Figure 4.
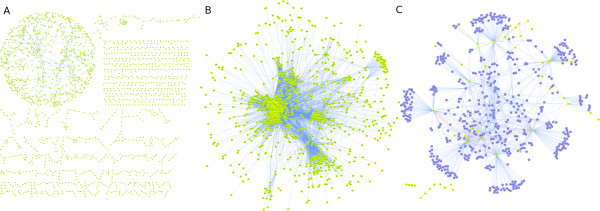
BIANA networks created from user provided datasets. BIANA Cytoscape plugin has been used to generate the relation networks of three different user specific datasets (data is available at http://sbi.imim.es/web/BIANA.php). Represented entities are: Proteins (green nodes), genes (blue nodes), interactions and metabolic relations (blue edges) and cooperation (red edges). A) Metabolic network reconstruction where a relation is established and scored between each pair of possible chained enzymatic reactions. A chained reaction between enzyme A and enzyme B is possible when there is at least one chemical compound in the intersection, acting at the same time as product of enzyme A and substrate of enzyme B. The network has been filtered with score greater than 1.2 (score is based on the plausibility of observing chemical compounds in the intersection, according to their own frequency and the frequency of other products of enzyme A and other substrates of enzyme B that do not take part in chaining reactions). B) Protein-protein interaction network predicted from sequences/structure distant patterns as described in Espadaler et al. [22]. Only human proteins are shown and interactions coming from set I3 (see a detailed explanation in the original work). C) Network representation of cooperative transcription factors and their regulated genes described at Aguilar et al. [23]. Only transcription factors cooperating with others have been represented.