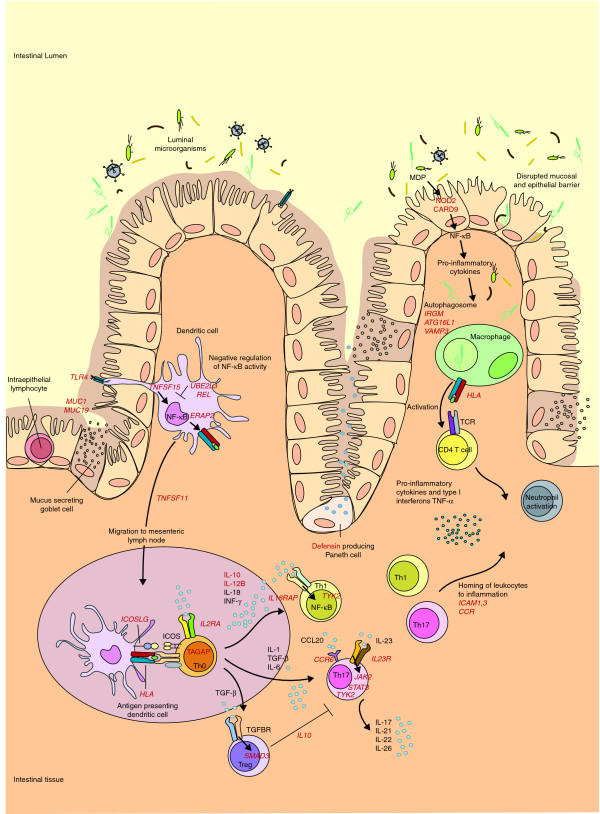
Figure 1.
Schematic representation of the genes and pathways associated with Crohn's disease pathogenesis. The ongoing inflammatory response in the gastrointestinal tract in patients with Crohn's disease (CD) is thought to be caused by an aberrant immune response to commensal microflora in the gut. In patients with CD, defects in first defense mechanisms (that is, disrupted epithelial and mucosal barrier) contribute to increased bacterial penetration (MUC1 and MUC19). Genes involved in pattern recognition (NOD2, TLR4 and CARD9) suggest an increased response of antigen-presenting cells to commensal microbes. Consequently, the NF-кB cascade is activated (TNFSF15), leading to production of pro-inflammatory cytokines. Association of REL and UBE2L3 suggest an impaired NF-кB negative feedback. Antigen-presenting cells migrate to Peyer's patches (intestinal mesenteric lymph nodes) (TNFSF11) to present antigens and stimulate T-cell proliferation (IL2RA and TAGAP) and differentiation. T cells of patients with CD, in turn, respond more intensely. Th0 cells are stimulated to differentiate into T-cell subtypes regulated by a variety of the produced cytokines and their receptors. Th17 cells are involved in many immune-related diseases, and they are activated through IL-23R, which, in turn, activates the JAK-STAT-TYK (Janus kinase- signal transducer and activator of transcription-tyrosine kinase) pathway that enhances pro-inflammatory cytokine production (JAK2, STAT3 and TYK2). Th1 and Th17 cells are pro-inflammatory, whereas Treg cells downregulate the immune response. Another major contribution to CD pathogenesis comes from autophagy. In autophagosomes, intracellular components, including phagocytosed microbes, are degraded, after which their antigens are presented to CD4+ cells. Autophagy is at least partly regulated by the CD risk genes ATG16L1, IRGM and VAMP3. The activation of CD4+ cells leads to the production of pro-inflammatory cytokines and the maintenance of the inflammation. All the displayed processes could finally lead to homing of leukocytes to inflammation sites (ICAM1,3, CCR cluster), and neutrophil recruitment. Consequently, chronic inflammation, ulceration and deeper microbial penetrance occur. The known associated genes are shown in red. Table 1 summarizes the associated loci shown here. CCL20, chemokine (C-C motif) ligand 20; ICOS, inducible T-cell co-stimulator; MDP, muramyl dipeptide; NF, nuclear factor; TCR, T-cell receptor; TGF, transforming growth factor; TGFBR, TGF β receptor; Th, T helper cell; TNF, tumor necrosis factor; Treg, regulatory T cell.