Figure 2.
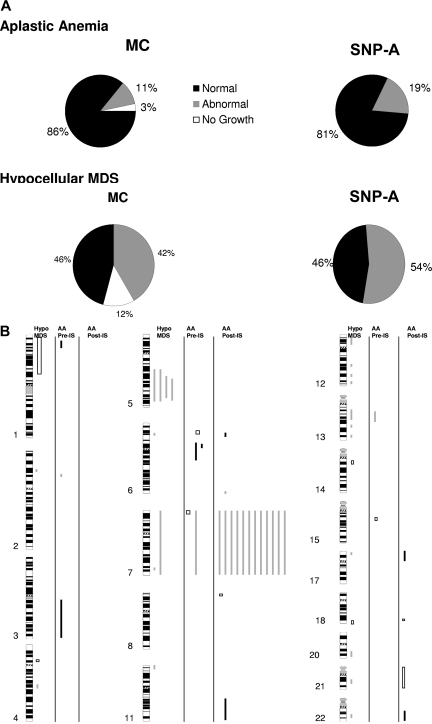
Frequency and genomic distribution of lesions detected by SNP-A. (A) Metaphase karyotyping identified lesions in a subset of patients with AA (top left); however, 3% of the patients had noninformative MC because of failure of growth. When MC and SNP-A karyotyping were combined, the detection rate for chromosomal lesions was increased (top right). In addition, the noninformative cases were resolved. For hMDS, when MC and SNP-A karyotyping were combined, the detection rate for was increased from 42% to 54% (bottom). (B) Genomic distribution of lesions detected by SNP-A in the analyses of hMDS and AA. □ and illustrate genomic gains and losses, respectively. ■ depicts regions of segmental UPD.