biochemical and molecular findings in four patients. Neuropediatrics. 2000;33:186–189. doi: 10.1055/s-2002-34493. [DOI] [PubMed] [Google Scholar]
- 5.Jaeken J, Wadman SK, Duran M, van Sprang FJ, Beemer FA, Holl RA, Theunissen PM, de Cock P, van den Bergh F, Vincent MF, van den Berghe G. Adenylosuccinase deficiency: an inborn error of purine nucleotide synthesis. Eur. J. Pediatr. 1988;148(1988):126–131. doi: 10.1007/BF00445919. [DOI] [PubMed] [Google Scholar]
- 6.Kmoch S, Hartmannová H, Stibůrková B, Krijt J, Zikánová M, Sebesta I. Human Adenylosuccinate Lyase (hADSL), cloning and characterization of full length cDNA and its Isoform, gene structure and molecular basis for ADSL deficiency in six patients. Hum. Mol. Genet. 2000;9:1501–1513. doi: 10.1093/hmg/9.10.1501. [DOI] [PubMed] [Google Scholar]
- 7.Spiegel EK, Colman RF, Patterson D. Adenylosuccinate lyase deficiency. Mol. Genet. Metab. 2006;89:19–31. doi: 10.1016/j.ymgme.2006.04.018. [DOI] [PubMed] [Google Scholar]
- 8.Gitiaux C, Ceballos-Picot I, Marie S, Valayannopoulos V, Rio M, Verrieres S, Benoist JF, Vincent MF, Desquerre I, Bahi-Buisson N. Misleading behavioural phenotype with adenylosuccinate lyase deficiency. Eur. J. Hum. Genet. 2009;17:133–136. doi: 10.1038/ejhg.2008.174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mouchegh K, Zikánová M, Hoffmann GF, Kretzschmar B, Kühn T, Mildenberger E, Stoltenburg-Didinger G, Krijt J, Dvoráková L, Honzík T, Zeman J, Kmoch S, Rossi R. Lethal fetal and early neonatal presentation of adenylosuccinate lyase deficiency: observation of 6 patients in 4 families. J. Pediatr. 2007;150:57–61. doi: 10.1016/j.jpeds.2006.09.027. [DOI] [PubMed] [Google Scholar]
- 10.Jurecka A, Zikanova M, Tylki-Szymanska A, Krijt J, Bogdanska A, Gradowska W, Mullerova K, Sykut-Cegielska J, Kmoch S, Pronicka E. Clinical, biochemical and molecular findings in seven Polish patients with adenylosuccinate lyase deficiency. Mol. Genet. Metab. 2008;94:435–442. doi: 10.1016/j.ymgme.2008.04.013. [DOI] [PubMed] [Google Scholar]
- 11.Chen BC, McGown IN, Thong MK, Pitt J, Yunus ZM, Khoo TB, Ngu LH, Duley JA. Adenylosuccinate lyase deficiency in a Malaysian patient, with novel adenylosuccinate lyase gene mutations. J. Inherit. Metab. Dis. 2010 Feb 23; doi: 10.1007/s10545-010-9056-z. [Epub ahead of print] [DOI] [PubMed] [Google Scholar]
- 12.Sempere A, Arias A, Farré G, García-Villoria J, Rodríguez-Pombo P, Desviat LR, Merinero B, García-Cazorla A, Vilaseca MA, Ribes A, Artuch R. J. Campistol, Study of inborn errors of metabolism in urine from patients with unexplained mental retardation. J. Inherit. Metab. Dis. 2010;33:1–7. doi: 10.1007/s10545-009-9004-y. [DOI] [PubMed] [Google Scholar]
- 13.Marie S, Race V, Nassogne MC, Vincent MF, Van den Berghe G. Mutation of a nuclear respiratory factor 2 binding site in the 5' untranslated region of the ADSL gene in three patients with adenylosuccinate lyase deficiency. Am. J. Hum. Genet. 2002;71:14–21. doi: 10.1086/341036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Marie S, Cuppens H, Heuterspreute M, Jaspers M, Tola EZ, Gu XX, Legius E, Vincent MF, Jaeken J, Cassiman JJ, Van den Berghe G. Mutation analysis in adenylosuccinate lyase deficiency: eight novel mutations in the re-evaluated full ADSL coding sequence. Hum. Mutat. 1999;13:197–202. doi: 10.1002/(SICI)1098-1004(1999)13:3<197::AID-HUMU3>3.0.CO;2-D. [DOI] [PubMed] [Google Scholar]
- 15.Race V, Marie S, Vincent MF, Van den Berghe G. Clinical, biochemical and molecular genetic correlations in adenylosuccinate lyase deficiency. Hum. Mol. Genet. 2000;9:2159–2165. doi: 10.1093/hmg/9.14.2159. [DOI] [PubMed] [Google Scholar]
- 16.Zikanova M, Skopova V, Hnizda A, Krijt J, Kmoch S. Biochemical and structural analysis of 14 mutant adsl enzyme complexes and correlation to phenotypic heterogeneity of adenylosuccinate lyase deficiency. Hum. Mutat. 2010;31:445–455. doi: 10.1002/humu.21212. [DOI] [PubMed] [Google Scholar]
- 17.Van den Bergh F, Vincent MF, Jaeken J, Van den Berghe G. Functional studies in fibroblasts of adenylosuccinase-deficient children. J. Inherit. Metab. Dis. 1993a;16:425–434. doi: 10.1007/BF00710293. [DOI] [PubMed] [Google Scholar]
- 18.Van den Bergh F, Vincent MF, Jaeken J, Van den Berghe G. Residual adenylosuccinase activities in fibroblasts of adenylosuccinase-deficient children: parallel deficiency with adenylosuccinate and succinyl-AICAR in profoundly retarded patients and non-parallel deficiency in a mildly retarded girl. J. Inherit. Metab. Dis. 1993b;16:415–424. doi: 10.1007/BF00710291. [DOI] [PubMed] [Google Scholar]
- 19.Mierzewska H, Schmidt-Sidor B, Jurkiewicz E, Bogdańska A, Kuśmierska K, Stepień T. Severe encephalopathy with brain atrophy and hypomyelination due to adenylosuccinate lyase deficiency--MRI, clinical, biochemical and neuropathological findings of Polish patients. Folia Neuropathol. 2009;47:314–320. [PubMed] [Google Scholar]
- 20.Patterson D. Biochemical genetics of Chinese hamster cell mutants with deviant purine metabolism, IV, Isolation and characterization of a mutant which accumulates adenylosuccinic acid and succinylaminoimidazole carboxamide ribotide, Somat. Cell Genet. 1976;2:189–203. doi: 10.1007/BF01538958. [DOI] [PubMed] [Google Scholar]
- 21.Tu AS, Patterson D. Biochemical genetics of Chinese hamster cell mutants with deviant purine metabolism. V1. Enzymatic studies of two mutants unable to convert inosinic acid to adenylic acid. Biochem. Genet. 1977;15:195–195. doi: 10.1007/BF00484561. [DOI] [PubMed] [Google Scholar]
- 22.Van Keuren ML, Hart IM, Kao F-T, Neve RL, Bruns GA, Kurnit DA, Patterson D. A somatic cell hybrid with a single human chromosome 22 corrects the defect in the CHO mutant (Ade-I) lacking adenylosuccinase activity. Cytogenet. Cell Genet. 1987;44:142–147. doi: 10.1159/000132358. [DOI] [PubMed] [Google Scholar]
- 23.Sivendran S, Colman RF. Effect of a new non-cleavable substrate analog on wild-type and serine mutants in the signature sequence of adenylosuccinate lyase of Bacillus subtilis and Homo sapiens. Protein Sci. 2008;17:1162–1174. doi: 10.1110/ps.034777.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bulusu V, Srinivasan B, Bopanna MP, Balaram H. Elucidation of the substrate specificity, kinetic and catalytic mechanism of adenylosuccinate lyase from Plasmodium falciparum. Biochim. Biophys. Acta. 2009;1794:642–654. doi: 10.1016/j.bbapap.2008.11.021. [DOI] [PubMed] [Google Scholar]
- 25.Frohman MA, Dush MK, Martin GR. Rapid production of full-length cDNAs from rare transcripts by amplification using a single gene-specific oligonucleotide primer. Proc. Natl. Acad. Sci. 1988;85:8998–9002. doi: 10.1073/pnas.85.23.8998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Loh EL, Elliott JF, Cwirla S, Lanier LL, Davis MM. Polymerase chain reaction with single sided specificity: Analysis of T cell receptor delta chain. Science. 1989;243:217–220. doi: 10.1126/science.2463672. [DOI] [PubMed] [Google Scholar]
- 27.Ohara O, Dorit RI, Gilbert W. One-sided PCR: The amplification of cDNA. Proc. Natl. Acad. Sci. 1989;86:5673–5677. doi: 10.1073/pnas.86.15.5673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang Z, Schwartz S, Wagner L, Miller W. A greedy algorithm for aligning DNA sequences. J. Comput. Biol. 2000;7:203–214. doi: 10.1089/10665270050081478. [DOI] [PubMed] [Google Scholar]
- 29.Knox AJ, Graham C, Bleskan J, Brodsky G, Patterson D. Mutations in the Chinese hamster ovary cell GART gene of de novo purine synthesis. Gene. 2009;429:23–30. doi: 10.1016/j.gene.2008.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Tornheim K, Lowenstein JM. The Purine Nucleotide Cycle: The Production of Ammonia from Aspartate by Extracts of Rat Skeletal Muscle. J. Biol. Chem. 1972;247:162–169. [PubMed] [Google Scholar]
- 31.Knox AJ. Ph.D. Thesis. 2007. The GART gene of purine biosynthesis: assessment of functional sites through mutagenesis in CHO cells and analysis of behavioral phenotypes in transgenic mice. [Google Scholar]
- 32.Spitzer J, Poolman B. The role of biomacromolecular crowding, ionic strength, and physicochemical gradients in the complexities of life’s emergence. Microbiol. Mol. Biol. Rev. 2009:371–388. doi: 10.1128/MMBR.00010-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhou H-X. Protein folding in confined and crowded environments. Arch. Biochem. Biophys. 2008;469:76–82. doi: 10.1016/j.abb.2007.07.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.You X, Nguyen AW, Jabaiah A, Sheff MA, Thorn KS, Daugherty PS. Intracellular protein interaction mapping with FRET hybrids. Proc. Natl. Acad. Sci. 2006;103:18458–18463. doi: 10.1073/pnas.0605422103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Zal T. Visualization of protein interactions in living cells. Adv. Exp. Med. Biol. 2008;640:183–197. doi: 10.1007/978-0-387-09789-3_14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ebbinghaus S, Dhar A, McDonald JD, Gruebele M. Protein folding stability and dynamics imaged in a living cell. Nat Methods. 2010;7:319–323. doi: 10.1038/nmeth.1435. [DOI] [PubMed] [Google Scholar]
- 37.Prudencio M, Durazo A, Whitelegge JP, Borchelt DR. Modulation of mutant superoxide dismutase 1 aggregation by co-expression of wild-type enzyme. J. Neurochem. 2009;108:1009–1018. doi: 10.1111/j.1471-4159.2008.05839.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Witan H, Gorlovoy P, Kaya AM, Koziollek-Drechsler I, Neumann H, Behl C, Clement AM. Wild-type Cu/Zn superoxide dismutase (SOD1) does not facilitate, but impedes the formation of protein aggregates of amyotrophic lateral sclerosis causing mutant SOD1. Neurobiol. Dis. 2009;36:331–342. doi: 10.1016/j.nbd.2009.07.024. [DOI] [PubMed] [Google Scholar]
- 39.An S, Kumar R, Sheets ED, Benkovic SJ. Reversible compartmentalization of de novo purine biosynthetic complexes in living cells. Science. 2008;320:103–106. doi: 10.1126/science.1152241. [DOI] [PubMed] [Google Scholar]
- 40.An S, Kyoung M, Allen JJ, Shokat KM, Benkovic SJ. Dynamic regulation of a metabolic multi-enzyme complex by protein kinase CK2. J. Biol. Chem. 2010;285:11093–11099. doi: 10.1074/jbc.M110.101139. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.