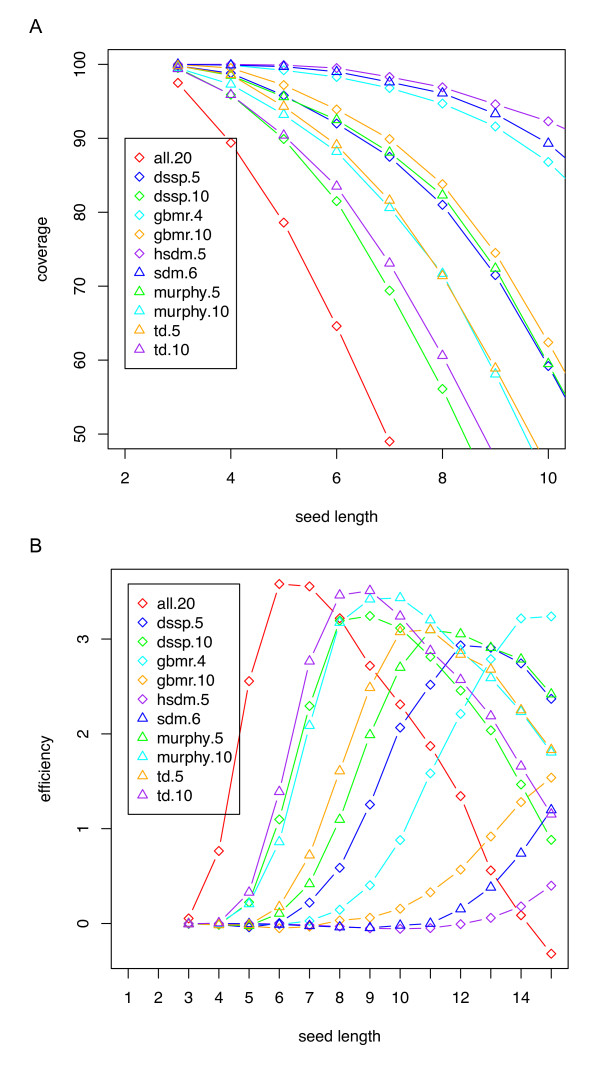
Figure 2.
Comparison of the performance of different reduced amino acid alphabets. The performance of a reduced alphabet of amino acids is measured by the coverage of seeds in the alphabet, defined as the percentage of pairs of alignments (of distantly related proteins of 20%-40% identify) that have at least one of the seeds of certain length (A), and the efficiency, defined as log ratio of the percentage of pairs of alignments that have at least one of the seeds of a defined length and the percentage of any pairs of unrelated sequences that share at least one of the seeds (B). See Table 1 for details of the alphabets. Note that only a subset of the alphabets is shown in the figure for clarity.