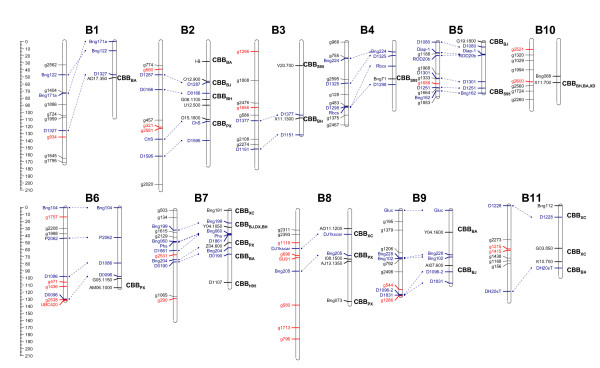
Figure 4.
The distribution of molecular markers co-localized with previously indentified QTLs associated to CBB resistance. For each linkage group, the map on the left is reproduced from McClean(2007) map (http://www.comparative-legumes.org/) [33], the map on the right is reproduced from comprehensive Freyre (1998) map (http://www.comparative-legumes.org/), adopted from Miklas et al (2006) [3]. Both maps are integrated by shared markers except for linkage group B10. In McClean (2007) map, only molecular markers used in association study and shared markers in blue were shown. The markers in red were found significantly (P ≤ 0.05) associated with CBB resistance. In Freyre (1998) map, loci placed on the left side of each chromosome were shared markers in blue and molecular markers closest to previous identified CBB-QTLs. To the right of each linkage group are previously identified CBB-QTLs in different populations [3]. Symbols in subscript represent the source population of the QTL: BA Belneb-RR-1/A55, BJ BAT93/JaloEEP558, BH BAC6/HT7719, DX DOR364/XAN176, H95 HR67/OAC95, PX PC50/XAN159, S95 Seaforth/OAC95 and XC XR-235-1-1/Calima. Marker UBC420, SU91, and QTL locations are approximate because most were not directly mapped in the BAT93/JaloEEP558 population. The total distance of each linkage group is expressed in cM (Kosambi mapping function).