Abstract
Recently, we reported a database (NCAD, Non-Coded Amino acids Database; http://recerca.upc.edu/imem/index.htm) that was built to compile information about the intrinsic conformational preferences of non-proteinogenic residues determined by quantum mechanical calculations, as well as bibliographic information about their synthesis, physical and spectroscopic characterization, the experimentally-established conformational propensities, and applications (J. Phys. Chem. B 2010, 114, 7413). The database initially contained the information available for α-tetrasubstituted α-amino acids. In this work, we extend NCAD to three families of compounds, which can be used to engineer peptides and proteins incorporating modifications at the –NHCO– peptide bond. Such families are: N-substituted α-amino acids, thio-α-amino acids, and diamines and diacids used to build retropeptides. The conformational preferences of these compounds have been analyzed and described based on the information captured in the database. In addition, we provide an example of the utility of the database and of the compounds it compiles in protein and peptide engineering. Specifically, the symmetry of a sequence engineered to stabilize the 310-helix with respect to the α-helix has been broken without perturbing significantly the secondary structure through targeted replacements using the information contained in the database.
Introduction
Apart from the genetically coded α-amino acids found in proteins, there is a large number of amino acids grouped under the common name of non-proteinogenic or non-coded amino acids (nc-aa). Some of these nc-aa are present in nature, occurring in non-ribosomally produced peptides (e.g. antibiotics); however, most are synthesized by chemists. Among the latter, those used for protein engineering1–6 and nanobiology7–12 are particularly interesting, and their applications extend to many fields such as pharmacology (drug design),13,14 biotechnology (biosensors),15,16 and nanomedicine (systems for drug delivery and diagnosis through imaging).17,18
In spite of the great significance and potential utility of nc-aa in modern biology, biomaterials engineering, and medicine, structural information is available only for a limited number of these. In the last two decades, the intrinsic conformational preferences of several tens of nc-aa have been investigated through sophisticated quantum mechanical calculations at the ab initio or Density Functional Theory (DFT) levels. Specifically, these reliable theoretical methods have been used to sample the potential energy hypersurface of small peptide systems incorporating such nc-aa in the absence of external forces. At the same time, crystallographic and spectroscopic structural data on peptide sequences incorporating nc-aa are becoming available for an increasing number of cases. Comparison of such experimental information with that derived from high-level theoretical calculations helps establish the influence of packing forces, solvent effects, and the chemical environment on the conformational propensities of nc-aa.
In practice, the use of nc-aa with well-characterized conformational properties is frequently limited because the relevant information is highly dispersed. Quantum mechanical calculations describing the intrinsic conformational preferences of these compounds are typically reported in physical chemistry journals, while their synthesis and crystalline structures developed by organic and peptide chemists are published in journals specialized in these fields. In contrast, many applications of nc-aa are tested by researchers working on protein engineering, medicine, or materials science. In order to integrate the dispersed contributions about nc-aa, which is expected to facilitate their practical application in many research fields, a database of nc-aa was recently built.19
NCAD (Non-Coded Amino acids Database) is a database conceived and created to identify the nc-aa that are compatible with a given structural motif, which is the key requirement for the application of these compounds in the life and materials sciences. The database, which is publicly accessible through the web page, http://recerca.upc.edu/imem/index.htm, integrates structural and energetic descriptors of those nc-aa whose intrinsic conformational properties have been previously studied using ab initio or DFT quantum mechanical calculations and reported in the literature. In our recent work19 we described the technical aspects of the database (structure, contents, informatics features and relationships among the different descriptors) and presented the first family of nc-aa integrated into it. Specifically, we discussed the relevance of α-tetrasubstituted α-amino acids in the design of peptides with well-defined conformational properties and described the 29 α-tetrasubstituted α-amino acids whose intrinsic conformational propensities have been determined to date using ab initio or DFT quantum mechanical methods according to a literature search.19
In this work, we extend NCAD to three new families of compounds: (i) N-substituted α-amino acids; (ii) thio-α-amino acids; and (iii) diamines and diacids surrogates of α-amino acids that are used to build pseudopeptides containing one (or more) retropeptide link(s), that is, a peptide bond in reversed direction to that of its immediate neighbors. The members of three such series can be used to engineer peptides and proteins incorporating modifications at the conventional –NHCO– peptide bond. Below, the work is organized as follows. First, a brief description of the contents and structure of NCAD is provided. After this, the three families of amino acids integrated in the database are presented. Next, the utility of the database is illustrated by engineering helical models using the information of the diamines and diacids included in NCAD. Finally, the conclusions of the work are outlined.
NCAD: Integrating the intrinsic conformational preferences of non-coded amino acids
NCAD integrates information about nc-aa whose intrinsic conformational preferences have been studied by applying ab initio or DFT quantum mechanical calculations to the smallest peptide systems which contains them (that is, the nc-aa bearing the amino and carboxylic acid termini as an amide, typically capped with the acetyl and methylamide groups). The information contained in NCAD is divided into three categories:
Minimum energy conformations. All the minima found for each compound contained in the database are described in detail. Dihedral angles, atomic coordinates and relative energies obtained at the highest level of theory among those used to study each nc-aa are stored through inter-connected descriptors. The backbone conformations of the different energy minima contained in NCAD have been identified using the nomenclature proposed by Perzcel et al.,20 which distinguish nine different conformations in the potential energy surface E=E(φ,ψ) of α-amino acids (i.e. γD,δD, αD, εD, βDL, εL, αL, δL, and γL). This information is essential to check the compatibility of the different energy minima with the secondary structural motifs typically found in peptides and proteins.
Bibliographic information describing the available experimental information. Specifically, the following items are reported for each amino acid: (i) synthesis and characterization of the free form or with the amino group adequately protected for use in peptide synthesis; and (ii) spectroscopic and structural characterization when incorporated into peptides, the atomic coordinates extracted from the X-ray crystal structures being also included in selected cases.
Applications. The most relevant publications and patents describing the use of nc-aa in related fields along with the biological properties and relevance in materials science.
The labels identifying the information stored in the database are connected through a relational scheme. This facilitates the search process, which can be performed using a single or several descriptors. NCAD is accessed through a user-friendly interface, which integrates a simple window system, thus facilitating the use by non-experts in informatics. The interface is divided into six folders, the first one being used to define the criteria applied to the search process. The available search criteria are: (i) the chemical nature of the side chain; (ii) the structural feature which distinguishes a particular family of nc-aa; (iii) the molecular formula; (iv) the type of backbone conformation according to the nomenclature established by Perczel et al.;20,21 (v) the values of the (φ,ψ) dihedral angles, with the tolerance being specified by the user; (vi) the name of the researcher who performed the theoretical or experimental investigation; (vii) the experimental data reported in the literature relating to the synthesis and characterization; and (viii) the applications described in the literature. The amino acids fulfilling all the search criteria are displayed in a list. Once the user selects a given amino acid from this list, the information stored in the database is presented in the remaining five folders of the interface. The specific information provided in each folder was detailed in our previous work.19
Amino-acid surrogates leading to peptides with modified peptide bonds
In our previous work,19 we provided evidence for the usefulness of the α-tetrasubstituted α-amino acids stored in NCAD in the control of peptide conformation. Another means of modulating the conformational properties of peptides can come from the modification of the peptide bond itself. This change should perturb the intramolecular hydrogen bonding network, the electronic distribution, and the steric hindrance, thus affecting the conformational preferences of the peptide chain.
N-substituted α-amino acids
In N-substituted amino acids, the hydrogen atom attached to nitrogen is replaced by a functional group. The substituents most frequently used are: alkyl (N-alkylation), hydroxyl (N-hydroxylation) or amino (N-amination). To the best of our knowledge, reliable theoretical studies at the ab initio or DFT quantum mechanical levels on the intrinsic conformational preferences of N-substituted amino acids have been performed only for N-methylalanine,22 N-hydroxyalanine23 and N-aminoglycine24 (Table 1).
Table 1.
N-substituted α-amino acids and thio-α-amino acids stored in NCAD. The systematic name (and the common name), the chemical structure, and the reference reporting the quantum mechanical study are given for each amino acid.
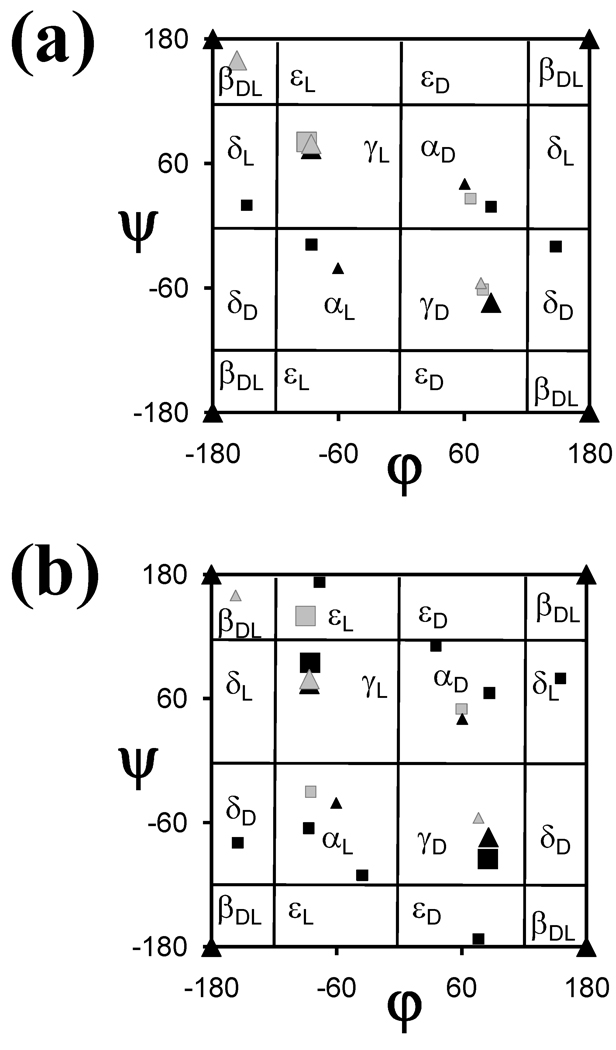
N-alkylation (in particular, N-methylation) has been largely used to change the pharmacological properties of peptides. Remarkable biological and pharmacological profiles have been observed for many N-methylated peptides.25 The conformational preferences of N-methylated amino acids are dictated by increased steric hindrance and lack of hydrogen-bond donor ability associated with the replacement of the hydrogen atom by a methyl group.26,27 Figure 1a presents the Ramachandran map corresponding to the (φ,ψ) backbone dihedral angles of all the energy minima reported for the N-acetyl-N’-methylamide derivatives of the N-methylated α-amino acid included in the database, namely N-methylalanine.22 The minima derived from ab initio quantum mechanical calculations for the analogous derivatives of glycine and alanine28 (MeCO-Gly-NHMe and MeCO-Ala-NHMe, respectively) are also shown in Figure 1a as a reference for proteinogenic amino acids. As it can be seen, the effect of the replacement of the amide hydrogen of Ala by a methyl group on the low-energy minimum (βDL) is very significant, with the minimum being annihilated. The disappearance of this minimum should be attributed to the repulsive interactions produced by the confrontation of the N-methyl group and the oxygen atom of C=O moiety.
Figure 1.
Ramachandran maps showing all the minimum energy conformations predicted by quantum mechanical calculations for the N-acetyl-N’-methylamide derivatives of the N-modified α-amino acids included in NCAD (see Table 1): (a) N-methylalanine (gray squares); and (b) N-aminoglycine (black squares) and N-hydroxyalanine (gray squares). The minimum energy conformations of Gly (black triangles) and Ala (grey triangles) have been included for comparison. Minima with relative energies lower and higher than 2.0 kcal/mol are represented by large and small symbols, respectively.
The N-hydroxylation and N-amination of peptides have been used to impart conformational rigidity through the formation of intramolecular hydrogen bonds with neighboring C=O groups.29,30 Besides its hydrogen-bonding abilities, the N-hydroxyl group has been proven able to chelate metal ions, which may be of use for the design of peptides which can bind specifically to proteins containing metals in the active site.29 The Ramachandran map in Figure 1b shows all the energy minima reported for the N-acetyl-N’-methylamide derivatives of N-hydroxyalanine23 and N-aminoglycine,24 which are compared to those of Ala and Gly,28 respectively. As can be seen, replacement of the amide hydrogen by a hydroxyl or amino group entails drastic conformational changes. The most remarkable ones can be summarized as follows: (i) the lowest energy minimum of Ala (γL) evolves towards εL upon N-hydroxylation; (ii) the extended βDL conformation of Ala is not a minimum in N-hydroxyalanine; (iii) the steric interactions induced by the N-amino group produces a destabilization of the βDL conformation in N-aminoglycine; and (iv) N-amination of Gly leads to the characterization of new local minima located at the εL, εD, δD, and δL regions of the Ramachandran map.
Thio-α-amino acids
Within the surrogates used to replace the amide bond of peptides, the thioamide functionality, in which the carbonyl oxygen is replaced by a sulfur atom, is amongst the most synthetically accessible.31–33 This modification not only affects the proteolytic stability and bioactivity of peptides;34–36 it also produces significant changes in the secondary structure relative to the parent peptides.37–38 Both amides and thioamides are planar and exhibit high energy barriers for rotation about the C–N bond.40,41 However, the length of the C=S bond in thioamides (~0.45 Å larger than the C=O linkage in amides) and the distinct stereoelectronic features of the sulfur atom (its van der Waals radius is 32% larger than that of oxygen and sulfur is less electronegative) affect the hydrogen-bonding abilities of the thioamide bond and translate into a reduced proton-acceptor character.42
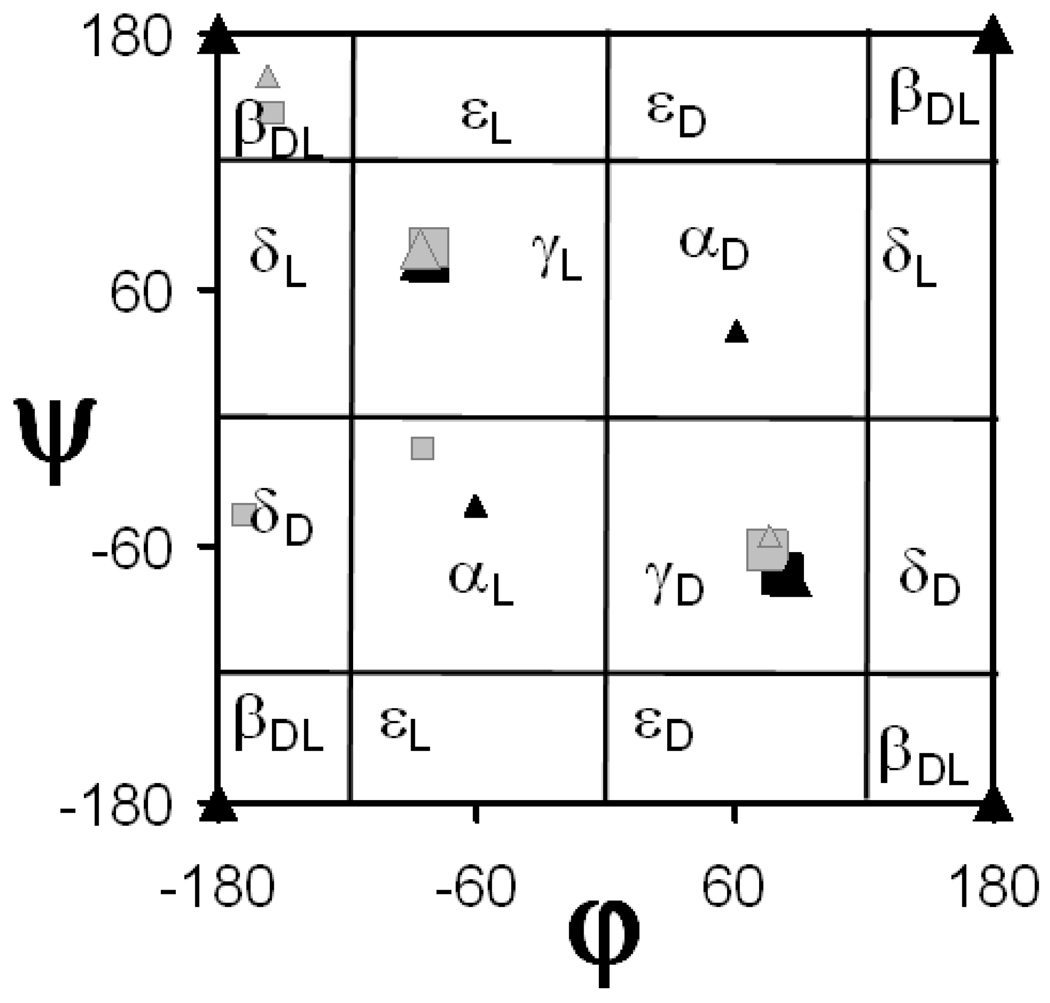
The influence of this amide-thioamide replacement on the intrinsic conformational properties has been studied only for Gly and Ala (Table 1).43 It has been found that the sulfur atom reduces drastically the conformational flexibility of thioglycine when compared to the coded amino acid. Thus, the only minimum energy conformation identified for thioglycine,43 which is two-fold degenerate, corresponds to a γ-turn motif (Figure 2). Therefore, the minima found for Gly28 at the βDL and α regions of the Ramachandran map are annihilated upon replacement of oxygen by sulfur. Figure 2 shows a Ramachandran map with the five minima found for thioalanine.43 The three minima detected for Ala28 were also identified as minimum energy conformations for thioalanine and exhibit similar relative energies. Thus, γL is the most stable conformation for the two amino acids while the relative energies of the γD and βDL structures differ in 0.6 and 1.4 kcal/mol, respectively. The relative energies of the two additional minima characterized for thioalanine, αL and δD, are above 4 kcal/mol. Accordingly, the changes produced in Ala by the presence of the sulfur atom are considerably less important than those observed for Gly.
Figure 2.
Ramachandran maps showing all the minimum energy conformations predicted by quantum mechanical calculations for the N-acetyl-N’-methylamide derivatives of the thio-α-amino acids included in NCAD (see Table 1): thioglycine (black squares) and thioalanine (gray squares). The minimum energy conformations of Gly (black triangles) and Ala (grey triangles) have been included for comparison. Minima with relative energies lower and higher than 2.0 kcal/mol are represented by large and small symbols, respectively.
Diamides and diacids used to build retropeptide bonds
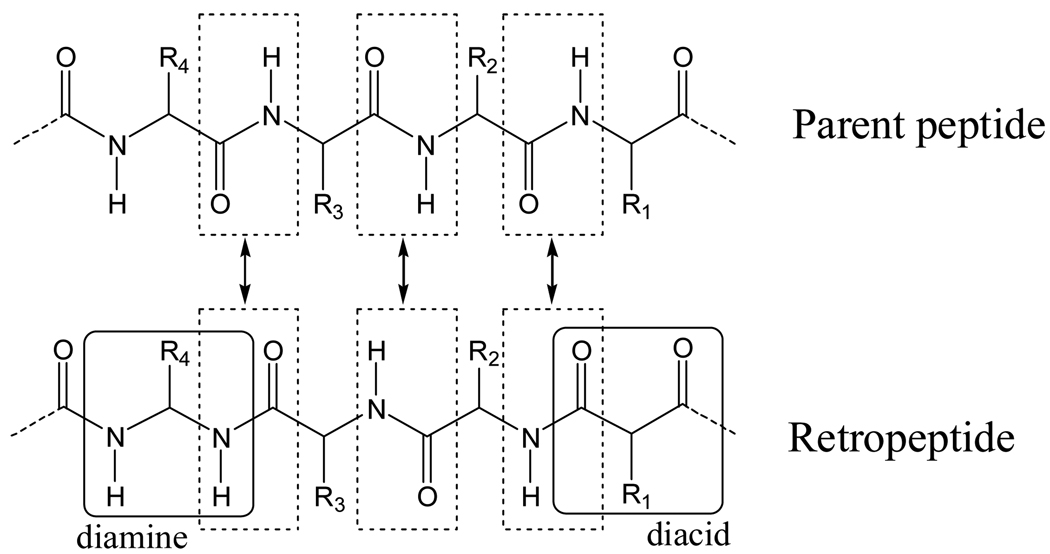
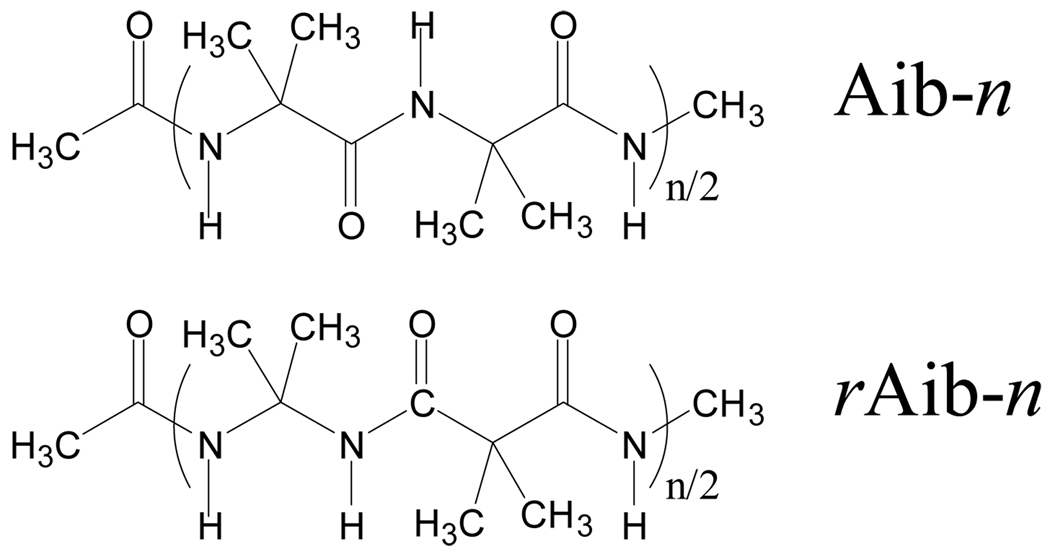
In the last decades, a great deal of effort has been devoted to the chemistry of the so-called retropeptides or partially modified retropeptides,44–50 in which the direction of all or some peptide bonds is reversed, i.e. NH–CO instead of CO–NH (Scheme 1). When the stereochemistry of one or more amino acids of the reversed segment is inverted, the resulting pseudopeptide is termed retroinverso. 1,1-Dicarboxylic acids (malonic acid derivatives) are used to initiate the retropeptide sequence, while the direction of the peptide bond can be restored by incorporating an 1,1-diamine unit (Scheme 1).
Scheme 1.
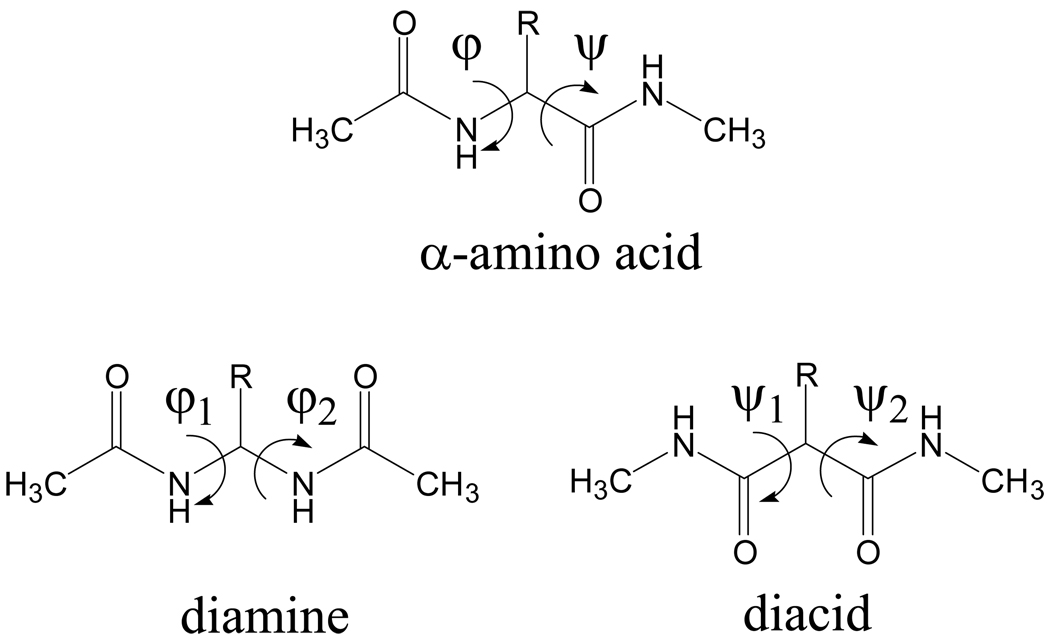
Reversal of the peptide bond has been used for designing bioactive pseudopeptides,50–54 for engineering and mirroring protein secondary structural motifs,55–58 and for the construction of synthetic (bio)polymers mimicking the structures found in fibrous proteins.59,60 NCAD includes the conformational preferences of 5 diacids and 4 diamines (Table 2) that are surrogates of the following α-amino acids: Gly,60,61 Ala,60 Val,62 α-aminoisobutyric acid (Aib), and dehydroalanine63 (ΔAla). Considering that the φ,ψ dihedral angles in α-amino acids define the rotation about the Cα–N and Cα–C(O) bonds, respectively, the conformation of the diacids is given by ψ1,ψ2 and that of the diamines by φ1,φ2. Scheme 2 compares the backbone dihedral angles of the N-acetyl-N’-methylamide derivatives, N,N’-dimethylamide derivatives, and N,N’-diacetyl derivatives of the α-amino acids, diamines and diacids, respectively.
Table 2.
Diacids and diamines analogues of α-amino acids stored in NCAD. The systematic name (and the abbreviated name used in this work), the chemical structure, and the reference reporting the quantum mechanical study are given for each amino acid.
| Systematic name (abbreviated name) |
Structure | Ref. |
|---|---|---|
| malonic acid (dc-Gly) |  |
61 |
| methylmalonic acid (dc-Ala) |  |
This work |
| dimethylmalonic acid (dc-Aib) |  |
This work |
| isopropylmalonic acid (dc-Val) |  |
62 |
| methylenemalonic acid (dc-ΔAla) |  |
64 |
| methylenediamine (dm-Gly) |  |
60 |
| 1,1-ethylenediamine (dm-Ala) |  |
60 |
| 2,2-propylenediamine(dm-Aib) |  |
This work |
| 1,1-isobutylenediamine(dm-Val) |  |
62 |
Scheme 2.
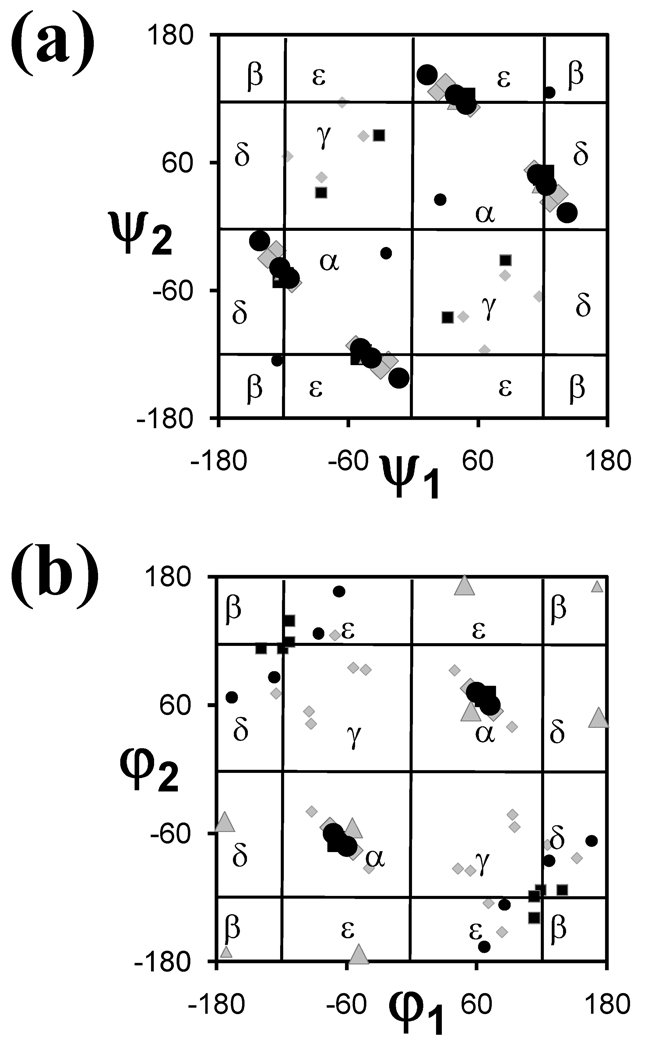
The Ramachandran map in Figure 3a shows the location of the minimum energy conformations found for the N,N’-dimethylamide derivatives of the diacid surrogates of Gly, Ala, Aib, Val, and ΔAla, hereafter denoted dc-Gly,61 dc-Ala, dc-Aib, dc-Val,62 and dc-ΔAla,63 respectively (Table 2). The minimum energy conformations of the dc-Ala and dc-Aib derivatives were not reported in the literature and have been calculated in this work at the B3LYP/6-31G(d) level (the dihedral angles and relative energies are given in the Supporting Information). The resulting conformations have been used for engineering a 310-helical retropeptide (see below).
Figure 3.
Ramachandran maps showing all the minimum energy conformations predicted by quantum mechanical calculations for (a) the N,N’-dimethylamide derivatives of the diacids and (b) the N,N’-diacetyl derivatives of the diamines surrogates of α-amino acids included in NCAD (see Table 2). (a) Diacids: dc-Gly (black squares), dc-Ala (black circles), dc-Aib (gray triangles), dc-Val (gray diamonds) and dc-ΔAla (empty circles); (b) Diamines: dm-Gly (black squares), dm-Ala (black circles), dm-Aib (gray triangles) and dm-Val (gray diamonds). The nomenclature used for the different diacids and diamines is discussed in the text. Minima with relative energies lower and higher than 2.0 kcal/mol are represented by large and small symbols, respectively.
Due to molecular symmetry, structures characterized by dihedral angles ψ1,ψ2; -ψ1,-ψ2; ψ2,ψ1; and -ψ2,-ψ1 are energetically degenerate (Figure 3a). The lowest energy conformation was found at ψ1,ψ2≈ 45°,120° for all the diacids but dc-ΔAla, in which case, the global minimum appears at ψ1,ψ2≈ 13°,142°. This small difference should be attributed to the hybridization state of the α carbon atom, namely sp2 for dc-ΔAla and sp3 for dc-Gly, dc-Ala, dc-Aib, and dc-Val. These non-symmetric conformations are very similar to those found in the crystal state for malonamide derivatives.64 On the other hand, Figure 3a shows that the minima of dc-ΔAla with relative energies larger than 2 kcal/mol (i.e. minima with ψ1,ψ2= ±25.0°, ±25.0° and ±126.0°, ±126.0°) are located in the map diagonals. However, the lowest energy conformation remains asymmetric (ψ1≠ψ2), even though this diacid is chemically symmetric.
Figure 3b shows the energy minima found for the N,N’-diacetyl derivatives of the Gly, Ala, Aib and Val diamine surrogates, hereafter denoted dm-Gly,60 dm-Ala,60 dm-Aib, and dm-Val,62 respectively. The minimum energy conformations of the dm-Aib derivative have been calculated in this work at the B3LYP/6-31G(d) level (see the Supporting Information). For the reason mentioned above, structures with φ1,φ2; -φ1,-φ2; φ2,φ1; and -φ2,-φ1 are energetically equivalent, as reflected by the map symmetry. As can be seen, the lowest energy conformations of the diamines are located in the α region at φ1,φ2≈ 60°,60°. In addition, dm-Aib shows a low energy minima destabilized by 1.0 kcal/mol at φ1,φ2= 49°,173°. Minima with relative energies higher than 2 kcal/mol are essentially located in the map diagonal as well as in the regions labeled as ε and δ. As expected, the γ regions are scarcely populated for diamines and diacids derivatives (Figure 3), whereas they are found to be preferred by the N-acetyl-N’-methylamide derivatives of proteinogenic amino acids. This is because in the latter compounds, the γ conformations are stabilized by an intramolecular hydrogen bond defining a seven-membered ring, whereas this interaction is not possible in diamines and diacids.
In silico molecular engineering of a 310-helical motif using NCAD: A test case
In this section, we present a computational design study aimed at constructing non-symmetric retropeptides exhibiting a 310-helix conformation by using information extracted from NCAD. This is not intended to provide a deep and rigorous investigation of the resulting retropeptides, but to illustrate the potential utility of the database in applications involving peptide and protein engineering through a test case.
It is well known that Aib-based peptides have a remarkable tendency to adopt helical conformations both in solution and in the solid state.65–69 Helices of the 310 type are overwhelmingly formed by Aib homopeptides, independently of the environmental conditions, whereas small peptides containing a mixture of Aib and proteinogenic α-amino acids are found to accommodate either α- or 310-helices depending mainly on the peptide size and the Aib content.
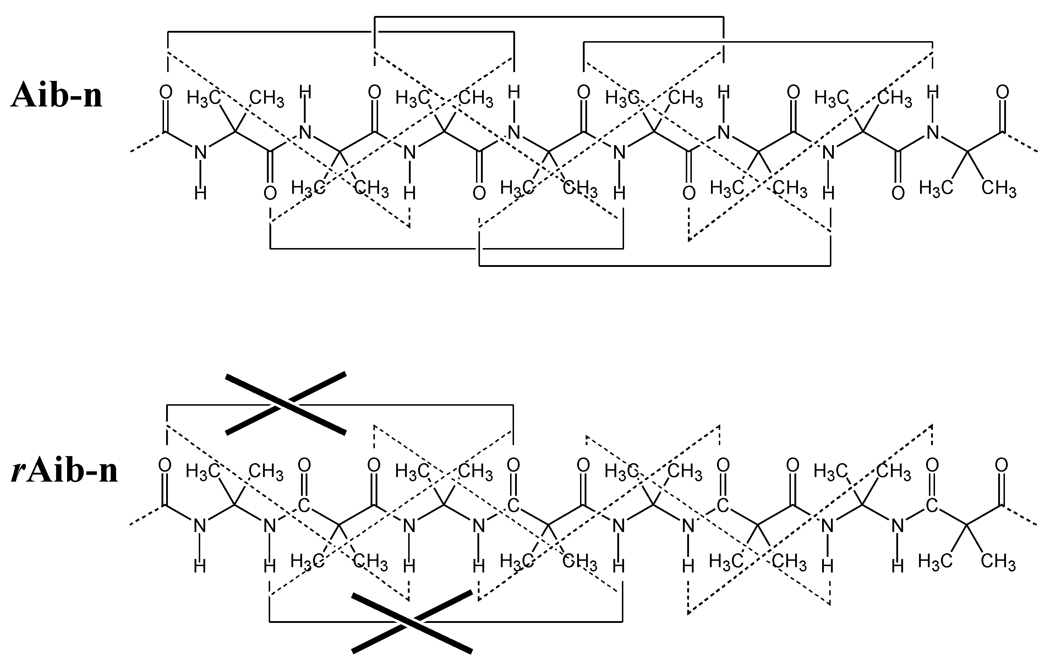
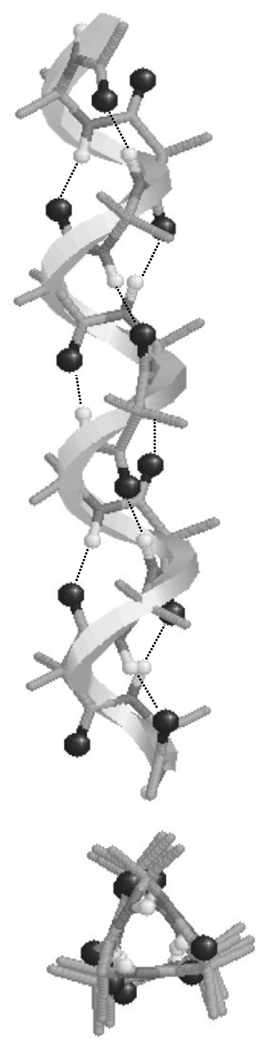
In an earlier work, we used retropeptide bonds to enhance the stability of the 310-helix with respect to the α-helix in peptides containing Aib.70 Specifically, Aib-containing homoretropeptides (hereafter denoted rAib-n, where n indicates the number of residues) cannot form the intramolecular hydrogen-bonding network that stabilizes the α-helix (i←i+4) and this results in unfavorable interactions between the C=O groups of residues i and i+4. In contrast, the molecular architecture of rAib-n (Scheme 3) in which diacid and diamine surrogates of Aib are alternated, (dm-Aib–dc-Aib)n/2, is fully compatible with the hydrogen-bonding scheme of the 310-helix (i←i+3). These features are schematically depicted in Figure 4, whereas Figure 5 shows the 310-helix adopted by rAib-n.
Scheme 3.
Figure 4.
Hydrogen bonding network for the α- and 310-helices (i←i+4 solid line and i←i+3 dashed line, respectively) of Aib-n and rAib-n.
Figure 5.
Side and top views of the 310-helix adopted by rAib-n.
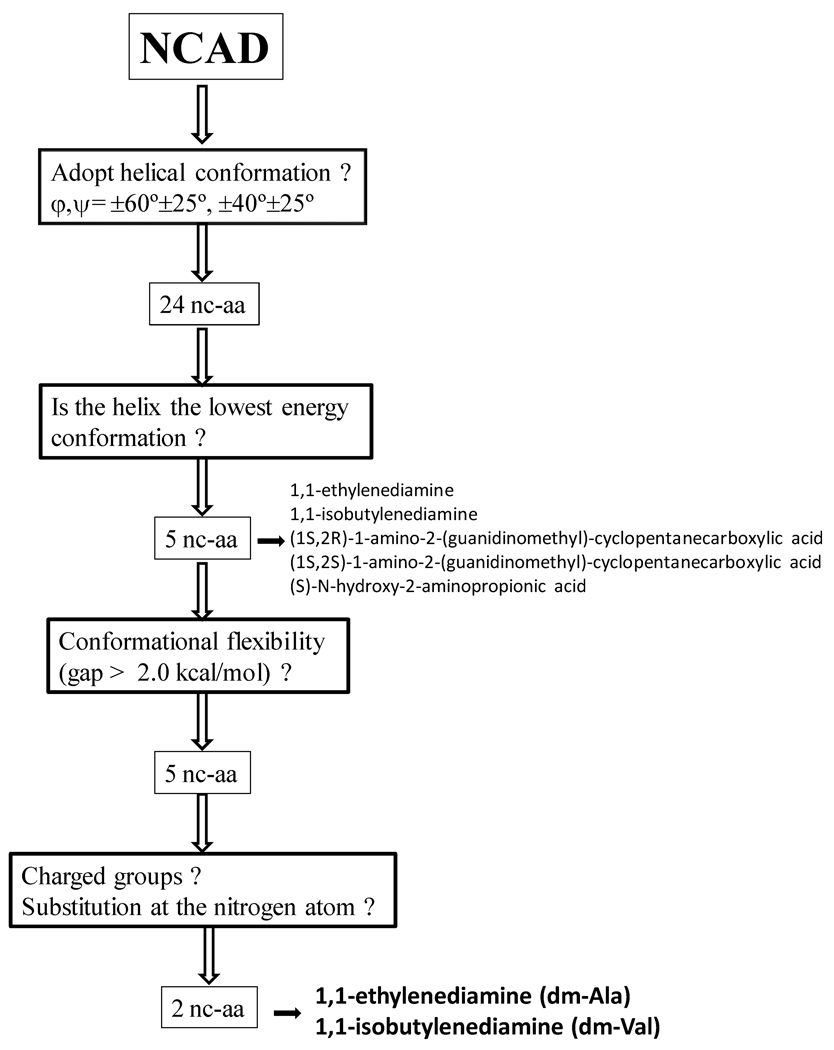
rAib-n are symmetric molecules, i.e. the two ends of the peptide are identical, which may be a limitation in some recognition processes. Accordingly, we used the information stored in the NCAD database to search for nc-aa that, after targeted replacements, facilitate the recognition process but retaining as much as possible the stability of 310-helix. The procedure followed for this purpose, which is schematized in Figure 6, can be summarized as follows:
NCAD was used to search for the nc-aa able to adopt a helical conformation with φ,ψ= +60°±25°, +40°±25° or φ,ψ= −60°±25°, −40°±25°. The number of nc-aa that satisfied this initial criterion was 24.
From the 24 nc-aa identified in (i), NCAD allowed identify those in which the helix correspond to the lowest energy conformation. This criterion was satisfied by 6 nc-aa including the diamine analogues of Ala, Aib and Val (dm-Ala, dm-Aib and dm-Val in Table 2). Obviously, dm-Aib was eliminated from the list since it is contained in the parent retropeptide.
From the 5 nc-aa provided by (ii), we looked for those with lower conformational flexibility. For this purpose, we used NCAD to exclude those nc-aa in which the energy gap between the helical global minimum and the most stable local minimum was lower than 2 kcal/mol. Unfortunately, no nc-aa was eliminated using this criterion since in all cases the gap was larger than 2 kcal/mol. However, the information provided by the database was considerably useful since it allowed ascertain that the lowest (2.4 kcal/mol) and highest (4.3 kcal/mol) value of this energy gap were for dm-Val and dm-Ala, respectively.
Inspection to the chemical structures stored in NCAD indicated that 2 of 5 nc-aa selected in (ii) and (iii) involved positively charged side groups [i.e. (1S,2R)- and (1S,2S)-1-amino-2-(guanidinomethyl)-cyclopentanecarboxylic acids]. These two nc-aa were eliminated in order to avoid undesirable repulsive electrostatic interactions, i.e. rAib-n does not involve negatively charged groups able to form attractive interactions with the positively charged groups. On the other hand, another nc-aa [i.e. (S)-N-hydroxy-2-aminopropionic acid] involves an N-hydroxylation that was expected to induce drastic distortions in the helix through alterations in the geometry of the hydrogen bonding network, being also removed from the list. Accordingly, dm-Ala and dm-Val were considered by NCAD as the more suitable candidates for the proposed substitutions.
Figure 6.
Search performed with the NCAD database to select nc-aa able to break the symmetry in rAib-n without alter significantly the stability of the 310-helix (see text).
The information stored in NCAD indicates that two different strategies can be followed to design a non-symmetric retropeptide based on rAib-n through a single mutation without decreasing significantly the stability of the 310-helix. The first strategy requires the replacement of dm-Aib by dm-Ala (i.e. removal of a methyl group in a diamide). The second involves the replacement of dm-Aib by the corresponding Val analogue. Although the helical conformation is energetically favored for the diamines of both Ala and Val, the larger conformational flexibility of the Val derivatives and the larger steric interactions expected for the isopropyl group in comparison to the methyl substituent suggest that the replacement of dm-Aib by the corresponding Ala analogue is more appropriate. Accordingly, the first strategy was selected for further investigation.
We first examined the most appropriate position to carry out the targeted replacement. For this purpose, three mutants of a retropeptide containing 12 residues, rAib-12 [i.e. (dm-Aib – dc-Aib)6], were considered, their sequences being:
Diamine-1: dm-Ala – dc-Aib – (dm-Aib – dc-Aib)5
Diamine-2: (dm-Aib – dc-Aib) – dm-Ala – dc-Aib – (dm-Aib – dc-Aib)4
Diamine-3: (dm-Aib – dc-Aib)2 – dm-Ala – dc-Aib – (dm-Aib – dc-Aib)3
The three retropeptides were built up in a 310-helix conformation71 and their molecular geometries were submitted to complete geometry optimizations using the B3LYP quantum mechanical method72,73 combined with the 6-31G(d) basis set. The stability of the three mutants under study was similar. Thus, the diamine-2 was more stable than diamines-1 and -3 by only 0.3 and 0.1 kcal/mol, respectively, indicating the lack of preference towards a given position for the mutation.
Comparison of the backbone dihedral angles obtained for the three optimized mutants to those derived from geometry optimization of rAib-12 at the same theoretical level indicates that the conformational distortion introduced by dm-Ala is almost negligible. Thus, the largest distortion was observed for diamine-1, where the φ1 dihedral of dm-Ala differs in 16.2° from the corresponding angle in rAib-12. For the rest of the mutants, no deviation was found to exceed 10°. Similarly, the end-to-end distance remained almost identical for the mutants and the parent peptide, the largest difference being 0.08 Å for diamine-1. These results confirm that the helical stability of rAib-12 is retained in the three mutants investigated. In order to quantitatively evaluate the effect of the mutation in the energetic stability of the 310-helix, the following isodesmic reaction has been considered:
| (1) |
Thus, the loss of stability of the helix (Δh) was defined according to:
| (2) |
where E refers to the energy of the different species. The Δh values obtained for diamines-1, -2 and -3 are 1.7, 1.4 and 1.5, respectively. Comparison between these Δh values and the relative stabilities mentioned above for the three mutants indicates that breaking the molecular symmetry of rAib-12 by replacement of one dm-Aib by dm-Ala produces a destabilization of 1.4 kcal/mol, independently of the position.
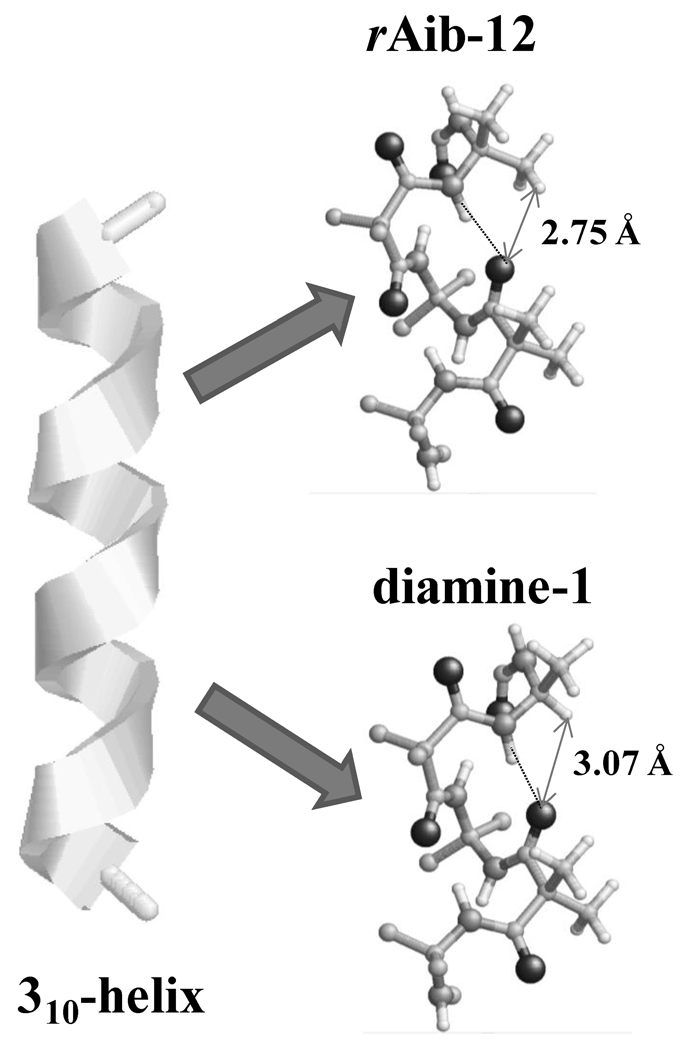
Inspection of the dipole moments calculated for the mutants indicates that incorporation of dm-Ala produces a very small variation with respect to rAib-12 (µ= 0.225 Debyes), the largest variation being of only 0.092 Debyes (diamine-1). Accordingly, the small destabilization detected by the substitution should not be attributed to electrostatic changes in the helix. However, a detailed analysis of the intramolecular distances may help explain this feature. Specifically, inspection of rAib-12 indicates that the methyl groups of the dm-Aib residues form a weak interaction with the oxygen atom of a proximal O=C moiety, which in turn, is involved in an intramolecular hydrogen bond with a neighboring amide hydrogen. This van der Waals interaction is illustrated in Figure 7, which also indicates that the average C-H⋯O=C distance is 2.75±0.01 Å. Evaluation of such a distance for the three mutants under study shows that removal of the methyl group induces a variation that becomes significantly larger at the mutated positions (Figure 7). Thus, the distance increases by 0.32±0.02 in the retropeptides mutated at the diamines, with respect to that found in rAib-12, and this may contribute to the loss of stability. It should be mentioned that the B3LYP method does not treat appropriately the dispersion effects. Therefore, the results described in this section illustrate the use of the database and should be considered only from a qualitative point view.
Figure 7.
Detail about the C-H⋯O=C interaction (arrows) in the 310-helix of rAib-12 and diamine-1. Averaged H⋯O distances are indicated.
Conclusions
In this work, NCAD has been extended through the incorporation of residues that may be used as building blocks for the construction of peptides with modified peptide bonds. The information related to N-substituted α-amino acids, thio-α-amino acids, and diamines and diacids used to build retropeptides described in the literature has been collected and integrated into the database. The conformational propensities of the new α-amino acid surrogates incorporated into NCAD have been compared to those of the corresponding coded residues. We also present an example illustrating the utility of NCAD and the usefulness of the new amino acid surrogates in peptide engineering. More specifically, the information contained in NCAD has been used to propose targeted replacements able to break the molecular symmetry of a retropeptide sequence that adopts the 310-helix.
Supplementary Material
Acknowledgements
Computer resources were generously provided by the Centre de Supercomputació de Catalunya (CESCA). Financial support from Ministerio de Ciencia e Innovación - FEDER (grants CTQ2007-62245, CTQ2008-00423-E/BQU, and CTQ2010-17436), Generalitat de Catalunya (research group 2009 SGR 925; XRQTC; ICREA Academia prize for excellence in research to C.A.), and Gobierno de Aragón (research group E40) is gratefully acknowledged. This project has been funded in part with Federal funds from the National Cancer Institute, National Institutes of Health, under contract number HHSN261200800001E. The content of this publication does not necessarily reflect the view of the policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organization imply endorsement by the U.S. Government. This research was supported [in part] by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research.
Footnotes
Supporting Information
Dihedral angles and relative energies of the minimum energy conformations found for dc-Ala, dc-Aib, and dm-Aib at the B3LYP/6-31G(d) level.
References
- 1.Voloshchuk N, Montclare JK. Incorporation of unnatural amino acids for synthetic biology. Mol BioSyst. 2010;6:65–80. doi: 10.1039/b909200p. [DOI] [PubMed] [Google Scholar]
- 2.Wu X, Schultz PG. Synthesis at the Interface of Chemistry and Biology. J Am Chem Soc. 2009;131:12497–12515. doi: 10.1021/ja9026067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hartman MCT, Josephson K, Szostak JW. Enzymatic aminoacylation of tRNA with unnatural amino acids. Proc Natl Acad Sci U.S.A. 2006;103:4356–4361. doi: 10.1073/pnas.0509219103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang L, Schultz PG. Expanding the genetic code. Angew Chem Int Ed. 2005;44:34–66. doi: 10.1002/anie.200460627. [DOI] [PubMed] [Google Scholar]
- 5.Hendrickson TL, De Crécy-Lagard V, Schimmel P. Incorporation of nonnatural amino acids into proteins. Annu Rev Biochem. 2004;73:147–176. doi: 10.1146/annurev.biochem.73.012803.092429. [DOI] [PubMed] [Google Scholar]
- 6.Wang L, Brock A, Herberich B, Schultz PG. Expanding the genetic code of Escherichia coli. Science. 2001;292:498–500. doi: 10.1126/science.1060077. [DOI] [PubMed] [Google Scholar]
- 7.Zanuy D, Ballano G, Jiménez AI, Casanovas J, Haspel N, Cativiela C, Curcó D, Nussinov R, Alemán C. Protein Segments with Conformationally Restricted Amino Acids Can Control Supramolecular Organization at the Nanoscale. J Chem Inf Model. 2009;49:1623–1629. doi: 10.1021/ci9001487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cejas MA, Kinney WA, Chen C, Leo GC, Tounge BA, Vinter JG, Joshi PP, Maryanoff BE. Collagen-related peptides: Self-assembly of short, single strands into a functional biomaterial of micrometer scale. J Am Chem Soc. 2007;129:2202–2203. doi: 10.1021/ja066986f. [DOI] [PubMed] [Google Scholar]
- 9.Yang ZM, Liang GL, Ma ML, Gao Y, Xu B. In vitro and in vivo enzymatic formation of supramolecular hydrogels based on self-assembled nanofibers of a beta-amino acid derivative. Small. 2007;3:558–562. doi: 10.1002/smll.200700015. [DOI] [PubMed] [Google Scholar]
- 10.Crisma M, Toniolo C, Royo S, Jiménez AI, Cativiela C. A helical, aromatic, peptide nanotube. Org Lett. 2006;8:6091–6094. doi: 10.1021/ol062600u. [DOI] [PubMed] [Google Scholar]
- 11.Brea RJ, Amorín M, Castedo L, Granja JR. Methyl-blocked dimeric alpha,gamma-peptide nanotube segments: Formation of a peptide heterodimer through backbone-backbone interactions. Angew Chem Int Ed. 2005;44:5710–5713. doi: 10.1002/anie.200501555. [DOI] [PubMed] [Google Scholar]
- 12.Hartgerink JD, Beniash E, Stupp SI. Self-assembly and mineralization of peptide-amphiphile nanofibers. Science. 2001;294:1684–1688. doi: 10.1126/science.1063187. [DOI] [PubMed] [Google Scholar]
- 13.Ersmark K, Del Valle JR, Hanessian S. Chemistry and biology of the aeruginosin family of serine protease inhibitors. Angew Chem Int Ed. 2008;47:1202–1223. doi: 10.1002/anie.200605219. [DOI] [PubMed] [Google Scholar]
- 14.Menard J, Patchett AA. Angiotensin-converting enzyme inhibitors. Adv Protein Chem. 2001;56:13–75. doi: 10.1016/s0065-3233(01)56002-7. [DOI] [PubMed] [Google Scholar]
- 15.Lee HS, Guo J, Lemke E, Dimla R, Schultz PG. Genetic Incorporation of a Small, Environmentally Sensitive, Fluorescent Probe into Proteins in Saccharomyces cerevisiae. J Am Chem Soc. 2009;131:12921–12923. doi: 10.1021/ja904896s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shinohara H, Kusaka T, Yokota E, Monden R, Sisido M. Electron transfer between redox enzymes and electrodes through the artificial redox proteins and its application for biosensors. Sens Actuators B. 2000;65:144–146. [Google Scholar]
- 17.Cellitti SE, Jones DH, Lagpacan L, Hao X, Zhang Q, Hu H, Brittain SM, Brinker A, Caldwell J, Bursulaya B, Spraggon G, Brock A, Ryu Y, Uno T, Schultz PG, Geierstanger BH. In vivo incorporation of unnatural amino acids to probe structure, dynamics, and ligand binding in a large protein by nuclear magnetic resonance spectroscopy. J Am Chem Soc. 2008;130:9268–8281. doi: 10.1021/ja801602q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Reid PJ, Loftus C, Beeson CC. Evaluating the potential of fluorinated tyrosines as spectroscopic probes of local protein environments: A UV resonance Raman study. Biochemistry. 2003;42:2441–2448. doi: 10.1021/bi0202676. [DOI] [PubMed] [Google Scholar]
- 19.Revilla-López G, Torras J, Curcó D, Casanovas J, Calaza MI, Zanuy D, Jiménez AI, Cativiela C, Nussinov R, Grodzinski P, Alemán C. NCAD, a Database Integrating the Intrinsic Conformational Preferences of Non-Coded Amino Acids. J Phys Chem B. 2010;114:7413–7422. doi: 10.1021/jp102092m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Perczel A, Angyán JG, Kajtar M, Viviani W, Rivail JL, Marcoccia JF, Csizmadia IG. Peptide models .1. topology of selected peptide conformational potential-energy surfaces (glycine and alanine derivatives) J Am Chem Soc. 1991;113:6256–6265. [Google Scholar]
- 21.Hudáky I, Kiss R, Perczel A. A nomenclature of peptide conformers. J Mol Struct. 2004;675:177–183. [Google Scholar]
- 22.Siodlak D, Gajewska M, Macedowska A, Rzeszotarska B. Conformational studies into N-methylation of alanine diamide models: A quantitative approach. J Mol Struct (Theochem) 2006;775:47–59. [Google Scholar]
- 23.Malde AK, Khedkar SA, Coutinho ECJ. The omega, phi, and psi space of N-hydroxy-N-methylacetamide and N-acety-N '-hydroxy-N '-methylamide of alanine and their boron isosteres. J Chem Theory Comput. 2006;2:312–321. doi: 10.1021/ct050242v. [DOI] [PubMed] [Google Scholar]
- 24.Alemán C. N-amination of peptides: A theoretical study. J Phys Chem A. 2002;106:1441–1449. [Google Scholar]
- 25.Chatterjee J, Gilon C, Hoffman A, Kessler H. N-Methylation of Peptides: A New Perspective in Medicinal Chemistry. Acc Chem Res. 2008;41:1331–1342. doi: 10.1021/ar8000603. [DOI] [PubMed] [Google Scholar]
- 26.Teixido M, Albericio F, Giralt EJ. Solid-phase synthesis and characterization of N-methyl-rich peptides. Pept Res. 2005;65:153–166. doi: 10.1111/j.1399-3011.2004.00213.x. [DOI] [PubMed] [Google Scholar]
- 27.Rainaldi M, Moretto V, Crisma M, Peggion E, Mammi S, Toniolo C, Cavicchioni G. Peptoid residues and beta-turn formation. J Pept Sci. 2002;8:241–252. doi: 10.1002/psc.392. [DOI] [PubMed] [Google Scholar]
- 28.Cornell WD, Gould IR, Kollman PA. The effects of basis set and blocking groups on the conformational energies of glycyl and alanyl dipeptides - A Hartree-Fock and MP2 study. J Mol Struct (Theochem) 1997;392:101–109. [Google Scholar]
- 29.Ye Y, Liu M, Kao JLK, Marshall GR. Peptide-bond modification for metal coordination: Peptides containing two hydroxamate groups. Biopolymers (Pept Sci) 2003;71:489–515. doi: 10.1002/bip.10471. [DOI] [PubMed] [Google Scholar]
- 30.Dupont V, Lecoq A, Mangeot JP, Aubry A, Boussard G, Marraud M. Conformational perturbations induced by N-amination and N-hydroxylation of peptides. J Am Chem Soc. 1993;115:8898–8906. [Google Scholar]
- 31.Clausen K, Thorsen M, Lawesson SO. Studies on amino acids and peptides .1. synthesis of N-benzyloxycarbonylendothiodipeptides esters. Tetrahedron. 1981;37:3635–3639. [Google Scholar]
- 32.Ilankumaran P, Ramesha AR, Chandrasekaran S. A facile conversion of amides and lactams to thioamides and thiolactams using tetrathiomolybdate. Tetrahedron Lett. 1995;36:8311–8314. [Google Scholar]
- 33.Yokoyama M, Hasegawa Y, Hatanaka H, Kawazoe Y, Imamoto T. Improved O/S exchange reagents. Synthesis. 1984;84:827–829. [Google Scholar]
- 34.Bartlett PA, Spear KL, Jacobsen NE. A thioamide substrate of carboxypeptidase-A. Biochemistry. 1982;21:1608–1611. doi: 10.1021/bi00536a022. [DOI] [PubMed] [Google Scholar]
- 35.Wildemann D, Schiene-Fischer C, Aumüller T, Bachmann A, Kiefhaber T, Lücke C, Fischer G. A nearly isosteric photosensitive amide-backbone substitution allows enzyme activity switching in ribonuclease S. J Am Chem Soc. 2007;129:4910–4918. doi: 10.1021/ja069048o. [DOI] [PubMed] [Google Scholar]
- 36.Kessler H, Matter H, Geyer A, Diehl HJ, Köck M, Kurz G, Opperdoes FR, Callens M, Wierenga RK. Selective inhibition of trypanosomal triosephosphate isomerase by a thiopeptide. Angew Chem Int Ed Engl. 1992;31:328–330. [Google Scholar]
- 37.Guthrie DJS, Williams CH, Elmore DT. Configuration of thionopeptide bond in solution. Int J Pept Protein Res. 1986;28:208–211. [Google Scholar]
- 38.Hollósi M, Majer Z, Zewdu M, Ruff F, Kajtár M, Kövér KE. Mixed intramolecular H-bonds of secondary thioamides. Tetrahedron. 1988;44:195–202. [Google Scholar]
- 39.Doi M, Takehara S, Ishida T, Inoue M. Effects of thioamide substitution for the enkephalin conformation -crystal-structure of Boc-Tyr-Gly-Gly-Phe-Psi[CSNH]Leu-Obzl. Int J Pep Protein Res. 1989;34:369–373. [PubMed] [Google Scholar]
- 40.Zhao J, Wildemann D, Jakob M, Vargas C, Schiene-Fischer C. Direct photomodulation of peptide backbone conformations. Chem Commun. 2003:2810–2811. doi: 10.1039/b309927j. [DOI] [PubMed] [Google Scholar]
- 41.Wiberg KB, Rush DJ. Solvent effects on the thioamide rotational barrier: An experimental and theoretical study. J Am Chem Soc. 2001;123:2038–2046. doi: 10.1021/ja003586y. [DOI] [PubMed] [Google Scholar]
- 42.Alemán C. On the ability of modified peptide links to form hydrogen bonds. J Phys Chem A. 2001;105:6717–6723. [Google Scholar]
- 43.Artis DR, Lipton MA. Conformations of thioamide-containing dipeptides: A computational study. J Am Chem Soc. 1998;120:12200–12206. [Google Scholar]
- 44.Fletcher MD, Campbell MM. Partially modified retro-inverso peptides: Development, synthesis, and conformational behavior. Chem Rev. 1998;98:763–795. doi: 10.1021/cr970468t. [DOI] [PubMed] [Google Scholar]
- 45.Goodman M, Chorev M. Concept of linear modified retro-peptide structures. Acc Chem Res. 1979;12:1–7. [Google Scholar]
- 46.Chorev M, Willson CG, Goodman M. General approach to retro-isomeric linear peptide-synthesis. J Am Chem Soc. 1977;99:8075-–8076. [Google Scholar]
- 47.Volonterio A, Bravo P, Zanda M. Synthesis of partially modified retro and retroinverso psi[NHCH(CF3)]-peptides. Org Lett. 2000;2:1827–1830. doi: 10.1021/ol005876p. [DOI] [PubMed] [Google Scholar]
- 48.Sani M, Bruche L, Chiva G, Fustero S, Piera J, Volonterio A, Zanda M. Highly stereoselective tandem aza-michael addition-enolate protonation to form partially modified retropeptide mimetics incorporating a trifluoroalanine surrogate. Angew Chem Int Ed. 2003;42:2060–2063. doi: 10.1002/anie.200250711. [DOI] [PubMed] [Google Scholar]
- 49.Ceretti S, Luppi G, De Pol S, Formaggio F, Crisma M, Toniolo C, Tomasini C. Total synthesis of sequential retro-peptide oligomers. Eur J Org Chem. 2004;20:4188–4196. [Google Scholar]
- 50.Ando S, Mitsuyasu K, Soeda Y, Hidaka M, Ito Y, Matsubara K, Shindo M, Uchida Y, Aoyagi H. Structure-activity relationship of indolicidin, a Trp-rich antibacterial peptide. J Pept Sci. 2010;16:171–177. doi: 10.1002/psc.1217. [DOI] [PubMed] [Google Scholar]
- 51.Phan-Chan-Du A, Petit MC, Guichard G, Briand JP, Muller S, Cung MT. Structure of antibody-bound peptides and retro-inverso analogues. A transferred nuclear overhauser effect spectroscopy and molecular dynamics approach. Biochemistry. 2001;40:5720–5727. doi: 10.1021/bi001151h. [DOI] [PubMed] [Google Scholar]
- 52.Pal-Bhowmick I, Pandey RP, Jarori GK, Kar S, Sahal D. Structural and functional studies on Ribonuclease S, retro S and retro-inverso S peptides. Biochem Biophys Res Commun. 2007;364:608–613. doi: 10.1016/j.bbrc.2007.10.056. [DOI] [PubMed] [Google Scholar]
- 53.Petit MC, Benkirane N, Guichard G, Phan-Chan-Du A, Marraud M, Cung MT, Briand JP, Muller S. Solution structure of a retro-inverso peptide analogue mimicking the foot-and-mouth disease virus major antigenic site - Structural basis for its antigenic cross-reactivity with the parent peptide. J Biol Chem. 1999;274:3686–3692. doi: 10.1074/jbc.274.6.3686. [DOI] [PubMed] [Google Scholar]
- 54.Sakurai K, Chung HS, Kahne D. Use of a retroinverso p53 peptide as an inhibitor of MDM2. J Am Chem Soc. 2004;126:16288–16289. doi: 10.1021/ja044883w. [DOI] [PubMed] [Google Scholar]
- 55.Guptasarma P. Reversal of peptide backbone direction may result in the mirroring of protein-structure. FEBS Lett. 1992;310:205–210. doi: 10.1016/0014-5793(92)81333-h. [DOI] [PubMed] [Google Scholar]
- 56.Preissner R, Goede A, Michalski E, Frömmel C. Inverse sequence similarity in proteins and its relation to the three-dimensional fold. FEBS Lett. 1997;414:425–429. doi: 10.1016/s0014-5793(97)00907-1. [DOI] [PubMed] [Google Scholar]
- 57.Shukla A, Raje M, Guptasarma P. A backbone-reversed form of an all-beta alpha-crystallin domain from a small heat-shock protein (retro-HSP12.6) folds and assembles into structured multimers. J Biol Chem. 2003;278:26505–26510. doi: 10.1074/jbc.M303123200. [DOI] [PubMed] [Google Scholar]
- 58.Alemán C. Chain conformation in polyretropeptides .3. Design of a 3(10) helix using alpha,alpha-dialkylated amino acids and retropeptide bonds. Proteins. 1997;29:575–582. doi: 10.1002/(sici)1097-0134(199712)29:4<575::aid-prot16>3.0.co;2-j. [DOI] [PubMed] [Google Scholar]
- 59.Alemán C, Franco L, Puiggalí J. Conformations of nylon 1, N according to the number of methylene carbons. J Macromolecules. 1994;27:4298–4303. [Google Scholar]
- 60.Alemán C, Puiggalí J. Retromodified residues - small peptides and polymers - interactions, force-field parametrizations and conformational - analyses. J Org Chem. 1995;60:910–924. [Google Scholar]
- 61.Sandrone G, Dixon DA, Hay BP. Conformational analysis of malonamide, N,N '-dimethylmalonamide, and N,N,N ',N '-tetramethylmalonamide. J Phys Chem A. 1999;103:3554–3561. [Google Scholar]
- 62.Alemán C. A computational study of partially modified retro-inverso valine dipeptides: Effect of the side chain on the conformational preferences of malonyl and gem-diamino residues. J Phys Chem B. 2001;105:860–866. [Google Scholar]
- 63.Alemán C. Effects of the φ[NHCO] retromodification on dehydroalanine dipeptide. J Biol Struct Dyn. 1996;14:193–199. doi: 10.1080/07391102.1996.10508108. [DOI] [PubMed] [Google Scholar]
- 64.Alemán C, Pérez JJ. SCF-MO study of the preferred conformation of a symmetrical malonamide derivative -N’N’-dimethyl malonamide. J Mol Struct (Theochem) 1993;2:221–227. [Google Scholar]
- 65.Di Blasio B, Pavone V, Lombardi A, Pedone C, Benedetti E. Noncoded residues as building-blocks in the design of specific secondary structures - symmetrically disubstituted glicines and beta-alanine. Biopolymers. 1993;33:1037–1049. doi: 10.1002/bip.360330706. [DOI] [PubMed] [Google Scholar]
- 66.Balaram P. Non-Standard amino acids in peptide design and protein engineering. Curr Opin Struct Biol. 1992;64:1061–1068. [Google Scholar]
- 67.Toniolo C, Crisma M, Bonora GM, Benedetti E, Di Blasio B, Pavone V, Pedone C, Santini A. Linear oligopeptides series .229. preferred conformation of the thermally blocked (Aib)10 homo-oligopeptide - a long regular 3(10) helix. Biopolymers. 1991;;31:129–138. [Google Scholar]
- 68.Otoda K, Kitagawa Y, Kimura S, Imanishi Y. Chain lenght dependent transition of 3(10)- to alpha-helix of Boc-(Ala Aib)(N)-Ome. Biopolymers. 1993;33:1337–1345. [Google Scholar]
- 69.Toniolo C, Bonora GM, Barone V, Bavoso A, Benedetti E, Di Blasio B, Grimaldi P, Lelj F, Pavone V, Pedone C. Linear oligopeptides .121. conformation of pleionomers of alpha-aminoisobutyric-acid. Macromolecules. 1985;18:895–902. [Google Scholar]
- 70.Alemán C. Chain conformation in polyretropeptides .3. Design of a 3(10) helix using alpha,alpha-dialkylated amino acids and retropeptide bonds. Proteins. 1997;29:575–582. doi: 10.1002/(sici)1097-0134(199712)29:4<575::aid-prot16>3.0.co;2-j. [DOI] [PubMed] [Google Scholar]
- 71.Alemán C, Roca R, Luque FJ, Orozco M. Helical preferences of alanine, glycine, and aminoisobutyric homopeptides. Proteins. 1997;28:83–93. [PubMed] [Google Scholar]
- 72.Becke AD. Density-functional thermochemistry .3. the role of exact exchange. J Chem Phys. 1993;98:5648–5652. [Google Scholar]
- 73.Lee C, Yang W, Parr RG. Development of the Colle-Salvetti correlation-energy formula into a functional of electron-density. Phys Rev B. 1988;37:785–789. doi: 10.1103/physrevb.37.785. [DOI] [PubMed] [Google Scholar]
- 74.Hariharan PC, Pople JA. Influence of polarization functions on molecular-orbital hydrogenation energies. Theor Chim Acta. 1973;28:213–222. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.