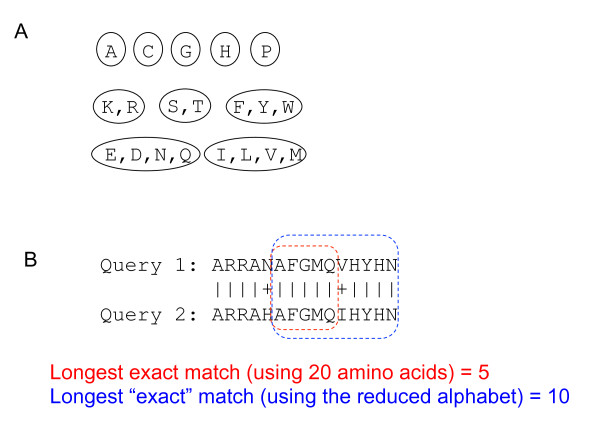
Figure 1.
Speeding up protein database searching by using a reduced alphabet of amino acids. (A) A reduced alphabet with ten symbols, in which, for example, K (Lysine) and R (Arginine) are grouped and represented by a single symbol because of their similar chemical properties. (B) The utilization of reduced alphabet will yield longer (and thus more efficient) sequence seeds that are common in homologous proteins. In this example, the maximum exact match (MEM) in the reduced alphabet is of length 10 between the pair of homologous proteins, whereas the MEM in 20-aa alphabet is of length 5. Hence, to retrieve this alignment in the database searching, one must retain the seeds of 10 or longer when using reduced alphabet, and retain the seeds of 5 or longer when using 20-aa alphabet. In this case, the efficiency of the seeds in reduced alphabet is much higher because only the ratio of the number of random seeds in these two cases is about 10-10/20-5 ≈ 10-2.