Abstract
Cortical interneurons, generated predominantly in the medial ganglionic eminence, migrate around and avoid the developing striatum in the subpallium en route to the cortex. This is attributable to the chemorepulsive cues of class 3 semaphorins expressed in the striatal mantle and acting through neuropilin (Nrp1 and Nrp2) receptors expressed in these cells. Cortical interneurons also express Robo receptors, and we show here that in mice lacking Robo1, but not Robo2, these cells migrate aberrantly through the striatum. In vitro experiments demonstrated that interneurons lacking Robo1 function are significantly less responsive to the effects of semaphorins. Failure to respond to semaphorin appears to be attributable to a reduction in Nrp1 and PlexinA1 receptors within these cells. Biochemical studies further demonstrated that Robo1 binds directly to Nrp1, but not to semaphorins, and this interaction is mediated by a region contained within its first two Ig domains. Thus, we show for the first time that Robo1 interacts with Nrp1 to modulate semaphorin signaling in the developing forebrain and direct the migration of interneurons through the subpallium and into the cortex.
Introduction
The medial ganglionic eminence (MGE) in the subpallium is the major source of GABA-containing cortical interneurons during corticogenesis (Lavdas et al., 1999; Corbin et al., 2001; Wichterle et al., 2001). En route to the cerebral cortex, MGE-derived interneurons encounter the developing striatum and migrate around it (Marín and Rubenstein, 2003; Métin et al., 2006). Work in the past decade has identified some of the molecular cues that guide migrating cortical interneurons through the subpallium (for review, see Marín and Rubenstein, 2003; Métin et al., 2006; Hernández-Miranda et al., 2010). These include two class 3 semaphorins, Sema3A and Sema3F, and their receptors, the neuropilins and plexins. Studies by Marín et al. (2001) have shown that interneurons destined to populate the cortex express both neuropilin 1 (Nrp1) and Nrp2 receptors, which enable them to respond to chemorepulsive Sema3A and Sema3F signals emanating from the striatal mantle. This repulsive activity in the developing striatum creates an exclusion zone for migrating interneurons to channel them into adjacent paths, leading to the formation of the migratory routes to the cortex.
We have previously shown that migrating cortical interneurons express Robo proteins (Robo1, Robo2, Robo3), the receptors for Slit ligands (Andrews et al., 2006, 2008; Barber et al., 2009). Our analysis of mice deficient for each receptor (Robo1−/−, Robo2−/−, Robo3−/− mice, respectively) revealed that Robo1−/−, but not Robo2−/− or Robo3−/−, animals contained significantly more calbindin-positive (CB+) cells in their cortices than wild-type littermates, indicating that more cortical interneurons were present in the cerebral cortex of these mice (Andrews et al., 2006, 2008; Barber et al., 2009). In addition, developing Robo1−/− mice contained more CB+ cells within the striatum than control littermates (Andrews et al., 2006), raising the possibility that loss of Robo1 causes cortical interneurons to migrate through this structure, as reported for neuropilin mutants (Marín et al., 2001). However, CB is also expressed by the vast majority of mature striatal projection neurons (Liu and Graybiel, 1992; Ouimet et al., 1998), and the exact nature of the increased CB+ cells in the developing striatum of Robo1−/− mice remains to be determined.
Here, we used a panel of different markers for striatal projection neurons and for cortical and striatal interneurons to demonstrate an influx of cortical interneurons into the developing striatum of Robo1−/− mice. Using chemotaxis assays, we found that MGE-derived interneurons of Robo1−/− mice were markedly less responsive to Sema3A and Sema3F and, furthermore, this effect was not attributable to a direct interaction between class 3 semaphorins and Robo1. Expression studies illustrated downregulation of class 3 semaphorin receptors, especially Nrp1 and PlexinA1, in MGE cells from Robo1−/− mice. Moreover, biochemical experiments revealed that Nrp1 is able to bind directly to the Ig domains 1 and 2 of Robo1. Together, our data demonstrate that Robo1 modulates semaphorin–neuropilin/plexin signaling to steer interneurons around the striatum and into the cortex.
Materials and Methods
Animals.
All experimental procedures were performed in accordance with the United Kingdom Animals (Scientific Procedures) Act 1986 and institutional guidelines. Wild-type animals were C57BL/6J mice obtained from Charles River. Slit1−/−/Slit2−/−, Robo2−/−, and Robo1−/− mice were generated as described previously (Plump et al., 2002; Hammond et al., 2005; Andrews et al., 2008, respectively). GAD67-GFP (Δneo) mice (Tamamaki et al., 2003) used in this study were also maintained in C57BL/6J background. The day the vaginal plug was found was considered to be embryonic day 0.5 (E0.5). Animals of either sex were used in our experiments.
Cell lines.
GN11 cells (Radovick et al., 1991) and COS-7 cells (American Type Culture Collection) were grown as a monolayer at 37°C in a humidified CO2 incubator in complete DMEM (Invitrogen) and supplemented with 10% fetal bovine serum (FBS) (Invitrogen). Subconfluent cells were harvested by trypsinization and cultured in 10 cm2 dishes. Cells within six passages were used in all experiments.
Immunohistochemistry.
Embryonic brains (E13.5–E18.5) were fixed in 4% paraformaldehyde (PFA), made in PBS, for 4–8 h at room temperature (RT). Adult mice were perfused through the heart with 4% PFA, and their brains removed and immersed in fixative solution at RT for 4 h. After fixation, embryonic and adult brains were cryoprotected in 30% sucrose in PBS, embedded and frozen in a mixture of 15% sucrose/50% Tissue-Tek OCT (Sakura Finetek), and sectioned coronally at 20 μm using a cryostat (Bright Instruments). Sections were washed in PBS and blocked in PBS containing 5% normal goat serum (NGS) (Sigma-Aldrich) (v/v) and 0.1% Triton X-100 (v/v) (Sigma-Aldrich) at RT for 2 h. They were subsequently incubated in primary antibodies at RT for 2 h and then, at 4°C overnight. The following antibodies were used: rabbit anti-calbindin (1:3000; CB-28; Swant), rabbit anti-Caspase3 (R&D Systems), goat anti-choline acetyltransferase (ChAT) (1:250; Millipore), rabbit anti-DARPP-32 (1:500; Millipore), rabbit anti-Forkhead box protein P2 (FOXP2) (1:700; Abcam), mouse anti-parvalbumin (PV) (1:250; Swant), and rabbit anti-somatostatin (SST) (1:100; Millipore). After incubation in primary antibodies, sections were washed in PBS, incubated in biotinylated anti-species (1:250; Vector Laboratories) for 2 h, and processed using conventional immunohistochemistry protocols described previously (Andrews et al., 2006).
GN11 cells were fixed in 1% PFA made in PBS for 15 min. They were then washed in PBS and blocked in a solution of 5% NGS (v/v) and 0.5% Triton X-100 (v/v) (Sigma-Aldrich) in PBS at RT for 2 h. Subsequently, they were incubated in primary antibodies at RT for 2 h and then at 4°C overnight. The following antibodies were used: rabbit anti-Robo1 (1:5000), rabbit anti-Robo2 (1:5000), and rabbit anti-Robo3 (1:1000) (all Robo antibodies were raised and characterized in the laboratory of Prof. F. Murakami, Osaka University, Osaka, Japan) (Tamada et al., 2008).
Quantification of labeled cells in the striatum.
All morphometric analyses were conducted separately for rostral, middle, and caudal levels of the striatum based on the following anatomical landmarks. In embryonic tissue, the rostral level was considered where the septum was clearly identifiable, the middle level was selected where the intraventricular foramen and the anterior-dorsal thalamus were present, and the caudal level was chosen where the telo-diencephalic junction was distinguishable and the caudal ganglionic eminence was present. In postnatal tissue, the rostral level was chosen where the septum was clearly visible, the middle level was selected where the anterior commissure crosses bilaterally, and the caudal level was considered where the hippocampus was present. To determine the size of the striatal area, sections were stained with 0.025% thionin solution for 2 min and rinsed in a series of alcohols (70–100%). Striatal area was estimated using the NIH ImageJ software, version 1.3. To assess the total number of immunoreactive cells throughout the rostral-caudal extent of the striatum, a minimum of three nonconsecutive sections were stained for each marker per animal, age, and condition.
Production of semaphorin- and Slit-conditioned media.
Full-length chicken Sema3A-myc cDNA and Flag-tagged mouse Sema3F-Flag cDNA were provided by A. Cariboni (Cariboni et al., 2007). Full-length human Slit1-myc cDNA was kindly provided by Dr. V. Sundaresan (Kings College London, London, UK) (Patel et al., 2001). PMT21-myc and p3xFlag-CMV-8 were chosen as control plasmids. For transfection, COS-7 cells (at 80% confluence) were grown in culture plates in complete culture medium for 24 h and incubated with the selected expression vector (1 μg/ml) for 4 h in the presence of Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions. COS-7 cells were transfected with Sema3A-myc plasmid (CM-Sema3A), Sema3F-Flag plasmid (CM-Sema3F), Slit1-myc plasmid (CM-Slit1), pCDNA3.1-myc plasmid (CM-myc), or p3xFlag-CMV-8 plasmid (CM-Flag) and left in serum-free medium for 24 h. Cell supernatants were collected in ice-cold tubes, centrifuged at 3000 × g for 5 min, and immediately used for chemotaxis assays. Secretion of Sema3A, Sema3F, and Slit1 was confirmed by immunohistochemistry and Western blot analysis using anti-myc and anti-Flag antibodies (data not shown) (Cariboni et al., 2007).
Dissociated cell cultures.
Dissociated cell cultures were prepared from E13.5 brains according to the method of Cavanagh et al. (1997). Briefly, MGEs were dissected in Hanks solution under a stereo-microscope and dissociated enzymatically in Neurobasal medium (Invitrogen) containing 0.1% trypsin (Sigma-Aldrich) and 0.001% DNaseI (Roche) at 37°C for 15 min. Trypsin was inactivated by 10% FBS in Neurobasal media at 37°C for 5 min, and cells were dissociated by trituration with a sterile pipette tip. The resulting suspension was centrifuged at 1000 × g for 3 min, the supernatant was discarded, and, subsequently, cells were resuspended in Neurobasal media containing B27 supplement (Invitrogen) and 100 μg/ml penicillin/streptomycin. They were then plated onto 13 mm coverslips coated with 10 μg/ml poly-l-lysine and 10 μg/ml laminin in 24-well plates at a density of 104 cells/50 μl.
Chemotaxis assays.
Chemotaxis assays were performed using a 48-well Boyden's chamber (NeuroProbe) as described previously (Cariboni et al., 2007). Briefly, either GN11 or dissociated MGE cells were suspended in serum-free medium (105 cells/50 μl) and placed in the open-bottom wells of the upper compartment of the chamber. These were separated from the lower chamber by a polycarbonate porous membrane (8 μm pores), precoated with either gelatin (0.2 mg/ml) or laminin (10 μg/ml) for GN11 or dissociated MGE cells, respectively. Twenty-seven microliters of chemotactic agents were placed into the lower compartment of the chamber. The chamber was kept in an incubator at 37°C for 4 h (GN11 cells) or overnight (MGE cells). After incubation, the migrated cells that adhered to the underside of the membrane were fixed and stained using the Diff-Quick kit (Reagena). For quantitative analysis, the membranes were observed using an Olympus light microscope with a 20× objective adapted with a 500 × 500 μm grid. Four random fields of stained cells were counted for each well, and the mean number of migrating cells per square millimeter for each experimental condition was estimated.
Matrigel explants.
Small pieces of E13.5 MGE were dissected, as described previously, and placed onto 13 mm coverslips coated with 10 μg/ml poly-l-lysine and 10 μg/ml laminin in 24-well plates. Explants were embedded in a mixture of 75% Matrigel solution (BD Biosciences) and 25% conditioned media (CM) or CM containing chemotactic cues supplemented with B27 (Invitrogen). After a period of 30 min to allow the Matrigel to set, 1 ml of CM or CM-chemotactic cues supplemented with B27 was added, depending on the experiment. Explants were cultured for 2 d in vitro (DIV) in a sterile incubator (37°C, 5% CO2). Cell migration from MGE explants was assessed by the Sholl analysis (Sholl, 1953). Briefly, 10 concentric circles of gradually increasing radius (spaced every 30 μm from the edge of the explants) were drawn on digitized pictures of MGE explants. Cell migration from the explants was estimated by counting the number of concentric circles (or intersections) that the cells reached after incubation.
Fluorescent-activated cell sorting.
GN11 cells were transfected with Robo1 dominant-negative-GFP construct as previously described (Hammond et al., 2005). Two days after transfection, cells were washed three times in PBS before addition of CM-Flag or CM-Sema3F for 2 h. They were, then, washed again in PBS and trypsinized. Trypsin was inactivated by 10% FBS in Neurobasal medium for 5 min, and cells were washed in Neurobasal medium (without FBS) to remove serum for fluorescent-activated cell sorting (FACS). GN11 and dissociated MGE Robo1+/−;GAD67-GFP and Robo1−/−;GAD67-GFP cells (see above) were resuspended in Neurobasal medium without phenol red (Invitrogen) containing l-glutamine (Invitrogen). FACS was performed at the Wolfson Scientific Support Services (University College London) using a MoFlo Sorter (Dako). Untransfected GN11 cells, or GAD67-GFP-negative MGE cells, were used as controls for fluorescence. Cells were excited using a 488 nm argon laser and detected using a 530/40 (FL1) bandpass filter. Cell purity of 95–98.5% was obtained for each sort.
Quantitative real-time PCR.
Total RNA was extracted from FACS dissociated MGE cells taken from E15.5 Robo1−/−;GAD67-GFP and Robo1+/−;GAD67-GFP mice, and GN11-GFP-positive (Robo1-DN-transfected) and -negative (mock) FACS-purified cells using the QIAGEN RNeasy Plus kit. RNA was treated with DNaseI to remove any remaining trace amounts of DNA. The quality of the RNA was assessed using an Agilent bioanalyzer nanochip (Agilent). cDNA was generated with 25 ng of RNA using the QIAGEN Whole Transcriptome Amplification kit, as described in the manufacturer's protocol. Primers for quantitative real-time PCR (QPCR) were designed by Sigma-Genosys and were as follows: β-Actin (forward, GGCTGTATTCCCCTCCATCG; reverse, CCAGTTGGTAACAATGCCATGT); Flt1 (forward, TGGCTCTACGACCTTAGACTG; reverse, CAGGTTTGACTTGTCTGAGGTT); Gapdh (forward, ATGACATCAAGAAGGTGGTG; reverse, CATACCAGGAAATGAGCTTG); Kdr (forward, TTTGGCAAATACAACCCTTCAGA; reverse, GCAGAAGATACTGTCACACC); Nrp1 (forward, GGATGGATTCCCTGAAGTTG; reverse, TGGATAGAACGCCTGAAGAG); Nrp2 (forward, GCTGGCTACATCACTTCCCC; reverse, CAATCCACTCACAGTTCTGGTG); PlexinA1 (forward, CAGCACAGACAACGTCAACAA; reverse, GCTTGAAGAGATCGTCCAACC); PlexinA2 (forward, AACCTGTCTGTGGTTCTGCTC; reverse, TCCAGTCACGATTCTCAGAGT); PlexinA4 (forward, TGAGGACAACCCCAAGTGTTA; reverse, ACGCGATCAGCCTGTTTTCT); Robo1 (forward, GACCTGATCGTCTCCAAAGGA; reverse, TTGTCGGTCTCCACTCTTTCC); Robo2 (forward, TGATGGATCTCGTCTTCGTCA; reverse, GTCGGCCCTCTGCTTTACAG); Robo3 (forward, GCGCTTCTCAGTGTCTCCAAG; reverse, TGGTCCCTGGAGGATGACA); Vegfr3 (forward, GGTTGTGCATGACTGTGAAGG; reverse, GCGTGTCAGGTTTGTTGATGAA). The QPCR was performed using Sybr Green reagent (Sigma-Aldrich) on a Chromo4 PTC-200 Real-Time PCR Detector System (Bio-Rad). PCR conditions were 94°C for 2 min, followed by 40 three-step cycles of 94°C, 15 s; 60°C, 30 s; 72°C, 30 s. Gapdh and β-Actin were used for endogenous reference gene controls. Each primer set amplified a single PCR product of predicted size as determined by melt-curve analysis after PCR and by agarose gel electrophoresis, and had approximately equal amplification efficiency when validated using a serial dilution of representative cDNA. Each QPCR was performed in triplicate, and relative quantification was determined according to the ΔΔCt method (Livak and Schmittgen, 2001; Faux et al., 2010).
In situ hybridization.
In situ hybridization was performed as described previously (Faux et al., 2010). Briefly, embryonic brains were dissected in PBS and fixed in PFA overnight, followed by cryoprotection in 30% sucrose treated with diethyl pyrocarbonate (DEPC) made in PBS at 4°C for 2 d. Brains were frozen in Tissue-Tek OCT and sectioned coronally at 12 μm. Sections were dried at RT for 2 h, before overnight incubation at 65°C in hybridization buffer 1× DEPC-treated “salts” (200 mm NaCl, 5 mm EDTA, 10 mm Tris, pH 7.5, 5 mm NaH2PO4·2H2O, 5 mm Na2HPO4; Sigma-Aldrich), 50% deionized formamide (Ambion), 0.1 mg/ml RNase-free yeast tRNA (Invitrogen), 1× Denhardts (RNase/DNase free; Invitrogen), 10% dextran sulfate (Sigma-Aldrich) containing 100–500 ng/ml digoxigenin (DIG)-labeled RNA probes. Probes were generated by linearization of plasmids with appropriate enzymes and reverse transcription polymerases to obtain antisense probes. Probes used were as follows: Lhx6, Lhx8, Nkx2.1, Dlx1, and Dlx2 (kindly provided by Dr. Nicoletta Kessaris, Wolfson Institute, University College London, London, UK). After hybridization, sections were washed three times in a solution containing 50% formamide 1× SSC (Ambion) and 0.1% Tween 20 (Sigma-Aldrich) at 65°C and two times at RT in 1× MABT (20 mm maleic acid, 30 mm NaCl, 0.1% Tween 20; Sigma-Aldrich) before incubating in a solution containing 2% blocking reagent (Roche) and 10% NGS in MABT, followed by overnight incubation in alkaline phosphatase (AP)-conjugated anti-DIG antibody (1:1500; Roche). Nitroblue tetrazolium chloride (NBT) (Roche)/5-bromo-4-chloro-3-indolyl phosphate (BCIP) (Roche) diluted 1:1000 in MABT with 5% polyvinyl alcohol (VWR International) was used for the colorimetric detection for 6 h. Sections were mounted using Glycergel Mounting Medium (Dako).
Semaphorin coimmunoprecipitation binding assays.
COS-7 cells were transfected with expression vectors for Nrp1, Nrp2, PlexinA1, PlexinA4, Robo1, and pCDNA3.1 (control), using Lipofectamine. After 2 DIV, cells were incubated with CM-Sema3F (Flag-tagged) at 37°C for 2 h. Cells were washed with PBS and lysed on ice with immunoprecipitation (IP) buffer [50 mm Tris-HCl, pH 7.4, 100 mm NaCl, 2.5 mm MgCl2, 1% Triton X-100 (Sigma-Aldrich), and mixtures of protease inhibitors (Boehringer)]. The lysate was incubated at 4°C for 30 min and centrifuged for 5 min. After preclearing with protein A-agarose (Invitrogen) at 4°C for 2 h, samples were immunoprecipitated with mouse anti-myc antibody (5 μl) on protein A preformed immunocomplexes for 16 h at 4°C. The beads were then washed, and precipitates were analyzed on immunoblots with anti-Flag (Sigma-Aldrich; 1:1000) and anti-myc (1:1000) antibodies.
Semaphorin-AP binding assays.
Sema3A-AP and Sema3F-AP were prepared as previously described (Vieira et al., 2007). COS-7 cells were fixed in absolute methanol for 5 min, washed five times with PBS, incubated in PBS containing 0.1% Triton X-100 (PBT) and 10% FBS for 30 min, and then reacted with AP-fusion protein at RT for 2 h. Sections were then washed 5 min each with PBS, fixed in 4% PFA at RT for 2 min, and washed again. Endogenous AP was heat inactivated by incubation at 65°C for 3 h. Tissue-bound heat-stable recombinant AP activity was detected as an insoluble reaction product after incubation with NBT and BCIP.
Expression of neuropilin and plexin receptors.
Dissociated MGE cells or GN11 cells were lysed in IP buffer. The lysate was incubated at 4°C for 30 min and centrifuged for 5 min. Cell lysates were processed for conventional SDS-PAGE and membrane transfer. To assess the protein levels of Nrp1, Nrp2, PlexinA1, Robo1, and β-Actin, membranes were incubated with the following polyclonal antibodies: Nrp1 (1:1000; Santa Cruz Biotechnology), Nrp2 (1:1000; Santa Cruz Biotechnology), PlexinA1 (1:1000; Santa Cruz Biotechnology), Robo1 (1:1000; R&D Systems), and β-Actin (1:500; Sigma-Aldrich), in 5% BSA-TBST, washed several times with TBST, and incubated with a horseradish peroxidase-conjugated secondary antibody (1:5000; Vector Laboratories). After intensive washing, the proteins were visualized with ECL detection reagent (GE Healthcare).
Robo coimmunoprecipitation assay.
E15.5 mouse forebrains were lysed by trituration in IP buffer, and incubated at 4°C for 30 min before centrifuging for 5 min to remove cellular debris. After preclearing with protein A-agarose at 4°C for 2 h, samples were immunoprecipitated with the following polyclonal antibodies: anti-Nrp1 antibodies (5 μl), anti-Nrp2 antibodies (5 μl), anti-PlexinA1 antibodies (5 μl), anti-Robo1 (C-terminal) antibodies (5 μl) (Prince et al., 2009), anti-Robo2 (C-terminal) antibodies (5 μl) (Cho et al., 2007), anti-VEGFR3 antibodies (5 μl; Abcam), on protein A preformed immunocomplexes at 4°C for 16 h. The beads were then washed, and precipitates were analyzed on immunoblots with the following polyclonal antibodies: anti-Nrp1 (1:1000), anti-Nrp2 (1:1000), anti-PlexinA1 (1:1000), and anti-Robo1 (N-terminal) (1:1000).
Fc binding assays.
COS-7 cells were transfected with expression vectors for Nrp1, Nrp2, PlexinA1, PlexinA4, Robo1, and pCDNA3.1 (control), using Lipofectamine. After 2 DIV, cells were incubated with either Nrp1-Fc (1 μg; R&D Systems), Robo1-Fc (1 μg; R&D Systems), or Robo1 deletion-Fc proteins (1 μg) (Liu et al., 2004) at 37°C for 2 h. Cells were washed with PBS and lysed on ice in IP buffer (containing 10% low Ig sera; Sigma-Aldrich). The lysate was incubated at 4°C for 30 min and centrifuged for 5 min. Protein A-agarose (50 μl) was added to lysates and mixed at 4°C for 16 h. The beads were then washed, and precipitates were analyzed on immunoblots with anti-myc (1:1000) antibody.
Covasphere aggregation assay.
The covasphere aggregation assay was performed as previously described (Liu et al., 2004). In brief, 50 μl samples of fluorescent beads (coated with anti-human Fc antibody; Spherotech) were washed with PBS and resuspended in 15 μl of PBS/5% FBS. The Fc-chimera (1 μg in 25 μl) was added and the beads were incubated at 37°C for 2 h. They were then washed and resuspended in 50 μl of PBS/5% FBS. To start the assay, the beads were first triturated and sonicated in an ice bath to break up any preformed aggregates. The sample was then warmed in a 37°C water bath, and the degree of bead aggregation was monitored as a function of time (taken as t = 0 min) using an inverted microscope (Liu et al., 2004).
Digital image acquisition and processing.
Optical and fluorescent images were collected using a Leica light microscope (DM5000B; Leica Microsystems). Images were reconstructed and digitized with Photoshop CS4 software (Adobe Systems).
Statistics.
Statistical analyses were performed by GraphPad 3 software (GraphPad Software). All data are reported as mean number and SEM. The statistical significance between group means was tested by one-way ANOVA (one-way ANOVA), followed by Bonferroni's post hoc test (for multiple-comparison tests). Significance was set at a value of p ≤ 0.05.
Results
The developing striatum of Robo1-, but not Robo2-null mice, contains ectopic cortical interneurons
We previously reported that deletion of exon 5 of the Robo1 gene in mice increased the number of CB+ cells (presumptive interneurons) in the embryonic cerebral cortex when compared with control littermates (Andrews et al., 2006). This observation was subsequently confirmed and extended in a new line of Robo1-deficient Dulox mice (hereafter Robo1−/− mice), in which the entire gene for the Robo1 receptor (exons 1–22 inclusive) was deleted (Andrews et al., 2008). Previous reports have also shown that expression of one copy of Robo1 gene is sufficient to promote normal embryonic development (Andrews et al., 2008). Consistent with this, in the present study we did not find any difference in the number of CB+ cells in cortical and subcortical structures of Robo1+/+ and Robo1+/− mice (data not shown). Therefore, we used here Robo1+/− mice as control animals.
To explore whether the increased number of interneurons was restricted to the cortices of Robo1−/− mice, we cross-mated Robo1−/− mice with GAD67-GFP animals to generate Robo1+/−;GAD67-GFP and subsequently Robo1−/−;GAD67-GFP mice (Robo1 control and knock-out animals, respectively). Examination of the forebrains of E15.5 Robo1−/−;GAD67-GFP mice revealed that, in addition to the greater number of GABAergic (GAD67-GFP+) cells in the cortex (Fig. 1A–D), the developing striatum also presented an excess of labeled cells when compared with Robo1+/−;GAD67-GFP animals (Fig. 1A,B,E–G). This is in agreement with our previous finding of increased CB staining in the striatum of Robo1 deficient (exon 5 deleted) mice (Andrews et al., 2006).
Figure 1.
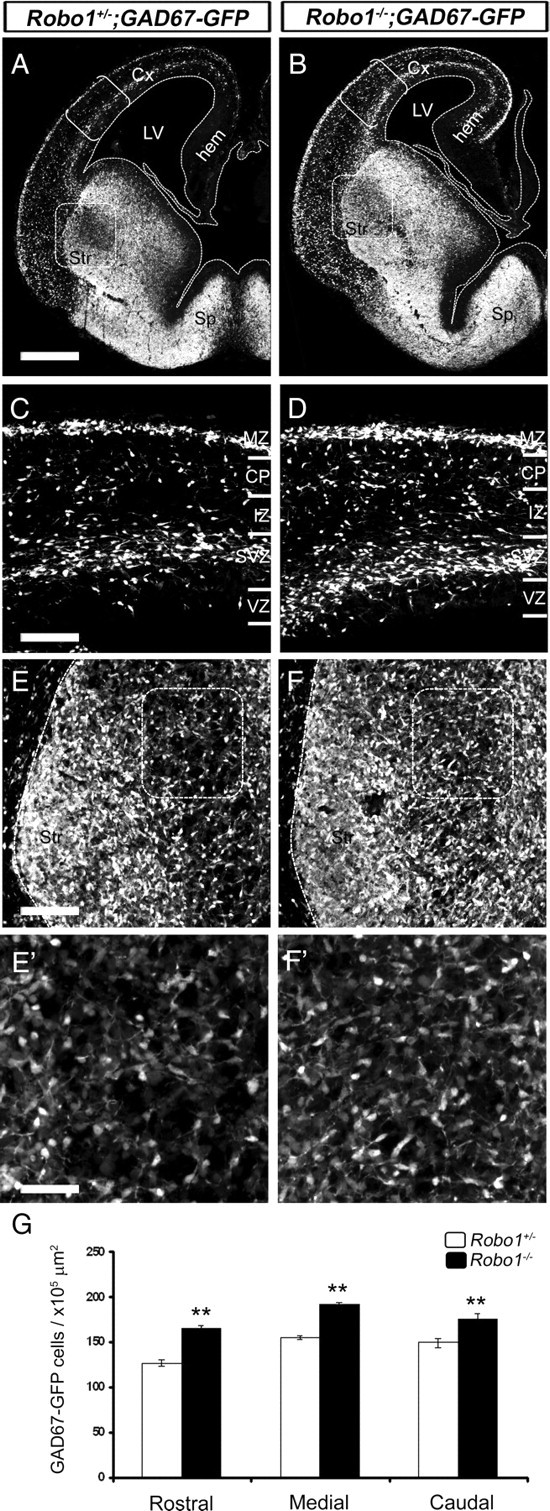
Deletion of Robo1 receptor increases the number of GABAergic neurons in the cerebral cortex and striatum. A, B, Coronal sections through the brains of a Robo1+/−;GAD67GFP mouse (A) and a Robo1−/−;GAD67-GFP littermate (B) at E15.5. The areas bracketed in A and B are shown at higher magnifications in C and D, respectively, whereas boxes in A and B are shown at higher magnifications in E and F, respectively. E′, F′, The boxed areas in E and F are shown at an even higher magnification. G, Counts of GAD67-GFP+ cells show increased number throughout the rostral-caudal extent of the striatum of Robo1−/− mice at E15.5. Scale bars: A, B, 500 μm; C–F′, 150 μm. **p < 0.01. Error bars indicate SEM. Abbreviations: CP, Cortical plate; Cx, cerebral cortex; hem, cortical hem; IZ, intermediate zone; LV, lateral ventricle; MZ, marginal zone; Sp, septum; Str, striatum; SVZ, subventricular zone; VZ, ventricular zone.
The striatum is a structure in which both projection neurons and interneurons express GABA. To assess whether the increased number of cells in the striatum of Robo1−/−;GAD67-GFP mice were striatal projection neurons, we immunostained coronal sections from Robo1−/− mice and Robo1+/− littermates for the transcription factor FOXP2, a marker of these cells in embryonic life (Takahashi et al., 2003). Counts of labeled cells at E15.5 and E18.5 (n = 4 per age for each condition) throughout the rostral-caudal extent of the striatum showed no difference between the two groups of animals at the two ages examined (Fig. 2A–F). Therefore, deletion of the Robo1 receptor does not alter the generation of striatal projection neurons.
Figure 2.
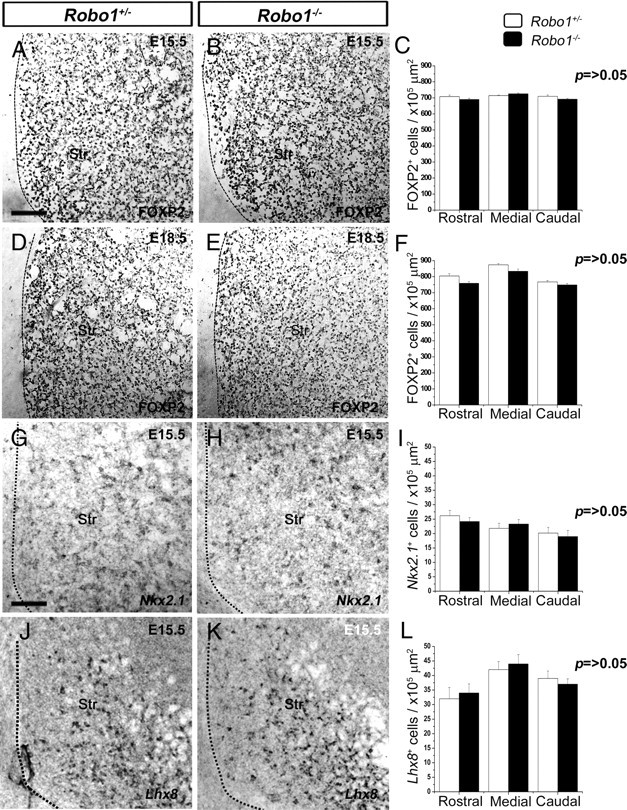
No differences in the number of developing striatal projection neurons or striatal interneurons between Robo1+/− and Robo1−/− embryos. A, B, D, E, Coronal sections from Robo1+/− and Robo1−/− mice at E15.5 (A, B) and at E18.5 (D, E) were immunostained for FOXP2. C, F, Counts of FOXP2+ cells showed no differences throughout the rostral-caudal extent of the developing striatum of Robo1−/− and Robo1+/− littermates at E15.5 (C) and E18.5 (F). G, H, Coronal brain sections taken from Robo1+/− and Robo1−/− mice at E15.5 and processed by in situ hybridization for Nkx2.1. J, K, Similar sections were processed by in situ hybridization for Lhx8. I, L, Counts of labeled cells showed no differences for either gene between the two groups of animals. Scale bars: A, B, D, E, 100 μm; G, H, J, K, 120 μm. Abbreviation: Str, striatum. Error bars indicate SEM.
We then explored whether the additional number of GAD67+ cells in the striatum of Robo1−/−;GAD67-GFP mice were developing striatal interneurons, a population of cells that normally accounts for <5% of all striatal neurons (Gerfen, 1992). Thus, we analyzed by in situ hybridization the striatum of Robo1+/− and Robo1−/− mice (n = 3 per condition) at E15.5 for the expression of Nkx2.1, a transcription factor expressed by striatal, but not cortical interneurons (Nóbrega-Pereira et al., 2008), and for Lhx8, a marker of striatal cholinergic cells (Zhao et al., 2003; Fragkouli et al., 2009). This analysis showed no difference in the number of Nkx2.1+ or Lhx8+ cells between Robo1−/− and control littermates (Fig. 2G–L), suggesting that deletion of Robo1 receptor does not affect the generation of striatal interneurons during development.
We also analyzed sections taken from adult (3 months) Robo1−/− (n = 3) and Robo1+/− (n = 3) mice and immunostained for CB and DARPP-32 (markers of ∼85 and 98% of mature striatal projection neurons, respectively) (Ouimet et al., 1998). This analysis showed no differences in the number of CB+ (Fig. 3A–C) or DARPP-32+ cells (Fig. 3D–F) at any level of the adult striatum of Robo1−/− mice compared with Robo1+/− littermates. We then assessed whether the number of mature striatal interneurons was changed in mice lacking Robo1 receptor by immunostaining sections of adult Robo1−/− (n = 3) and Robo1+/− (n = 3) mice for three markers of nonoverlapping subpopulations of striatal interneurons, PV, SST, and ChAT (Kawaguchi et al., 1995). Counts of the three cell types also showed no differences between the two groups of animals throughout the rostral-caudal extent of the striatum (Fig. 3G–O). Together, our analysis of developing and adult animals suggests that deletion of Robo1 receptor does not impair the establishment of striatal projection neurons or interneurons.
Figure 3.
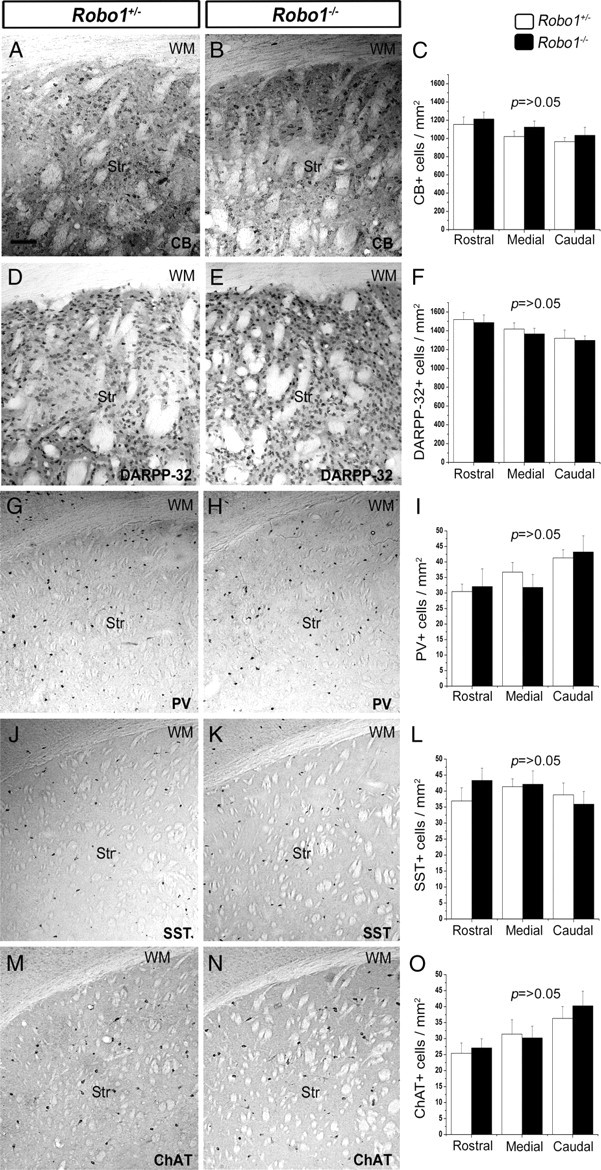
No differences in the number of striatal projection neurons or interneurons between Robo1+/− and Robo1−/− adult mice. Immunostaining of coronal brain sections from adult Robo1+/− and Robo1−/− mice for the striatal projection neuron markers CB (A, B) and DARP-32 (D, E), and for the interneuron markers PV (G, H), SST (J, K), and ChAT (M, N). The quantitation of immunopositive neurons for each marker is shown adjacent to the sections (C, F, I, L, O). Scale bar, 100 μm. Abbreviations: Str, Striatum; WM, white matter. Error bars indicate SEM.
We then wanted to quantify the previously observed increase in CB staining in the developing striatum of Robo1-deficient mice (Andrews et al., 2006). Thus, we counted labeled neurons at E15.5, E18.5, and postnatal day 0 (P0) (n = 3 per age for each condition) and found a significant increase throughout the rostral-caudal extent of the striatum of Robo1−/− mice when compared with Robo1+/− littermates at all three ages (Fig. 4). At E15.5, the increase was ∼23% (p < 0.01) (Fig. 4A–C); at E18.5, ∼39% (p < 0.01) (Fig. 4D–F); and at P0, ∼16% (p < 0.05) (Fig. 4G–I). The decrease in the number of CB+ cells over time within the developing striatum of Robo1−/− mice could be attributable to either an increase in cell death or migration through this structure. Previous studies had reported a very small number of dying cells within the striatum during embryogenesis, increasing somewhat during the first postnatal week (Fentress et al., 1981). To test the first possibility, we stained the striatum of Robo1+/− and Robo1−/− littermates at E15.5 and E18.5 with the apoptotic marker Caspase3 (Hoshi et al., 1998). Although we observed immunostaining in the marginal zone (MZ) and ventricular zone (VZ) of all animals tested, we failed to observe any Caspase3+ cells within the developing striatum of Robo1+/− and Robo1−/− littermates of either age (data not shown). The observed increase in the number of CB+ cells in the striatum only during the period of cortical interneuron migration through the ventral forebrain, coupled with the finding of no change in the number of striatal projection neurons or interneurons, led us to hypothesize that the excess of CB+ and GAD67+ cells represents aberrantly migrating cortical interneurons.
Figure 4.
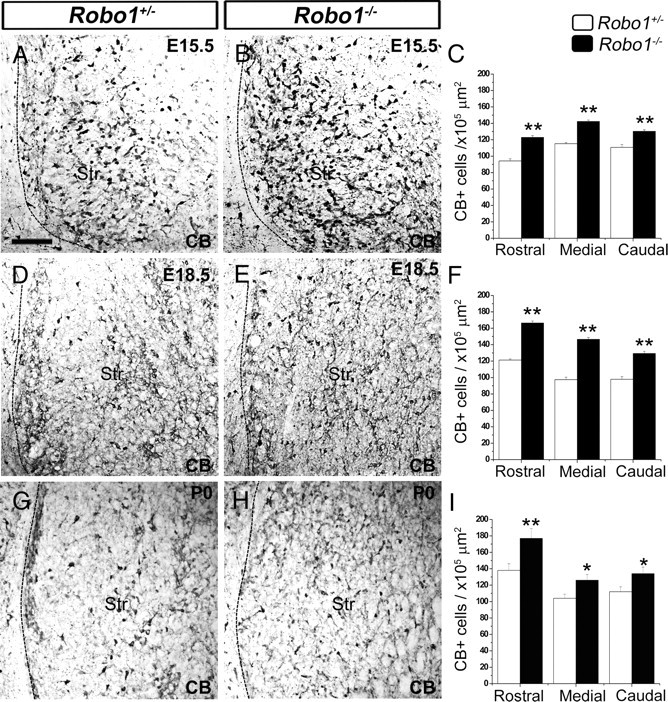
Increased number of CB+ cells in the striatum of Robo1−/− mouse embryos. Immunostaining of coronal sections of Robo1+/− and Robo1−/− brains for CB at E15.5 (A, B), E18.5 (D, E), and P0 (G, H). The quantification of the number of CB+ cells in the striatum of Robo1−/− animals and Robo1+/− littermates at each age is shown adjacent to the sections (C, F, I, respectively). Scale bar, 100 μm. *p < 0.05; **p < 0.01. Abbreviation: Str, Striatum. Error bars indicate SEM.
We previously demonstrated that cortical interneurons also express Robo2 receptors, but that Robo2−/− mice do not show significant differences in the number and position of these cells in their cortices compared with Robo2+/− littermates, suggesting that this receptor is not involved in their migration (Andrews et al., 2008). Here, we analyzed the developing striatum and found no significant difference in the number of differentiating projection neurons (FOXP2+ cells) in Robo2−/− (n = 4) mice compared with Robo2+/− (n = 4) littermates throughout the rostral-caudal extent at E15.5 (Fig. 5A–C). Counts of CB+ cells also showed no difference at middle and caudal levels of the striatum between Robo2−/− and Robo2+/− animals (n = 4 for each condition) (Fig. 5D–F). However, there was a significant increase in the number of CB+ cells in Robo2-deficient mice rostrally at E15.5 (Robo2+/−, 96.8 ± 2.4 cells/×105 μm2; Robo2−/−, 127.5 ± 2.3 cells/×105 μm2; p < 0.01). Thus, our previous and current observations suggest that, unlike the Robo1 receptor, Robo2 does not play an important role in the migration of interneurons through the developing striatum or into the cortex.
Figure 5.
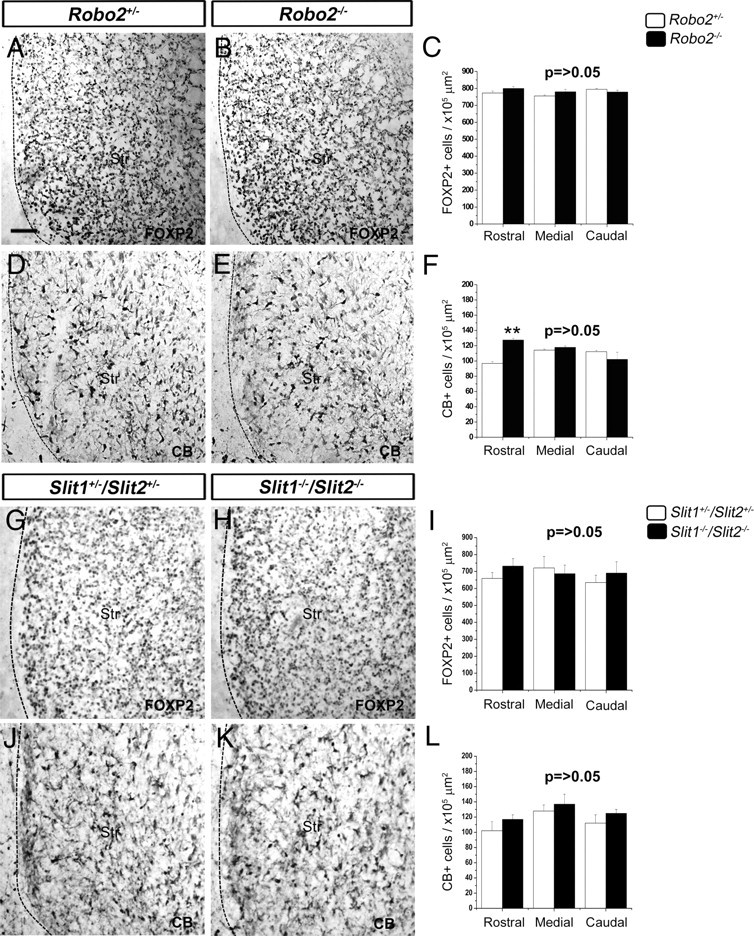
No differences in the number of CB+ or FOXP2+ cells in the striatum of Robo2−/−, Slit1−/−;Slit2−/− mice, and control littermates. A, B, D, E, Coronal sections taken from the brains of Robo2+/+ (A, D) and Robo2−/− (B, E) mice at E15.5 were immunostained for FOXP2 (A, B) and CB (D, E). C, F, Quantitation of the number of FOXP2+ and CB+ cells in the striatum at E15.5. G, H, J, K, Coronal sections from brains of Slit+/−;Slit2+/− (G, J) and Slit1−/−;Slit2−/− (H, K) mice at E15.5 were immunostained for FOXP2 (G, H) and CB (J, K). I, L, Quantitation of the number of FOXP2+ and CB+ cells in the striatum at E15.5. Scale bar, 100 μm. **p < 0.01. Abbreviation: Str, Striatum. Error bars indicate SEM.
We next asked whether the striatal defect in Robo1−/− mice is Slit dependent, as Slits are expressed in the basal telencephalon (Marillat et al., 2002). However, we found no significant difference in the number of FOXP2+ or CB+ cells at any level in E15.5 Slit1−/−;Slit2−/− double null mutants, compared with Slit1+/−;Slit2+/− controls (n = 3 for each condition) (Fig. 5G–L), suggesting that Slit1/Slit2 signaling does not play a role in striatal development. Ectopic interneuron migration into the developing striatum of Robo1−/− mice is, therefore, a Slit1/Slit2-independent event and must be attributable to the action of other directional guidance cues.
MGE-derived cells are less responsive to the class 3 semaphorin-induced chemorepulsion in the absence of Robo1
Because Sema3A and Sema3F have been implicated in the striatal repulsion of cortical interneurons, we asked whether Robo1 modulates semaphorin signaling in chemomigration assays. First, we assessed the migratory potential of neurons of MGE explants prepared from E13.5 Robo1−/− (n = 260) and Robo1+/− (n = 209) mice and treated with control CM (CM-myc or CM-Flag) after 2 DIV. We quantified cell movement away from the explants by applying the analysis of Sholl (see Materials and Methods). Our evaluation revealed no significant differences in the migratory potential between Robo1−/− and Robo1+/− cells in MGE explants treated with CM-myc (Robo1+/−, 6.36 ± 0.71 intersections; Robo1−/−, 7.08 ± 0.42 intersections; p > 0.05) or CM-Flag (Robo1+/−, 7.55 ± 0.32 intersections; Robo1−/−, 6.74 ± 0.29 intersections; p > 0.05), suggesting that the motility of Robo1−/− MGE-derived cells is not impaired (Fig. 6A,B,E,F). We then examined the response of cells from explants taken from Robo1+/− mice to CM-Sema3A (n = 63) or CM-Sema3F (n = 110) treatment. We observed a significant reduction in the migration of MGE-derived cells compared with those treated with CM-myc (n = 42) or CM-Flag (n = 130), respectively (CM-myc, 6.36 ± 0.71 intersections; CM-Sema3A, 1 ± 0.071 intersections; p < 0.001; CM-Flag, 7.55 ± 0.32 intersections; CMSema3F, 4.52 ± 0.52 intersections; p < 0.01) (Fig. 6A–F). In contrast, MGE explants taken from Robo1−/− mutants and treated with CM-Sema3A (n = 54) or CM-Sema3F (n = 129) showed no differences in their migratory capacity when compared with similar explants treated with CM-myc (n = 42) or CM-Flag (n = 111), respectively (CM-myc, 7.08 ± 0.42 intersections; CM-Sema3A, 7.48 ± 0.08 intersections; CM-Flag, 6.34 ± 0.3 intersections; CM-Sema3F, 6.48 ± 0.32 intersections; p > 0.05) (Fig. 6B,D–F). This finding suggested that MGE neurons derived from Robo1−/− mice are less responsive to Sema3A- or Sema3F-induced chemorepulsion.
Figure 6.
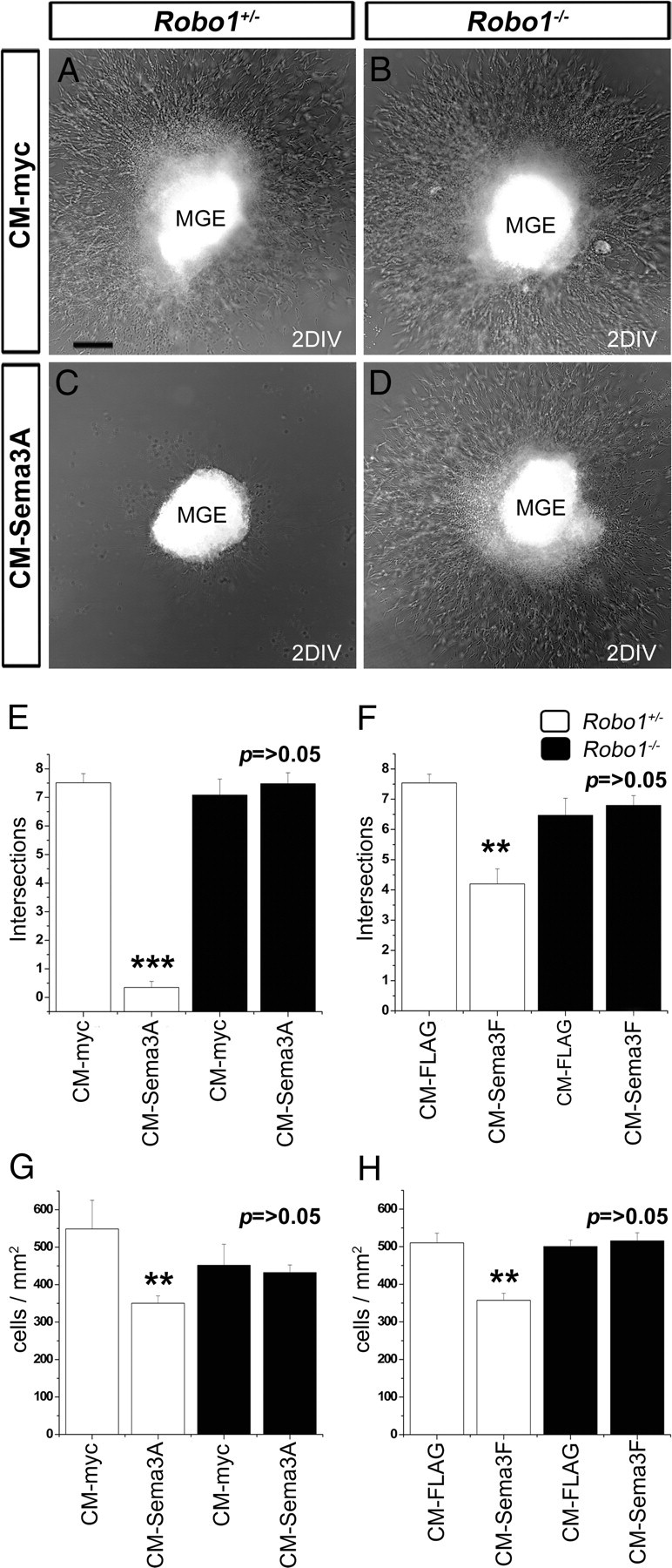
MGE-derived cells from Robo1−/− mutants do not respond to Sema3A or Sema3F. A–D, Migration of cells away from E13.5 MGE explants prepared from Robo1+/− (A, C) and Robo1−/− (B, D) littermates treated with control conditioned media (CM-myc) (A, B) or CM-Sema3A (C, D). E, F, Quantification of migration from Robo1+/− and Robo1−/− MGE explants treated with Sema3A (E) or Sema3F (F). G, H, Quantification of the number of dissociated MGE cells from Robo1+/− and Robo1−/− mice used in a Boyden's chamber assay to test their response to Sema3A (G) or Sema3F (H). Scale bar, 100 μm. **p < 0.01; ***p < 0.001. Error bars indicate SEM.
To provide additional evidence for this hypothesis, we compared how dissociated cells taken from the MGE of Robo1−/− and Robo1+/− littermates (n = 12) at E13.5 respond to control-CM, CM-Sema3A, or Sema3F in a Boyden's chamber assay. We found that cells taken from Robo1−/− and Robo1+/− animals responded similarly to control-CM (CM-myc Robo1+/−, 548.7 ± 76.3 cells/mm2; CM-myc Robo1−/−, 451.8 ± 55.99 cells/mm2; CM-Flag Robo1+/−, 510.3 ± 25.4 cells/mm2; CM-Flag Robo1−/−, 500.6 ± 16.8 cells/mm2; p > 0.05) (Fig. 6G,H), supporting the notion that deletion of the Robo1 gene does not impair the normal motility of MGE cells, and confirming our previous observations using explants (Fig. 6E,F). We also observed, as expected, a significant reduction in the number of MGE-derived cells taken from Robo1+/− mice that migrated toward compartments containing either CM-Sema3A or CM-Sema3F when compared with control-CM (CM-myc, 548.7 ± 76.3 cells/mm2; CM-Sema3A, 350.3 ± 19.7 cells/mm2; p < 0.01; CM-Flag, 510.3 ± 25.4 cells/mm2; CM-Sema3F, 357.3 ± 18.7 cells/mm2; p < 0.01) (Fig. 6G,H). Interestingly, Robo1−/− MGE dissociated cells did not respond to either Sema3A (CM-myc, 451.8 ± 55.99 cells/mm2; CM-Sema3A, 432.34 ± 20.21 cells/mm2; p > 0.05), or Sema3F (CM-Flag, 500.6 ± 16.8 cells/mm2; CM-Sema3F, 515.7 ± 21.1 cells/mm2; p > 0.05). Together, these in vitro data indicate that Robo1 deficiency reduces the responsiveness of MGE cells to Sema3A and Sema3F.
GN11 cells: a cell model for studying Slit-Robo and semaphorin signaling
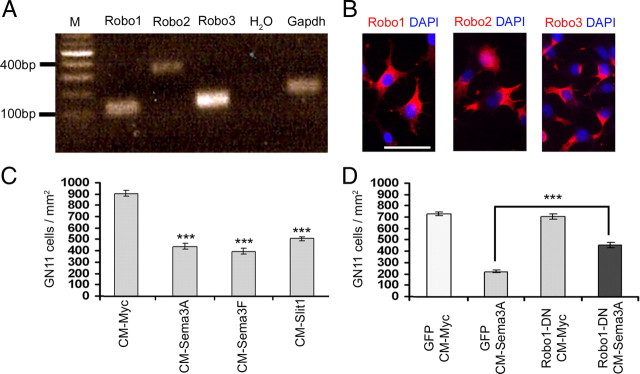
To further characterize the putative role of Robo1 receptor in semaphorin signaling, we wanted to determine whether the observed reduction in response was specific to cortical interneurons, or whether it applied to other migrating neuronal cell types. In a previous study, we had demonstrated that immortalized gonadotropin-releasing hormone secreting (GnRH) neurons (GN11 cells) express Nrp1 and Nrp2 receptors and respond to Sema3A and Sema3F (Cariboni et al., 2007). Here, we explored whether GN11 cells also possess Robo receptors and respond to the chemotactic action of Slit. We found that these cells, like interneurons, expressed mRNA for Robo1, Robo2, and Robo3 by reverse transcription-PCR (Fig. 7A) and by immunohistochemistry with Robo-specific antibodies (Fig. 7B).
Figure 7.
GN11 cells express Robo receptors and respond to class 3 semaphorins and Slit1. A, Reverse transcription-PCR analysis showed expression of Robo1–3 in GN11 cells. B, Immunohistochemistry confirmed the expression of Robo receptors in GN11 cells. C, D, Quantification of GN11 cell migration in a Boyden's chamber, containing Sema3A, Sema3F, Slit1, or control (CM-myc) conditioned media (C). D, Robo1-DN-transfected GN11 cells are less responsive to Sema3A than control (GFP)-transfected cells. Scale bar, 100 μm. ***p < 0.001. Error bars indicate SEM.
We then treated GN11 cells in chemomigration assays with either control CM-myc, CM-Sema3A, CM-Sema3F, or Slit1 (CM-Slit1). This analysis showed a reduction in the number of migrating GN11 cells into compartments containing either Sema3A (CM-myc, 855.8 ± 24.1 cells/mm2; CM-Sema3A, 415.3 ± 25.3 cells/mm2; p < 0.001) or Sema3F (CM-myc, 855.8 ± 24.1 cells/mm2; CM-Sema3F, 375.3 ± 25.3 cells/mm2; p < 0.001) compared with control media (Fig. 7C), confirming previous results (Cariboni et al., 2007). Similarly, Slit1 reduced significantly the number of migrating cells (CM-myc, 855.8 ± 24.1 cells/mm2; CM-Slit1, 481 ± 17.6 cells/mm2; p < 0.001) (Fig. 7C), as did Slit2 and Slit3 (data not shown). These results indicated that GN11 cells are repelled by both class 3 semaphorins and Slits.
Because GN11 cells express Nrp and Robo receptors and respond to class 3 semaphorins and Slits, we asked whether Robo1 modulates their response to Sema3A and Sema3F as observed for cortical interneurons. We, therefore, transfected GN11 cells with a control construct (GFP; n = 4) or a construct encoding GFP-tagged dominant-negative Robo1 (Robo1-DN; n = 4) (Hammond et al., 2005). After transfection, we assessed their ability to migrate in the presence of control CM (CM-myc) or Sema3A (CM-Sema3A) in a Boyden's chamber assay. We found that GN11 cells transfected with Robo1-DN migrated similar to GFP-transfected cells in the presence of control CM (GFP, 731.5 ± 15.4 cells/mm2; Robo1-DN, 708 ± 23.2 cells/mm2; p > 0.05). In contrast, cells transfected with Robo1-DN construct were less responsive to Sema3A chemorepulsion compared with cells transfected with GFP alone (Robo1-DN, 458.2 ± 24.12 cells/mm2; GFP, 222.5.5 ± 9.8 cells/mm2; p < 0.001) (Fig. 7D). Similar results were obtained with CM-Sema3F (data not shown). These findings suggest that, similar to cortical interneurons, loss of Robo1 function perturbs semaphorin signaling in GN11 cells.
Given that disruption of Robo1 signaling affects the response of Robo1−/− MGE-derived cells and Robo1-DN GN11 cells to Sema3A and Sema3F, we next asked whether Robo1 receptor could bind directly to class 3 semaphorins. To address this possibility, we transfected COS-7 cells with constructs for myc-tagged Robo1, Nrp1, Nrp2, PlexinA1, PlexinA4, or with pCDNA3.1 (control). After 2 DIV, conditioned media containing Sema3F (Flag-tagged) was added. Cell lysates were immunoprecipitated with anti-myc antibody, and immunoblotted with anti-Flag antibody to detect Sema3F. In agreement with previous results (Chen et al., 1997), Sema3F bound strongly Nrp2-transfected cells. In contrast, Sema3F did not bind to Robo1-transfected cells (Fig. 8A). To confirm these results, we performed a Sema3F-AP binding assay with transfected COS-7 cells. Again, Sema3F appeared to bind strongly Nrp2-transfected cells, but not Robo1-transfected or control cells (Fig. 8B). To assess the ability of Sema3A to bind to Robo1, we repeated the AP binding assay using Sema3A-AP and observed strong binding to Nrp1-transfected, but no binding to Robo1-expressing cells (Fig. 8B). Thus, neither Sema3A nor Sema3F appear to bind to Robo1.
Figure 8.
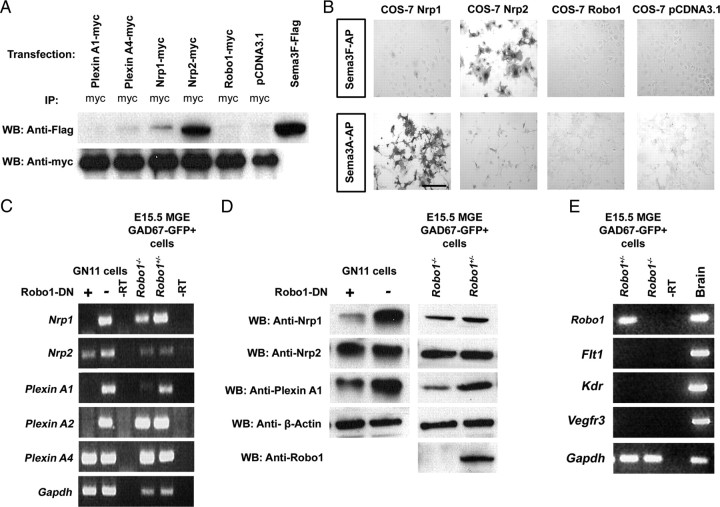
Downregulation of Nrp and PlexinA in MGE cells derived from Robo1−/−;GAD67-GFP mice and Robo1-DN-transfected GN11 cells. A, B, Sema3A and Sema3F do not bind to Robo1. Cell lysates from COS-7 cells, which had been transiently transfected with myc-tagged full-length Plexin, Nrp, or Robo1 and treated with Flag-tagged Sema3F in the culture medium, were immunoprecipitated with anti-myc antibody and immunoblotted with anti-Flag and anti-myc antibodies (A). B, Sema3A-AP or Sema3F-AP was added to COS-7 cells transiently transfected with full-length Nrp1, Nrp2, Robo1, or control. Sema3A binding was only observed with cells expressing Nrp1, and Sema3F binding was only observed with cells expressing Nrp2; no binding was observed to cells expressing Robo1. C, QPCR for semaphorin receptors was performed on GN11 cells transfected with Robo1-DN or control (GFP) constructs and of MGE cells derived from E15.5 Robo1+/−;GAD67-GFP and Robo1−/−;GAD67-GFP mice. D, Immunoblot analyses of MGE cells derived from E15.5 Robo1+/−;GAD67-GFP and Robo1−/−;GAD67-GFP mice, and of GN11 cells transfected with Robo1-DN cells showed reduced levels of Nrp1 and PlexinA1, but not Nrp2. Abbreviations: −RT, Without reverse transcriptase (negative control). Genes examined are listed next to the gel bands. E, QPCR for VEGF receptors was performed on MGE-derived cells from E15.5 Robo1+/−;GAD67-GFP and Robo1−/−;GAD67-GFP mice. Genes examined are listed next to the gel bands. Lack of expression of VEGF receptors was observed in MGE cells derived from both groups of animals but were present in total brain extract.
Downregulation of Nrp1, PlexinA1, and PlexinA2 receptors in Robo1−/− derived MGE and Robo1-DN-transfected GN11 cells
It has been reported that loss of Nrp function results in abnormal migration of cortical interneurons through the striatum (Marín et al., 2001). Since our data showed that Robo1−/− MGE-derived cells and Robo1-DN-transfected GN11 cells are less responsive to semaphorin chemorepulsion, we next asked whether Robo1 regulates the expression of Nrp or PlexinA receptors. QPCR on FACS MGE-derived cells taken from Robo1−/−;GAD67-GFP (n = 3) and Robo1+/−;GAD67-GFP (n = 3) littermates at E15.5 revealed a significant reduction in the expression of Nrp1 (−3.56-fold; p < 0.01) and PlexinA1 (−11.2-fold; p < 0.001), but no significant changes in the levels of Nrp2, PlexinA2, or PlexinA4 (Fig. 8C). Similar results were obtained with GN11 cells transfected with Robo1-DN. Specifically, we observed a significant decrease in the expression of Nrp1, Nrp2, PlexinA1, and PlexinA2, whereas the level of PlexinA4 was unaffected.
To determine whether the differences in the levels of class 3 semaphorin receptor mRNAs observed in MGE-derived cells of Robo1−/− animals translate into corresponding changes in protein levels, we performed Western blot analysis for Nrp1, PlexinA1 (the receptors that showed the greatest reduction in our QPCR), and Nrp2 on dissociated cells taken from the MGE of E15.5 Robo1−/− and Robo1+/− animals, as well as GN11 cells transfected with Robo1-DN. Our analysis confirmed that loss of Robo1 signaling reduces the levels of Nrp1 and PlexinA1, but not Nrp2 protein (Fig. 8D). Together, our data suggest that absence of Robo1 signaling in MGE-derived neurons and in GN11 cells downregulates Nrp1, PlexinA1, and PlexinA2 receptors, and seems to explain the similarity of the cortical interneuron defect in mice lacking Robo1 or semaphorin signaling through neuropilins.
In addition to modulating class 3 semaphorins, neuropilin receptors have been shown to function as coreceptors with Kdr, Flt1, and VEGFR3 to modulate vascular endothelial growth factor (VEGF) signaling in endothelial cells (Giraudo et al., 1998; Fuh et al., 2000; Gluzman-Poltorak et al., 2001; Catalano et al., 2004). Therefore, we asked whether reduction in Nrp1 levels in cortical interneurons of Robo1−/− mice affects VEGF receptor expression. However, we found that none of the VEGF receptors were expressed in FACS cells from the MGE of Robo1−/−;GAD67-GFP (n = 3) and Robo1+/−;GAD67-GFP (n = 3) littermates at E15.5. This indicates that cortical interneurons do not rely on VEGF signaling, at least during the peak of their migration (Fig. 8E).
Interaction between Robo1 and Nrp1
To assess whether there exist physical interactions between Nrp and PlexinA receptors with Robo1, we immunoprecipitated E15.5 forebrain lysates with antibodies specific for Nrp1, Nrp2, PlexinA1, VEGFR3, C-terminal Robo1 (Prince et al., 2009), and C-terminal Robo2 (Cho et al., 2007). We then immunoblotted with antibodies specific for Nrp1, Nrp2, PlexinA1, or Robo1 N terminus. Robo1 immunoprecipitated with Nrp1, Nrp2, and PlexinA1 antibodies, suggesting Robo1 receptor exists in complexes with all of these components of the semaphorin receptor (Fig. 9A). Robo1 also immunoprecipitated with Robo2, confirming previously observed heterophilic interactions (Liu et al., 2004), but VEGFR3 only shows a weak interaction. To confirm these results in a heterologous system, we transfected GN11 cells with expression vectors for myc-tagged Robo1, Nrp1, Nrp2, or PlexinA1. After 2 DIV, cell lysates were immunoprecipitated with anti-myc antibody and subjected to immunoblotting with antibodies for Nrp1, Nrp2, PlexinA1, or Robo1. Again, Robo1 coimmunoprecipitated with Nrp1, Nrp2, and PlexinA1 antibodies (data not shown).
Figure 9.
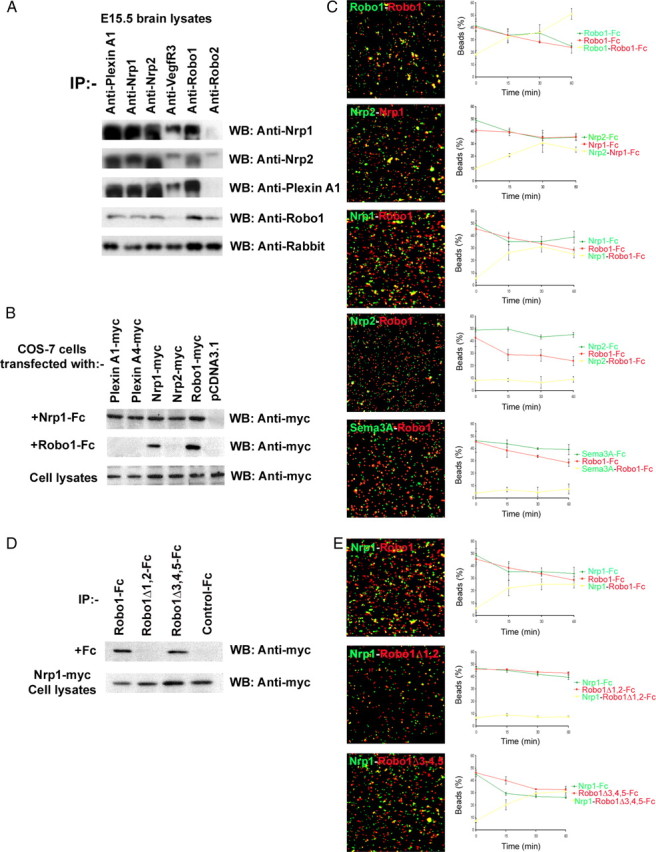
Robo1 interacts directly with Nrp1. A, E15.5 brain lysates were immunoprecipitated and immunoblotted with the indicated antibodies; note that Robo1 forms complexes with Robo2, Nrp1, Nrp2, and PlexinA1, but not VEGFR3. B, Cell lysates from COS-7 cells, transiently transfected with constructs for the indicated proteins or control expression vector, were immunoprecipitated with Fc-tagged Nrp1 or Robo1, and immunoblotted with anti-myc antibody. Robo1 bound homophilically to itself and heterophilically to Nrp1. C, Covasphere aggregation assay to identify Robo1 heterophilic interactions. Beads coated with the indicated Fc proteins were imaged with a 25× objective at the end of the aggregation assay. Histograms show the corresponding changes in the proportion of beads (red, green, and yellow) over time; heterophilic interactions are indicated by yellow. Robo1 appears to bind homophilically to itself, and heterophilically to Nrp1, but not Nrp2 or Sema3A. D, Cell lysates from COS-7 cells, which were transiently transfected with myc-tagged full-length Nrp1 and incubated with Robo1-Fc, Robo1Δ1,2-Fc, or Robo1Δ3,4,5-Fc in the culture medium, were immunoprecipitated with the same Fc-tagged protein and immunoblotted with anti-myc antibody. Robo1Δ1,2-Fc failed to coimmunoprecipitate Nrp1, suggesting that the neuropilin binding domain resides within the first two Ig domains of Robo1. E, Covasphere aggregation assays using Nrp-Fc (green) and Robo1-Fc, Robo1Δ1,2-Fc, and Robo1Δ3,4,5-Fc (red) confirmed these findings. Error bars indicate SEM.
To identify Robo1 binding partners, COS-7 cells were transfected with constructs expressing myc-tagged Nrp1, Nrp2, Robo1, PlexinA1, PlexinA4, or with pCDNA3.1 (control) and incubated with either Nrp1-Fc or Robo1-Fc chimeras. After immunoprecipitation with Fc tag, immunoblots were probed with anti-myc antibody. Homophilic Robo1 interactions were observed between Robo1-myc/Robo1-Fc, confirming previous reports (Hivert et al., 2002; Liu et al., 2004; Camurri et al., 2005). Heterophilic complexes formed between Nrp2-myc/Nrp1-Fc, Nrp1-myc/Robo1-Fc, and Robo1-myc/Nrp1-Fc, suggesting that Robo1 is able to specifically interact with itself and Nrp1, but not with Nrp2, PlexinA1, or PlexinA4 (Fig. 9B).
To confirm these findings, we used the covasphere assay, which has previously been used to identify Robo receptor interactions (Liu et al., 2004; Camurri et al., 2005). In these experiments, two types of Fc proteins were coated onto red and green fluorescent beads and mixed. If the covaspheres bind homophilically, the beads would show red or green fluorescent aggregates when viewed under epifluorescence. If heterophilic interactions take place, then there would be an overlap of the red and green fluorescence, resulting in yellow aggregates. We found that beads coated with Robo1 (red) and Robo1 (green) showed coaggregation, as did beads coated with Nrp1 (red) and Nrp2 (green) (Fig. 9C). These results indicate that Robo1 can bind homophilically and Nrp1–Nrp2 can bind heterophilically, in agreement with our immunoprecipitation data. When beads coated with Robo1 (red) and Nrp1 (green) were mixed, they coaggregated (yellow), indicating that they bind to each other (Fig. 9C). However, when Robo1 (red) beads were mixed with either Sema3A or Nrp2 (green) covasphere beads, small aggregates formed, and they were composed entirely of either red or green cluster of beads, indicating that there is no heterophilic interaction between Robo1 and Sema3A or Nrp2, in agreement with our previous experiments.
Using Robo1 Ig domain deletion constructs in coimmunoprecipitation and covasphere assays, we further found that Robo1 receptor lacking the first two Ig domains (Robo1Δ1,2-Fc) did not bind to Nrp1, whereas Robo1 lacking Ig domains 3, 4, 5 (Robo1Δ3,4,5-Fc) bound to Nrp1, suggesting that the critical binding region lies within the first two Ig domains of Robo1 (Fig. 9D,E). Together, our in vitro data suggest that Robo1 binds specifically to Nrp1, and not to Sema3A, Sema3F, Nrp2, or any of its other (co)receptors, and that the first two Ig domains of Robo1 are essential for this interaction.
Discussion
Cortical interneurons, generated predominantly in the MGE, migrate around the developing striatum en route to the cortex (Corbin et al., 2001; Marín and Rubenstein, 2003; Métin et al., 2006; Hernández-Miranda et al., 2010). These neurons express Robo receptors (Robo1, Robo2, Robo3), and our analyses of mice lacking each of the receptors (Robo1−/−, Robo2−/−, Robo3−/−) showed that only Robo1−/− animals contained significantly more CB+ cells, presumptive interneurons, in their cortex compared with wild-type littermates, both during development and in adulthood (Andrews et al., 2006, 2008; Barber et al., 2009). We also observed that developing Robo1−/− mice had increased CB staining within the striatum compared with control littermates and speculated that these cells may represent cortical interneurons aberrantly migrating through the striatum on the way to the cortex (Andrews et al., 2006). This effect appears to be specific to Robo1 receptor, as no changes in striatal projection cells or interneurons were observed in Robo2−/− mice. Robo3 is also unlikely to play a role, as it is expressed in a subpopulation of cortical interneurons at the early stages of corticogenesis and is markedly downregulated in the developing forebrain by E14.5 (Camurri et al., 2004; Barber et al., 2009).
However, our previous studies did not establish whether the excess CB staining in the Robo1−/− mice were ectopic cortical interneurons migrating through the striatum or cells destined to become striatal projection neurons, which are known to express the calcium binding protein (Liu and Graybiel, 1992; Ouimet et al., 1998). In the present study, we have provided several pieces of evidence that together support the former possibility. First, examination of Robo1−/−;GAD67-GFP animals showed an excess of labeled cells, most likely cortical interneurons, in the striatum compared with Robo1+/−;GAD67-GFP mice. Previous studies had reported that cortical interneurons contain higher levels of GAD67, unlike striatal cells that preferentially express GAD65 (Greif et al., 1992; Mercugliano et al., 1992; Feldblum et al., 1993). Second, using specific markers for striatal projection neurons and interneurons in developing and mature animals, we found no differences between Robo1−/− animals and control littermates. Thirdly, the quantification of CB+ cells showed an increased number in the developing striatum that mirrored the wave of newly generated cortical interneurons migrating through the subpallium en route to the cortex, reaching a peak at E18.5 and diminishing in later ages. The diminution in excess CB+ cells in the striatum of Robo1−/− mice around the perinatal period did not appear to be attributable to cell death, as it was not accompanied by an increase in apoptotic cells. Moreover, the ∼30% increase in CB+ cells in the developing striatum of Robo1−/− embryos matched the increased number of interneurons reported in the mature cortex of these animals (Andrews et al., 2008). The increase in interneuron numbers may be attributed, at least in part, to the observed increase in proliferation in the MGE of Robo1−/− animals (Andrews et al., 2008). Interestingly, this event is independent of the Slit ligands as previous (Marín et al., 2003; Andrews et al., 2008) and the present analysis did not show differences in the number and positioning of interneurons in the developing cortex and of striatal cells between Slit1−/−/Slit2−/− mice and the corresponding controls. Together, our results strongly suggest that interneurons generated in the MGE of mice lacking Robo1 receptors do not avoid the developing striatum but, instead, migrate through it to reach the cortex.
What steers interneurons clear of the striatum during their migration to the cortex? Previous work by Marín et al. (2001) had suggested that Sema3A and Sema3F, abundantly present in this region, repel neuropilin-expressing cortical interneurons as they migrate through the subpallium. Because a proportion of cortical interneurons do not respect this exclusion zone in Robo1−/− mice and instead migrate through it, we reasoned that these cells are less responsive to the chemorepulsive effects of the semaphorins. Our in vitro studies with MGE cells from Robo1−/− mice and the GN11 cell line transfected with Robo1-DN confirmed our hypothesis. Interestingly, ∼30% of the MGE-derived cells taken from Robo1−/− mice were not responsive to the effect of Sema3A and Sema3F, which corresponded to the observed increase of CB+ cells in the developing striatum of these mice.
Since deletion of Robo1 renders migrating cortical interneurons less sensitive to Sema3A and Sema3F, we speculated that this receptor may potentially interact directly with the class 3 semaphorins or with neuropilin–plexin complexes to regulate their response to these molecules. However, our in vitro experiments eliminated the former possibility, as they showed lack of binding between Robo1 and Sema3A or Sema3F, but demonstrated strong binding between semaphorins and neuropilins.
Previous studies by Marín et al. (2001) had suggested that loss or disruption of neuropilin function perturbs cortical interneuron migration. Specifically, loss-of-function studies, using focal electroporation of Nrp1 DN plasmids in the MGE of wild-type embryos or similar delivery of GFP plasmids in Nrp2−/− mice, revealed an influx of cortical interneurons in the developing striatum. Our experiments showed that loss of Robo1 function significantly reduces the expression of neuropilin and plexin receptors, especially Nrp1 and PlexinA1. The reduction in Nrp1/PlexinA1, which specifically mediate the effects of Sema3A (He and Tessier-Lavigne, 1997), appears to explain why Robo1−/− MGE-derived cells are less sensitive to Sema3A compared with Sema3F, which primarily signals through Nrp2/PlexinA3–4 (Chen et al., 1997). Interestingly, the role of Robo1 in modulating semaphorin responsiveness does not appear to be selectively important for cortical interneurons, as GN11 cells transfected with Robo1-DN were also less responsive to semaphorins. It is tempting to speculate that this mechanism may be relevant for many other Robo1-expressing cell populations. Examples include neurons that send axons across the midline, which rely on the concerted actions of Slit/Robo signaling and class 3 semaphorins for correct guidance (Zou et al., 2000).
Recent evidence suggests that Dlx1 and Dlx2 control the migration of telencephalic interneurons by direct repression of the Nrp2 receptor and, thus, their response to class 3 semaphorins (Le et al., 2007). More recently, Nkx2.1 was also shown to repressively affect neuropilin levels. Specifically, ectopic expression of Nkx2.1 in migrating MGE-derived cells was shown to render them insensitive to Sema3A/Sema3F chemorepulsion as a result of a mild reduction in the expression of Nrp1 and a prominent decrease in the expression of Nrp2 transcripts in these cells, with no effect on the expression of PlexinA3 or PlexinA4 (Nóbrega-Pereira et al., 2008). The fact that we observed reduced levels of Nrp and PlexinA receptors in MGE-derived cells of Robo1−/− mice prompted us to ask whether the absence of Robo1 receptor also disrupts the expression of these transcription factors. In situ hybridization experiments and QPCR failed to show a change in the expression of Lhx6, Lhx8, Nkx2.1, Dlx1, and Dlx2 (data not shown), suggesting that loss of Robo1 function does not affect expression of these genes. Thus, Robo1 does not appear to exert its effects on neuropilin receptor levels at the transcriptional level.
Robo1 is a member of the Ig CAM (cell adhesion molecule) superfamily (Sundaresan et al., 1998). Other members of this family, such as the cell adhesion molecules L1 and NrCAM (neuron–glia-related cell adhesion molecule), are required for the formation and signaling of neuropilin–PlexinA receptor complexes, through which class 3 semaphorins signal (Winberg et al., 1998; Gelfand et al., 2009). Here, we provided novel evidence for a direct interaction between Robo1 and Nrp1 and, as is the case for L1 (Castellani et al., 2000, 2002), Robo1 is required for effective semaphorin signaling. Although we observed an interaction between Robo1 and Nrp1, additional mechanisms may further modulate Robo1–semaphorin signaling. For example, there are shared intracellular molecules involved in both Robo1 and Sema3A/Nrp1 functions, such as the small Rho GTPases (Rho, Rac, Cdc42) that regulate the actin cytoskeleton (Rohm et al., 2000; Zanata et al., 2002; Guan and Rao, 2003; Murray et al., 2010), and it is tempting to speculate that regulation of Rho GTPases might be a point of additional cross talk between Robo1 and class 3 semaphorins. Thus, additional studies are required to elucidate the nature of the Robo1–Nrp1 interaction, and the exact role of Robo1 in semaphorin signaling.
In summary, we have demonstrated quantitatively an influx of cortical interneurons within the developing striatum of Robo1−/−, but not Robo2−/− mice, and shown in vitro that cortical interneurons lacking the Robo1 receptor fail to respond to class 3 semaphorin-induced chemorepulsion. Failure to respond to semaphorin appears to be attributable to a decrease in the levels of Nrp1 and PlexinA1 receptors within these cells. Thus, we have shown for the first time that Robo1 interacts with Nrp1 to modulate semaphorin signaling in the developing forebrain and, as such, plays a crucial role in the directed migration of cortical interneurons through the subpallium.
Footnotes
This work was supported by Wellcome Trust Programme Grants 074549 (J.G.P.) and 089775 (J.G.P., C.R., W.D.A.). L.R.H.-M. was a recipient of a PhD scholarship from the Mexican Council for Science and Technology (Consejo Nacional de Ciencia y Tecnologia). J.-F.C. is a Canada Research Chair and is supported by grants from Canadian Institutes of Health Research, Centre of Excellence in Commercialization and Research of the Montreal Neurological Institute. J.H.C. was supported by a studentship from Le Fonds de la Recherche en Santé de Québec. We thank Drs. Y. Yanagawa and K. Obata for providing the GAD67-GFP transgenic mice, Dr. P. Rabbitts for the ROBO1 knock-out animals, Dr. V. Sundaresan for the Robo constructs, Dr. N. Kessaris for a number of in situ probes, and Prof. F. Murakami for Robo-specific antibodies.
References
- Andrews W, Liapi A, Plachez C, Camurri L, Zhang J, Mori S, Murakami F, Parnavelas JG, Sundaresan V, Richards LJ. Robo1 regulates the development of major axon tracts and interneuron migration in the forebrain. Development. 2006;133:2243–2252. doi: 10.1242/dev.02379. [DOI] [PubMed] [Google Scholar]
- Andrews W, Barber M, Hernandez-Miranda LR, Xian J, Rakic S, Sundaresan V, Rabbitts TH, Pannell R, Rabbitts P, Thompson H, Erskine L, Murakami F, Parnavelas JG. The role of Slit-Robo signaling in the generation, migration and morphological differentiation of cortical interneurons. Dev Biol. 2008;313:648–658. doi: 10.1016/j.ydbio.2007.10.052. [DOI] [PubMed] [Google Scholar]
- Barber M, Di Meglio T, Andrews WD, Hernández-Miranda LR, Murakami F, Chédotal A, Parnavelas JG. The role of Robo3 in the development of cortical interneurons. Cereb Cortex. 2009;19(Suppl 1):i22–i31. doi: 10.1093/cercor/bhp041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camurri L, Mambetisaeva E, Sundaresan V. Rig-1 a new member of Robo family genes exhibits distinct pattern of expression during mouse development. Gene Expr Patterns. 2004;4:99–103. doi: 10.1016/s1567-133x(03)00142-x. [DOI] [PubMed] [Google Scholar]
- Camurri L, Mambetisaeva E, Davies D, Parnavelas J, Sundaresan V, Andrews W. Evidence for the existence of two Robo3 isoforms with divergent biochemical properties. Mol Cell Neurosci. 2005;30:485–493. doi: 10.1016/j.mcn.2005.07.014. [DOI] [PubMed] [Google Scholar]
- Cariboni A, Hickok J, Rakic S, Andrews W, Maggi R, Tischkau S, Parnavelas JG. Neuropilins and their ligands are important in the migration of gonadotropin-releasing hormone neurons. J Neurosci. 2007;27:2387–2395. doi: 10.1523/JNEUROSCI.5075-06.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castellani V, Chédotal A, Schachner M, Faivre-Sarrailh C, Rougon G. Analysis of the L1-deficient mouse phenotype reveals cross-talk between Sema3A and L1 signaling pathways in axonal guidance. Neuron. 2000;27:237–249. doi: 10.1016/s0896-6273(00)00033-7. [DOI] [PubMed] [Google Scholar]
- Castellani V, De Angelis E, Kenwrick S, Rougon G. cis and trans interactions of L1 with neuropilin-1 control axonal responses to semaphorin 3A. EMBO J. 2002;21:6348–6357. doi: 10.1093/emboj/cdf645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Catalano A, Caprari P, Rodilossi S, Betta P, Castellucci M, Casazza A, Tamagnone L, Procopio A. Cross-talk between vascular endothelial growth factor and semaphorin-3A pathway in the regulation of normal and malignant mesothelial cell proliferation. FASEB J. 2004;18:358–360. doi: 10.1096/fj.03-0513fje. [DOI] [PubMed] [Google Scholar]
- Cavanagh JF, Mione MC, Pappas IS, Parnavelas JG. Basic fibroblast growth factor prolongs the proliferation of rat cortical progenitor cells in vitro without altering their cell cycle parameters. Cereb Cortex. 1997;7:293–302. doi: 10.1093/cercor/7.4.293. [DOI] [PubMed] [Google Scholar]
- Chen H, Chédotal A, He Z, Goodman CS, Tessier-Lavigne M. Neuropilin-2, a novel member of the neuropilin family, is a high affinity receptor for the semaphorins Sema E and Sema IV but not Sema III. Neuron. 1997;19:547–559. doi: 10.1016/s0896-6273(00)80371-2. [DOI] [PubMed] [Google Scholar]
- Cho JH, Lépine M, Andrews W, Parnavelas J, Cloutier JF. Requirement for Slit-1 and Robo-2 in zonal segregation of olfactory sensory neuron axons in the main olfactory bulb. J Neurosci. 2007;27:9094–9104. doi: 10.1523/JNEUROSCI.2217-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corbin JG, Nery S, Fishell G. Telencephalic cells take a tangent: non-radial migration in the mammalian forebrain. Nat Neurosci. 2001;4(Suppl):1177–1182. doi: 10.1038/nn749. [DOI] [PubMed] [Google Scholar]
- Faux C, Rakic S, Andrews W, Yanagawa Y, Obata K, Parnavelas JG. Differential gene expression in migrating cortical interneurons during mouse forebrain development. J Comp Neurol. 2010;518:1232–1248. doi: 10.1002/cne.22271. [DOI] [PubMed] [Google Scholar]
- Feldblum S, Erlander MG, Tobin AJ. Different distributions of GAD65 and GAD67 mRNAs suggest that the two glutamate decarboxylases play distinctive functional roles. J Neurosci Res. 1993;34:689–706. doi: 10.1002/jnr.490340612. [DOI] [PubMed] [Google Scholar]
- Fentress JC, Stanfield BB, Cowan WM. Observation on the development of the striatum in mice and rats. Anat Embryol. 1981;163:275–298. doi: 10.1007/BF00315705. [DOI] [PubMed] [Google Scholar]
- Fragkouli A, van Wijk NV, Lopes R, Kessaris N, Pachnis V. LIM homeodomain transcription factor-dependent specification of bipotential MGE progenitors into cholinergic and GABAergic striatal interneurons. Development. 2009;136:3841–3851. doi: 10.1242/dev.038083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuh G, Garcia KC, de Vos AM. The interaction of neuropilin-1 with vascular endothelial growth factor and its receptor flt-1. J Biol Chem. 2000;275:26690–26695. doi: 10.1074/jbc.M003955200. [DOI] [PubMed] [Google Scholar]
- Gelfand MV, Hong S, Gu C. Guidance from above: common cues direct distinct signaling outcomes in vascular and neuronal patterning. Trends Cell Biol. 2009;19:99–110. doi: 10.1016/j.tcb.2009.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerfen CR. The neostriatal mosaic: multiple levels of compartmental organization. Trends Neurosci. 1992;15:133–139. doi: 10.1016/0166-2236(92)90355-c. [DOI] [PubMed] [Google Scholar]
- Giraudo E, Primo L, Audero E, Gerber HP, Koolwijk P, Soker S, Klagsbrun M, Ferrara N, Bussolino F. Tumor necrosis factor-alpha regulates expression of vascular endothelial growth factor receptor-2 and of its co-receptor neuropilin-1 in human vascular endothelial cells. J Biol Chem. 1998;273:22128–22135. doi: 10.1074/jbc.273.34.22128. [DOI] [PubMed] [Google Scholar]
- Gluzman-Poltorak Z, Cohen T, Shibuya M, Neufeld G. Vascular endothelial growth factor receptor-1 and neuropilin-2 form complexes. J Biol Chem. 2001;276:18688–18694. doi: 10.1074/jbc.M006909200. [DOI] [PubMed] [Google Scholar]
- Greif KF, Tillakaratne NJ, Erlander MG, Feldblum S, Tobin AJ. Transient increase in expression of a glutamate decarboxylase (GAD) mRNA during the postnatal development of the rat striatum. Dev Biol. 1992;153:158–164. doi: 10.1016/0012-1606(92)90100-u. [DOI] [PubMed] [Google Scholar]
- Guan KL, Rao Y. Signaling mechanisms mediating neuronal responses to guidance cues. Nat Rev Neurosci. 2003;4:941–956. doi: 10.1038/nrn1254. [DOI] [PubMed] [Google Scholar]
- Hammond R, Vivancos V, Naeem A, Chilton J, Mambetisaeva E, Andrews W, Sundaresan V, Guthrie S. Slit-mediated repulsion is a key regulator of motor axon pathfinding in the hindbrain. Development. 2005;132:4483–4495. doi: 10.1242/dev.02038. [DOI] [PubMed] [Google Scholar]
- He Z, Tessier-Lavigne M. Neuropilin is a receptor for the axonal chemorepellent Semaphorin III. Cell. 1997;90:739–751. doi: 10.1016/s0092-8674(00)80534-6. [DOI] [PubMed] [Google Scholar]
- Hernández-Miranda LR, Parnavelas JG, Chiara F. Molecules and mechanisms involved in the generation and migration of cortical interneurons. ASN Neuro. 2010;2:e00031. doi: 10.1042/AN20090053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hivert B, Liu Z, Chuang CY, Doherty P, Sundaresan V. Robo1 and Robo2 are homophilic binding molecules that promote axonal growth. Mol Cell Neurosci. 2002;21:534–545. doi: 10.1006/mcne.2002.1193. [DOI] [PubMed] [Google Scholar]
- Hoshi T, Sasano H, Kato K, Yabuki N, Ohara S, Konno R, Asaki S, Toyota T, Tateno H, Nagura H. Immunohistochemistry of Caspase3/CPP32 in human stomach and its correlation with cell proliferation and apoptosis. Anticancer Res. 1998;8:4347–4353. [PubMed] [Google Scholar]
- Kawaguchi Y, Wilson CJ, Augood SJ, Emson PC. Striatal interneurons: chemical, physiological, and morphological characterization. Trends Neurosci. 1995;18:527–535. doi: 10.1016/0166-2236(95)98374-8. [DOI] [PubMed] [Google Scholar]
- Lavdas AA, Grigoriou M, Pachnis V, Parnavelas JG. The medial ganglionic eminence gives rise to a population of early neurons in the developing cerebral cortex. J Neurosci. 1999;19:7881–7888. doi: 10.1523/JNEUROSCI.19-18-07881.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Le TN, Du G, Fonseca M, Zhou QP, Wigle JT, Eisenstat DD. Dlx homeobox genes promote cortical interneuron migration from the basal forebrain by direct repression of the semaphorin receptor neuropilin-2. J Biol Chem. 2007;282:19071–19081. doi: 10.1074/jbc.M607486200. [DOI] [PubMed] [Google Scholar]
- Liu FC, Graybiel AM. Heterogeneous development of calbindin-D28K expression in the striatal matrix. J Comp Neurol. 1992;320:304–322. doi: 10.1002/cne.903200304. [DOI] [PubMed] [Google Scholar]
- Liu Z, Patel K, Schmidt H, Andrews W, Pini A, Sundaresan V. Extracellular Ig domains 1 and 2 of Robo are important for ligand (Slit) binding. Mol Cell Neurosci. 2004;26:232–240. doi: 10.1016/j.mcn.2004.01.002. [DOI] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using realtime quantitative PCR and the 2(−ΔΔC(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- Marillat V, Cases O, Nguyen-Ba-Charvet KT, Tessier-Lavigne M, Sotelo C, Chédotal A. Spatiotemporal expression patterns of slit and robo genes in the rat brain. J Comp Neurol. 2002;442:130–155. doi: 10.1002/cne.10068. [DOI] [PubMed] [Google Scholar]
- Marín O, Rubenstein JL. Cell migration in the forebrain. Annu Rev Neurosci. 2003;26:441–483. doi: 10.1146/annurev.neuro.26.041002.131058. [DOI] [PubMed] [Google Scholar]
- Marín O, Yaron A, Bagri A, Tessier-Lavigne M, Rubenstein JL. Sorting of striatal and cortical interneurons regulated by semaphorin/neuropilin interactions. Science. 2001;293:872–875. doi: 10.1126/science.1061891. [DOI] [PubMed] [Google Scholar]
- Marín O, Plump AS, Flames N, Sánchez-Camacho C, Tessier-Lavigne M, Rubenstein JL. Directional guidance of interneuron migration to the cerebral cortex relies on subcortical Slit1/2-independent repulsion and cortical attraction. Development. 2003;130:1889–1901. doi: 10.1242/dev.00417. [DOI] [PubMed] [Google Scholar]
- Mercugliano M, Soghomonian JJ, Qin Y, Nguyen HQ, Feldblum S, Erlander MG, Tobin AJ, Chesselet MF. Comparative distribution of messenger RNAs encoding glutamic acid decarboxylases (Mr 65,000 and Mr 67,000) in the basal ganglia of the rat. J Comp Neurol. 1992;318:245–254. doi: 10.1002/cne.903180302. [DOI] [PubMed] [Google Scholar]
- Métin C, Baudoin JP, Rakić S, Parnavelas JG. Cell and molecular mechanisms involved in the migration of cortical interneurons. Eur J Neurosci. 2006;23:894–900. doi: 10.1111/j.1460-9568.2006.04630.x. [DOI] [PubMed] [Google Scholar]
- Murray A, Naeem A, Barnes SH, Drescher U, Guthrie S. Slit and Netrin-1 guide cranial motor axon pathfinding via Rho-kinase, myosin light chain kinase and myosin II. Neural Dev. 2010;5:16. doi: 10.1186/1749-8104-5-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nóbrega-Pereira S, Kessaris N, Du T, Kimura S, Anderson SA, Marín O. Postmitotic Nkx2–1 controls the migration of telencephalic interneurons by direct repression of guidance receptors. Neuron. 2008;59:733–745. doi: 10.1016/j.neuron.2008.07.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ouimet CC, Langley-Gullion KC, Greengard P. Quantitative immunocytochemistry of DARPP-32-expressing neurons in the rat caudatoputamen. Brain Res. 1998;808:8–12. doi: 10.1016/s0006-8993(98)00724-0. [DOI] [PubMed] [Google Scholar]
- Patel K, Nash JA, Itoh A, Liu Z, Sundaresan V, Pini A. Slit proteins are not dominant chemorepellents for olfactory tract and spinal motor axons. Development. 2001;128:5031–5037. doi: 10.1242/dev.128.24.5031. [DOI] [PubMed] [Google Scholar]
- Plump AS, Erskine L, Sabatier C, Brose K, Epstein CJ, Goodman CS, Mason CA, Tessier-Lavigne M. Slit1 and Slit2 cooperate to prevent premature midline crossing of retinal axons in the mouse visual system. Neuron. 2002;33:219–232. doi: 10.1016/s0896-6273(01)00586-4. [DOI] [PubMed] [Google Scholar]
- Prince JE, Cho JH, Dumontier E, Andrews W, Cutforth T, Tessier-Lavigne M, Parnavelas J, Cloutier JF. Robo-2 controls the segregation of a portion of basal vomeronasal sensory neuron axons to the posterior region of the accessory olfactory bulb. J Neurosci. 2009;29:14211–14222. doi: 10.1523/JNEUROSCI.3948-09.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radovick S, Wray S, Lee E, Nicols DK, Nakayama Y, Weintraub BD, Westphal H, Cutler GB, Jr, Wondisford FE. Migratory arrest of gonadotropin-releasing hormone neurons in transgenic mice. Proc Natl Acad Sci U S A. 1991;88:3402–3406. doi: 10.1073/pnas.88.8.3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rohm B, Rahim B, Kleiber B, Hovatta I, Püschel AW. The semaphorin 3A receptor may directly regulate the activity of small GTPases. FEBS Lett. 2000;486:68–72. doi: 10.1016/s0014-5793(00)02240-7. [DOI] [PubMed] [Google Scholar]
- Sholl DA. Dendritic organization in the neurons of the visual and motor cortices of the cat. J Anat. 1953;87:387–406. [PMC free article] [PubMed] [Google Scholar]
- Sundaresan V, Roberts I, Bateman A, Bankier A, Sheppard M, Hobbs C, Xiong J, Minna J, Latif F, Lerman M, Rabbitts P. The DUTT1 gene, a novel NCAM family member is expressed in developing murine neural tissues and has an unusually broad pattern of expression. Mol Cell Neurosci. 1998;11:29–35. doi: 10.1006/mcne.1998.0672. [DOI] [PubMed] [Google Scholar]
- Takahashi K, Liu FC, Hirokawa K, Takahashi H. Expression of Foxp2, a gene involved in speech and language, in the developing and adult striatum. J Neurosci Res. 2003;73:61–72. doi: 10.1002/jnr.10638. [DOI] [PubMed] [Google Scholar]
- Tamada A, Kumada T, Zhu Y, Matsumoto T, Hatanaka Y, Muguruma K, Chen Z, Tanabe Y, Torigoe M, Yamauchi K, Oyama H, Nishida K, Murakami F. Crucial roles of Robo proteins in midline crossing of cerebellofugal axons and lack of their up-regulation after midline crossing. Neural Dev. 2008;3:29. doi: 10.1186/1749-8104-3-29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamamaki N, Yanagawa Y, Tomioka R, Miyazaki J, Obata K, Kaneko T. Green fluorescent protein expression and co-localisation with calretinin, parvalbumin, and somatostatin in the GAD67-GFP knock-in mouse. J Comp Neurol. 2003;467:60–79. doi: 10.1002/cne.10905. [DOI] [PubMed] [Google Scholar]
- Vieira JM, Schwarz Q, Ruhrberg C. Selective requirements for NRP1 ligands during neurovascular patterning. Development. 2007;134:1833–1843. doi: 10.1242/dev.002402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wichterle H, Turnbull DH, Nery S, Fishell G, Alvarez-Buylla A. In utero fate mapping reveals distinct migratory pathways and fates of neurons born in the mammalian basal forebrain. Development. 2001;128:3759–3771. doi: 10.1242/dev.128.19.3759. [DOI] [PubMed] [Google Scholar]
- Winberg ML, Noordermeer JN, Tamagnone L, Comoglio PM, Spriggs MK, Tessier-Lavigne M, Goodman CS. Plexin A is a neuronal semaphorin receptor that controls axon guidance. Cell. 1998;95:903–916. doi: 10.1016/s0092-8674(00)81715-8. [DOI] [PubMed] [Google Scholar]
- Zanata SM, Hovatta I, Rohm B, Püschel AW. Antagonistic effects of Rnd1 and RhoD GTPases regulate receptor activity in Semaphorin 3A-induced cytoskeletal collapse. J Neurosci. 2002;22:471–477. doi: 10.1523/JNEUROSCI.22-02-00471.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Y, Marín O, Hermesz E, Powell A, Flames N, Palkovits M, Rubenstein JL, Westphal H. The LIM-homeobox gene Lhx8 is required for the development of many cholinergic neurons in the mouse forebrain. Proc Natl Acad Sci U S A. 2003;100:9005–9010. doi: 10.1073/pnas.1537759100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zou Y, Stoeckli E, Chen H, Tessier-Lavigne M. Squeezing axons out of the gray matter: a role for slit and semaphorin proteins from midline and ventral spinal cord. Cell. 2000;102:363–375. doi: 10.1016/s0092-8674(00)00041-6. [DOI] [PubMed] [Google Scholar]