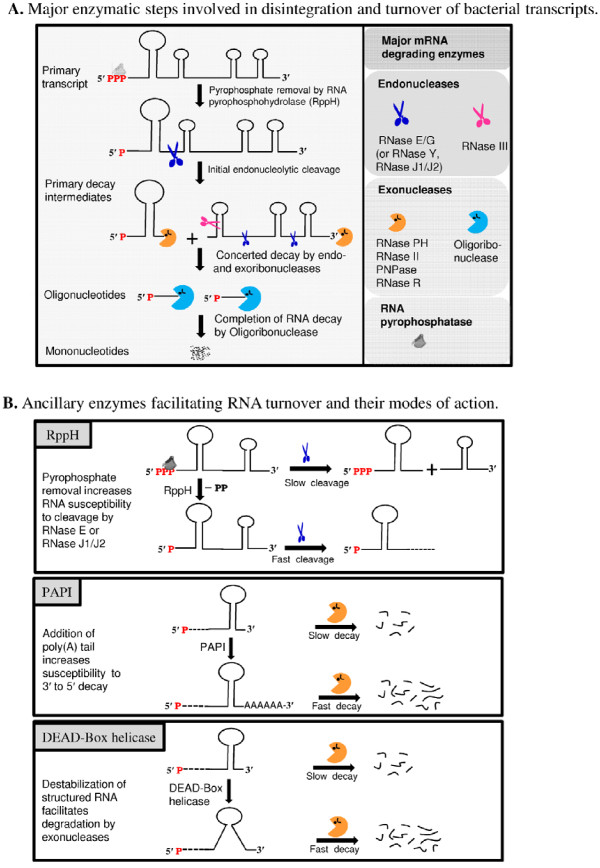
Figure 4.
Current unified model of mRNA decay pathways in Escherichia coli. (A) Schematic representation of major enzymatic steps involved in the disintegration and complete turnover of primary transcripts in E. coli. The decay of a regular transcript is usually initiated by endonucleolytic cleavage to generate primary decay intermediates that are further converted to short oligoribonucleotides by the combined action of exo- and endoribonucleases. The oligoribonucleotides are further degraded into mononucleotides by oligoribonuclease. (B) Ancillary enzymes facilitating mRNA turnover and their modes of action. Degradation of mRNA can be stimulated via pyrophosphate removal by RppH, which converts 5'-triphosporylated primary transcripts into their monophosphorylated variants, thus facilitating their endoribonucleolytic cleavage by RNase E [22,76] or by RNases J1/J2 [12] or by RNase Y [16] in B. subtilis. As the action of exoribonucleases can be inhibited by 3'-terminal stem-loop structures, two groups of ancillary RNA-modifying enzymes, PAPI and RhlB, help exonucleases to overcome this inhibitory effect. PAPI exerts its action by adding short stretches of adenosine residues, thereby facilitating exonuclease binding and subsequent cleavage of structured RNAs [10]. Enzymes of the second group, DEAD-box helicases such as E. coli RhlB, increase the efficiency of the exonuclease-dependent decay by unwinding double-stranded RNA regions in an ATP-dependent fashion.