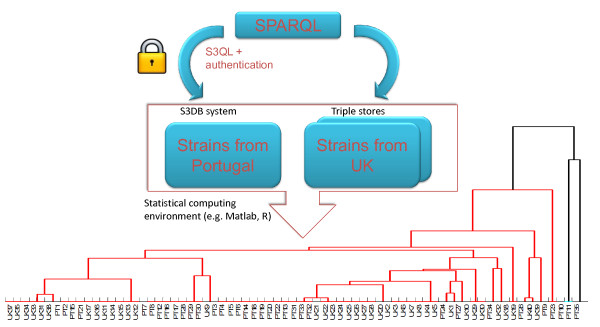
Figure 7.
Workflow for the creation of a hierarchical cluster of MLST profiles from strains collected in Portugal and in the UK. Two sources are chosen to perform the MLST profile assembly - an S3DB deployment, which holds the ITQB Staphylococcus database, and the SPARQL endpoint configured to host the Staphylococcus aureus MLST database. Although data in the MLST SPARQL endpoint is publicly available, to access data in S3DB, the user needs to provide an authentication token and a user_id as well as assemble the S3QL queries to retrieve the data [select(S|R172930) and select(S|R167271)], which may also be formulated as SPARQL. A data structure is assembled that can finally be analysed using hierarchical clustering methods such as the ones made available by Matlab. The complete MLST dataset used to obtain the graph is provided in additional file 2.