Table 2. Clustering Parameters and MCA Set Sizes.
| Parameter Type | Original Set | Pruned |
| K | 2, 4, 6, 8, 10, 12, 14 | 2, 4 |
| Transform | raw (untransformed) | |
| center (zero centered) | ||
| zscore (mean center, sample standard deviation of 1) | ||
| normMax (normalized to maximum value) | ||
| rangeScale (full range scaled to 1) | ||
| log10 (log base 10) | ||
| pow (power to 0.5) | ||
| pareto | ||
| FFT | ||
| diff (differential) | diff | |
| normMax_log10 (normMax follwed by log10) | ||
| zscore_log10 (zscore followed by log10) | zscore_log10 | |
| Distance | euclidean | |
| correlation | correlation | |
| cityblock | ||
| cosine | cosine | |
| chebychev | ||
| Algorithm | Ncut (Ncut Segmentation) | |
| AP (Affinity Propagation [18]) | ||
| SOM (Self-Organized Map [25]) | ||
| Kmeans | ||
| Hierarchical | Hierarchical | |
| Set Size | 1320 | 331 |
The parameters of clustering used in application to the EGF4 dataset including the short names used throughout this work as well as increased description of the parameters. Pruned parameters are parameter sets removed from 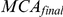 because their removal improved overall biological enrichment by at least 2% and did not negatively affect any one category by more than 10%.
because their removal improved overall biological enrichment by at least 2% and did not negatively affect any one category by more than 10%.
