Abstract
Mahella australiensis Bonilla Salinas et al. 2004 is the type species of the genus Mahella, which belongs to the family Thermoanaerobacteraceae. The species is of interest because it differs from other known anaerobic spore-forming bacteria in its G+C content, and in certain phenotypic traits, such as carbon source utilization and relationship to temperature. Moreover, it has been discussed that this species might be an indigenous member of petroleum and oil reservoirs. This is the first completed genome sequence of a member of the genus Mahella and the ninth completed type strain genome sequence from the family Thermoanaerobacteraceae. The 3,135,972 bp long genome with its 2,974 protein-coding and 59 RNA genes is a part of the Genomic Encyclopedia of Bacteria and Archaea project.
Keywords: strictly anaerobic, motile, spore-forming, Gram-positive, moderately thermophilic, chemoorganotrophic, Thermoanaerobacteraceae, GEBA
Introduction
Strain 50-1 BONT (= DSM 15567 = CIP 107919) is the type strain of Mahella australiensis, and the type and only species of the monotypic genus Mahella [1,2]. The genus name is derived from the Neo-Latin word Mahella (named in honor of the American microbiologist R. A. Mah, for his important contribution to the taxonomy of anaerobes) [2]. The species epithet is derived from the Neo-Latin word australiensis (related to Australia) [1]. Strain 50-1 BONT was isolated from the Riverslea Oil Field in the Bowen-Surat basin in Queensland, eastern Australia [1]. No further isolates have been reported for M. australiensis. Here we present a summary classification and a set of features for M. australiensis 50-1 BONT, together with the description of the complete genomic sequencing and annotation.
Classification and features
A representative genomic 16S rRNA sequence of M. australiensis was compared using NCBI BLAST under default settings (e.g., considering only the high-scoring segment pairs (HSPs) from the best 250 hits) with the most recent release of the Greengenes database [3] and the relative frequencies, weighted by BLAST scores, of taxa and keywords (reduced to their stem [4] were determined. The three most frequent genera were Clostridium (76.6%), Mahella (18.5%) and Pelotomaculum (4.8%) (36 hits in total). Regarding the two hits to sequences from members of the species, the average identity within HSPs was 99.9%, whereas the average coverage by HSPs was 100.0%. Among all other species, the one yielding the highest score was Pelotomaculum isophthalicicum, which corresponded to an identity of 88.5% and a HSP coverage of 49.0%. (Note that the Greengenes databases uses the INSDC (= EMBL/NCBI/DDBJ) annotation, which is not an authoritative source for nomenclature or classification.) The highest-scoring environmental sequence was DQ378192 ('oil-polluted soil clone F28 Pitesti'), which showed an identity of 98.5% and a HSP coverage of 98.0%. The five most frequent keywords within the labels of environmental samples which yielded hits were 'microbi' (3.7%), 'anaerob' (2.9%), 'digest' (2.2%), 'soil' (2.0%) and 'thermophil' (1.7%) (213 hits in total). The five most frequent keywords within the labels of environmental samples which yielded hits of a higher score than the highest scoring species were 'microbi' (4.4%), 'anaerob' (3.3%), 'digest' (3.2%), 'soil' (2.6%) and 'condit, denitrification-induc, paddi, popul, respons, rice' (1.9%) (123 hits in total). These keywords reflect some of the ecological and physiological properties reported for strain 50-1 BONT in the original description [1].
Figure 1 shows the phylogenetic neighborhood of M. australiensis 50-1 BONT in a 16S rRNA based tree. The sequences of the three 16S rRNA gene copies in the genome differ from each other by up to two nucleotides, and differ by up to four nucleotides from the previously published 16S rRNA sequence (AY331143).
Figure 1.
Phylogenetic tree highlighting the position of M. australiensis strain 50-1 BONT relative to the other type strains within the order Thermoanaerobacterales. The tree was inferred from 1,275 aligned characters [5,6] of the 16S rRNA gene sequence under the maximum likelihood criterion [7] and rooted in accordance with the current taxonomy. The branches are scaled in terms of the expected number of substitutions per site. Numbers to the right of bifurcations are support values from 950 bootstrap replicates [8] if larger than 60%. Lineages with type strain genome sequencing projects registered in GOLD [9] are labeled with one asterisk, those registered as 'Complete and Published' with two asterisks [10,11]. Apparently, even the best BLAST hits show a low degree of similarity to M. australiensis (see above), in agreement with the isolated position of the species in the latest version of the 16S rRNA phylogeny from the All-Species-Living-Tree Project [12]. The species selection for Figure 1 was based on the current taxonomic classification (Table 1).
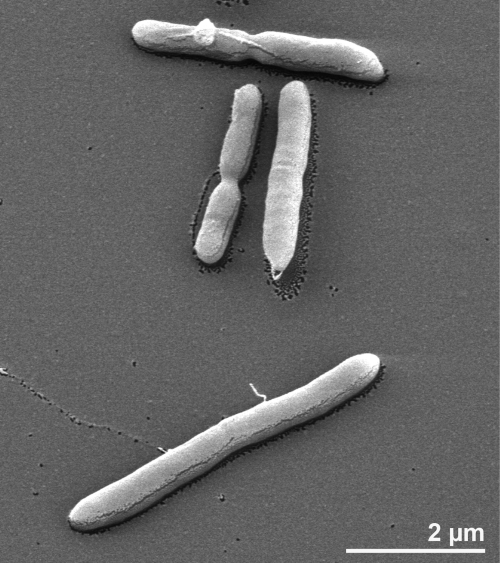
The cells of strain 50-1 BONT are generally rod-shaped with a size of 3–20 x 0.5 µm (Figure 2). They occur singly or in pairs [1]. Strain 50-1 BONT stains Gram-positive and is spore-forming (Table1). The organism is described to be motile by peritrichous flagella, with a mean of four flagella per cell [1] (not visible in Figure 2). Strain 50-1 BONT was found to be a strictly anaerobic chemoorganotroph which requires 0.1% NaCl for optimal growth [1], but is also able to grow in the presence of up to 4% NaCl [1]. The organism can use a wide range of carbohydrates as carbon and energy sources, including arabinose, cellobiose, fructose, galactose, glucose, mannose, sucrose, xylose and yeast extract [1]. Lactate, formate, ethanol, acetate, H2, and CO2 are the end products of the glucose metabolism [1]. The temperature range for growth is between 30°C and 60°C, with the optimum at 50°C [1]. Mesothermophilia distinguishes M. australiensis from its closest relatives, such as the members if the genus Thermoanaerobacterium [1]. After seven days of incubation at 50°C, round colonies (1–2 mm diameter) were found in roll tubes [1]. The pH range for growth is between 5.5 and 8.8, with an optimum at pH 7.5 [1]. Strain 50-1 BONT was not able to reduce thiosulfate or to hydrolyze starch [1]. Moreover, it does not use elemental sulfur, sulfate, sulfite, nitrate or nitrite as electron acceptors [1]. The generation time of the strain 50-1 BONT was 11 h [1].
Figure 2.
Scanning electron micrograph of M. australiensis 50-1 BONT
Table 1. Classification and general features of M. australiensis 50-1 BONT according to the MIGS recommendations [13] and the NamesforLife database [14].
| MIGS ID | Property | Term | Evidence code |
|---|---|---|---|
| Current classification | Domain Bacteria | TAS [15] | |
| Phylum Firmicutes | TAS [16,17] | ||
| Class Clostridia | TAS [18,19] | ||
| Order Thermoanaerobacterales | TAS [18,20] | ||
| Family Thermoanaerobacteraceae | TAS [18,21] | ||
| Genus Mahella | TAS [1] | ||
| Species Mahella australiensis | TAS [1] | ||
| Type strain 50-1 BON | TAS [1] | ||
| Gram stain | positive | TAS [1] | |
| Cell shape | rod-shaped | TAS [1] | |
| Motility | motile by peritrichous flagella | TAS [1] | |
| Sporulation | swollen sporangia, terminal spores | TAS [1] | |
| Temperature range | 30°C–60°C | TAS [1] | |
| Optimum temperature | 50°C | TAS [1] | |
| Salinity | 0.1%-4% NaCl | TAS [1] | |
| MIGS-22 | Oxygen requirement | strictly anaerobic | TAS [1] |
| Carbon source | arabinose, cellobiose, fructose, galactose, glucose, mannose, sucrose, xylose and yeast extract |
TAS [1] | |
| Energy metabolism | chemoorganotroph | TAS [1] | |
| MIGS-6 | Habitat | oil fields | TAS [1] |
| MIGS-15 | Biotic relationship | free-living | NAS |
| MIGS-14 | Pathogenicity | not reported | |
| Biosafety level | 1 | TAS [22] | |
| Isolation | oil well in Queensland | TAS [1] | |
| MIGS-4 | Geographic location | Riverslea Oil Field in the Bowen-Surat basin, Queensland, Australia | TAS [1] |
| MIGS-5 | Sample collection time | 1997 | NAS |
| MIGS-4.1 | Latitude | roughly -27.32 | NAS |
| MIGS-4.2 | Longitude | roughly 148.72 | NAS |
| MIGS-4.3 | Depth | not reported | |
| MIGS-4.4 | Altitude | not reported |
Evidence codes - IDA: Inferred from Direct Assay (first time in publication); TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from of the Gene Ontology project [23]. If the evidence code is IDA, the property was directly observed by one of the authors or an expert mentioned in the acknowledgements.
Chemotaxonomy
No chemotaxonomic information is currently available for the strain 50-1 BONT.
Genome sequencing and annotation
Genome project history
This organism was selected for sequencing on the basis of its phylogenetic position [24], and is part of the Genomic Encyclopedia of Bacteria and Archaea project [25]. The genome project is deposited in the Genome On Line Database [9] and the complete genome sequence is deposited in GenBank. Sequencing, finishing and annotation were performed by the DOE Joint Genome Institute (JGI). A summary of the project information is shown in Table 2.
Table 2. Genome sequencing project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | Finished |
| MIGS-28 | Libraries used | Three genomic libraries: one 454 pyrosequence standard library, one 454 PE library (10 kb insert size), one Illumina library |
| MIGS-29 | Sequencing platforms | Illumina GAii, 454 GS FLX Titanium |
| MIGS-31.2 | Sequencing coverage | 52.1 × Illumina; 35.9 × pyrosequence |
| MIGS-30 | Assemblers | Newbler version 2.3, Velvet, phrap |
| MIGS-32 | Gene calling method | Prodigal 1.4, GenePRIMP |
| INSDC ID | CP002360 | |
| Genbank Date of Release | May 13, 2011 | |
| GOLD ID | GC01760 | |
| NCBI project ID | 42243 | |
| Database: IMG-GEBA | 2503508009 | |
| MIGS-13 | Source material identifier | DSM 15567 |
| Project relevance | Tree of Life, GEBA |
Growth conditions and DNA isolation
M. australiensis 50-1 BONT, DSM 15567, was grown anaerobically in DSMZ medium 339 (Wilkins-Chalgreen anaerobe broth, Oxoid CM 643) [26] at 50°C. DNA was isolated from 0.5-1 g of cell paste using Jetflex Genomic DNA Purification Kit (GENOMED 600100) following the standard protocol as recommended by the manufacturer. Cell lysis was enhanced by adding 20 µl proteinase K for two hours at 58°C. DNA is available through the DNA Bank Network [27].
Genome sequencing and assembly
The genome was sequenced using a combination of Illumina and 454 sequencing platforms. All general aspects of library construction and sequencing can be found at the JGI website [28]. Pyrosequencing reads were assembled using the Newbler assembler (Roche). The initial Newbler assembly consisting of 40 contigs in one scaffold was converted into a phrap [29] assembly by making fake reads from the consensus, to collect the read pairs in the 454 paired end library. Illumina GAii sequencing data (444 Mb) was assembled with Velvet [30] and the consensus sequences were shredded into 1.5 kb overlapped fake reads and assembled together with the 454 data. The 454 draft assembly was based on 108.4 Mb 454 draft data and all of the 454 paired end data. Newbler parameters are -consed -a 50 -l 350 -g -m -ml 20. The Phred/Phrap/Consed software package [29] was used for sequence assembly and quality assessment in the subsequent finishing process. After the shotgun stage, reads were assembled with parallel phrap (High Performance Software, LLC). Possible mis-assemblies were corrected with gapResolution [28], Dupfinisher, or sequencing cloned bridging PCR fragments with subcloning [31]. Gaps between contigs were closed by editing in Consed, by PCR and by Bubble PCR primer walks (J.-F. Chang, unpublished). A total of 279 additional reactions were necessary to close gaps and to raise the quality of the finished sequence. Illumina reads were also used to correct potential base errors and increase consensus quality using a software Polisher developed at JGI [32]. The error rate of the completed genome sequence is less than 1 in 100,000. Together, the combination of the Illumina and 454 sequencing platforms provided 88.0 × coverage of the genome. The final assembly contained 364,783 pyrosequence and 4,541,603 Illumina reads.
Genome annotation
Genes were identified using Prodigal [33] as part of the Oak Ridge National Laboratory genome annotation pipeline, followed by a round of manual curation using the JGI GenePRIMP pipeline [34]. The predicted CDSs were translated and used to search the National Center for Biotechnology Information (NCBI) non-redundant database, UniProt, TIGR-Fam, Pfam, PRIAM, KEGG, COG, and InterPro databases. Additional gene prediction analysis and functional annotation was performed within the Integrated Microbial Genomes - Expert Review (IMG-ER) platform [35].
Genome properties
The genome consists of a 3,135,972 bp long chromosome with a G+C content of 43.5% (Table 3 and Figure 3). Of the 3,033 genes predicted, 2,974 were protein-coding genes, and 59 RNAs; 104 pseudogenes were also identified. The majority of the protein-coding genes (70.4%) were assigned with a putative function while the remaining ones were annotated as hypothetical proteins. The distribution of genes into COGs functional categories is presented in Table 4.
Table 3. Genome Statistics.
| Attribute | Value | % of Total |
|---|---|---|
| Genome size (bp) | 3,135,972 | 100.00% |
| DNA coding region (bp) | 2,822,780 | 90.01% |
| DNA G+C content (bp) | 1,362,640 | 43.45% |
| Number of replicons | 1 | |
| Extrachromosomal elements | 0 | |
| Total genes | 3,033 | 100.00% |
| RNA genes | 59 | 1.95% |
| rRNA operons | 3 | |
| Protein-coding genes | 2,974 | 98.05% |
| Pseudo genes | 104 | 3.43% |
| Genes with function prediction | 2,135 | 70.39% |
| Genes in paralog clusters | 103 | 3.40% |
| Genes assigned to COGs | 2,154 | 71.02% |
| Genes assigned Pfam domains | 2,341 | 77.18% |
| Genes with signal peptides | 596 | 19.65% |
| Genes with transmembrane helices | 813 | 26.81% |
| CRISPR repeats | 2 |
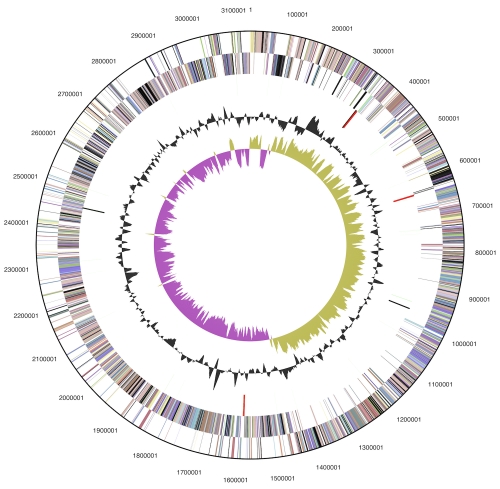
Figure 3.
Graphical circular map of the chromosome. From outside to the center: Genes on forward strand (color by COG categories), Genes on reverse strand (color by COG categories), RNA genes (tRNAs green, rRNAs red, other RNAs black), GC content, GC skew.
Table 4. Number of genes associated with the general COG functional categories.
| Code | value | %age | Description |
|---|---|---|---|
| J | 135 | 5.7 | Translation, ribosomal structure and biogenesis |
| A | 0 | 0.0 | RNA processing and modification |
| K | 164 | 7.0 | Transcription |
| L | 138 | 5.9 | Replication, recombination and repair |
| B | 1 | 0.0 | Chromatin structure and dynamics |
| D | 34 | 1.4 | Cell cycle control, cell division, chromosome partitioning |
| Y | 0 | 0.0 | Nuclear structure |
| V | 59 | 2.5 | Defense mechanisms |
| T | 127 | 5.4 | Signal transduction mechanisms |
| M | 121 | 5.1 | Cell wall/membrane/envelope biogenesis |
| N | 57 | 2.4 | Cell motility |
| Z | 0 | 0.0 | Cytoskeleton |
| W | 0 | 0.0 | Extracellular structures |
| U | 51 | 2.8 | Intracellular trafficking, secretion, and vesicular transport |
| O | 62 | 2.6 | Posttranslational modification, protein turnover, chaperones |
| C | 130 | 5.5 | Energy production and conversion |
| G | 382 | 16.2 | Carbohydrate transport and metabolism |
| E | 160 | 6.8 | Amino acid transport and metabolism |
| F | 63 | 2.7 | Nucleotide transport and metabolism |
| H | 123 | 5.2 | Coenzyme transport and metabolism |
| I | 37 | 1.6 | Lipid transport and metabolism |
| P | 87 | 3.7 | Inorganic ion transport and metabolism |
| Q | 25 | 1.1 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 244 | 10.4 | General function prediction only |
| S | 153 | 6.5 | Function unknown |
| - | 879 | 29.0 | Not in COGs |
Insights from the genome sequence
Comparative genomics
Lacking an available genome sequence of the closest relative of M. australiensis, (Thermoanaerobacterium thermosulfurogenes, Figure 1), the following comparative analyses were done with Thermoanaerobacterium thermosaccharolyticum (GenBank CP002171), the closest related organism with a publicly available genome. While the two genomes are similar in size (M. australiensis 3.1 Mb, 2,974 genes; T. thermosaccharolyticum 2.8 Mb, 2,757 genes), they differ significantly in their G+C content (43% vs. 34%). An estimate of the overall similarity between M. australiensis, T. thermosaccharolyticum and Caldicellulosiruptor saccharolyticus [11] (GenBank EKD00000000.1, as an equidistant outgroup, Figure 1), was generated with the GGDC-Genome-to-Genome Distance Calculator [36,37]. This system calculates the distances by comparing the genomes to obtain HSPs (high-scoring segment pairs) and inferring distances from the set of formulae (1, HSP length / total length; 2, identities / HSP length; 3, identities / total length). Table 5 shows the results of the pair wise comparison between the three genomes.
Table 5. Pairwise comparison of M. australiensis, T. thermosaccharolyticum and C. saccharolyticus using the GGDC-Calculator.
| HSP length / total length [%] |
identities / HSP length [%] |
identities / total length [%] |
||
|---|---|---|---|---|
| M. australiensis | T. thermosaccharolyticum | 2.02 | 86.8 | 1.84 |
| M. australiensis | C. saccharolyticus | 1.16 | 86.9 | 1.01 |
| C. saccharolyticus | T. thermosaccharolyticum | 2.37 | 85.5 | 2.03 |
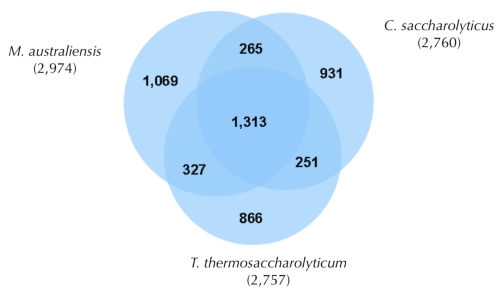
The fraction of shared genes in the three genomes is shown in a Venn diagram (Figure 4). The numbers of pairwise shared genes were calculated with the phylogenetic profiler function of the IMG ER platform [35]. The homologous genes within the genomes were detected with a maximum E-value of 10-5 and a minimum identity of 30%. About half of all the genes in the genomes (1,313 genes) are shared among the three genomes, with equivalent numbers of genes (265 to 327) shared pairwise to the exclusion of the third genome or occurring in only one genome (866 to 1,069). Within the 1,069 unique genes of M. australiensis that have no detectable homologs in the genomes of T. thermosaccharolyticum and C. saccharolyticus (under the sequence similarity thresholds used for the comparison) the 16 genes encoding xylose isomerases appear to be noteworthy; for seven of these isomerase genes no homologs were detected in the other two genomes; only nine genes were identified in C. saccharolyticus, and five in T. thermosaccharolyticum. The high number of xylose isomerise genes suggests a strong utilization of pentoses by M. australiensis. It is already known that several members of the order Thermoanaerobacterales are capable of xylose metabolism [38]. In addition, a number of extracellular solute-binding proteins were found in the genome of M. australiensis. These proteins belong to a high affinity transport system, which is involved in active transport of solutes across the cytoplasmic membrane. The M. australiensis genome contains 54 genes coding for solute-binding proteins belonging to family 1, whereas in C. saccharolyticus and T. thermosaccharolyticum contain only 16 and 13 solute-binding protein family 1 coding genes, respectively.
Figure 4.
Venn diagram depicting the intersections of protein sets (total number of derived protein sequences in parentheses) of M. australiensis, T. thermosaccharolyticum and C. saccharolyticus.
T. thermosaccharolyticum probably transports sugars via a phosphotransferase system (PTS). A total of 29 genes coding for proteins belonging to the PTS specific for different sugars were found in the genome of T. thermosaccharolyticum. The PTS of Thermoanaerobacter tengcongensis was recently studied in detail [39], with 22 proteins identified as participants in the PTS. In contrast, no genes coding for PTS proteins were identified in the genome of M. australiensis, and only one fructose specific PEP-dependent PTS gene was reported in C. saccharolyticus [11]. In conclusion, the number and distribution of these transport mechanisms seems to be highly variable within the Thermoanaerobacteraceae.
Acknowledgements
We would like to gratefully acknowledge the help of Katja Steenblock for growing M. australiensis cultures, and Susanne Schneider for DNA extractions and quality control (both at DSMZ). This work was performed under the auspices of the US Department of Energy Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344, and Los Alamos National Laboratory under contract No. DE-AC02-06NA25396, UT-Battelle and Oak Ridge National Laboratory under contract DE-AC05-00OR22725, as well as German Research Foundation (DFG) INST 599/1-2.
References
- 1.Bonilla Salinas MB, Fardeau ML, Thomas P, Cayol JL, Patel BKC, Ollivier B. Mahella australiensis gen. nov., sp. nov., a moderately thermophilic anaerobic bacterium isolated from an Australian oil well. Int J Syst Evol Microbiol 2004; 54:2169-2173 10.1099/ijs.0.02926-0 [DOI] [PubMed] [Google Scholar]
- 2.Euzéby JP. List of bacterial names with standing in nomenclature: A folder available on the Internet. Int J Syst Bacteriol 1997; 47:590-592 10.1099/00207713-47-2-590 [DOI] [PubMed] [Google Scholar]
- 3.DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie E, Keller K, Huber T, Dalevi D, Hu P, Andersen G. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 2006; 72:5069-5072 10.1128/AEM.03006-05 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Porter MF. An algorithm for suffix stripping. Program: electronic library and information systems. 1980; 14:130-137.
- 5.Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 2000; 17:540-552 [DOI] [PubMed] [Google Scholar]
- 6.Lee C, Grasso C, Sharlow MF. Multiple sequence alignment using partial order graphs. Bioinformatics 2002; 18:452-464 10.1093/bioinformatics/18.3.452 [DOI] [PubMed] [Google Scholar]
- 7.Stamatakis A, Hoover P, Rougemont J. A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 2008; 57:758-771 10.1080/10635150802429642 [DOI] [PubMed] [Google Scholar]
- 8.Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A. How many bootstrap replicates are necessary? Lect Notes Comput Sci 2009; 5541:184-200 10.1007/978-3-642-02008-7_13 [DOI] [PubMed] [Google Scholar]
- 9.Liolios K, Chen IM, Mavromatis K, Tavernarakis N, Hugenholtz P, Markowitz VM, Kyrpides NC. The Genomes On Line Database (GOLD) in 2009: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 2009; 38:D346-D354 10.1093/nar/gkp848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Pitluck S, Yasawong M, Munk C, Nolan M, Lapidus A, Lucas S, Glavina Del Rio T, Tice H, Cheng JF, Bruce D, et al. Complete genome sequence of Thermosediminibacter oceani type strain (JW/IW-1228PT). Stand Genomic Sci 2010; 3:108-116 10.4056/sigs.1133078 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.van de Werken HJ, Verhaart MR, VanFossen AL, Willquist K, Lewis DL, Nichols JD, Goorissen HP, Mongodin EF, Nelson KE, van Niel EW, et al. Hydrogenomics of the extremely thermophilic bacterium Caldicellulosiruptor saccharolyticus. Appl Environ Microbiol 2008; 74:6720-6729 10.1128/AEM.00968-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yarza P, Ludwig W, Euzéby J, Amman R, Schleifer KH, Glöckner FO, Rosselló-Mora R. Updates of the All-Species Living Tree Project based on 16S and 23S rRNA sequence analyses. Syst Appl Microbiol 2010; 33:291-299 10.1016/j.syapm.2010.08.001 [DOI] [PubMed] [Google Scholar]
- 13.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Garrity G. NamesforLife. BrowserTool takes expertise out of the database and puts it right in the browser. Microbiol Today 2010; 37:9 [Google Scholar]
- 15.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Garrity GM, Holt JG. The Road Map to the Manual. In: Garrity GM, Boone DR, Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 1, Springer, New York, 2001, p. 119-169. [Google Scholar]
- 17.Gibbons NE, Murray RGE. Proposals concerning the higher taxa of Bacteria. Int J Syst Bacteriol 1978; 28:1-6 10.1099/00207713-28-1-1 [DOI] [Google Scholar]
- 18.Validation list 132. List of new names and new combinations previously effectively, but not validly, published. Int J Syst Evol Microbiol 2010; 60:469-472 10.1099/ijs.0.022855-0 [DOI] [PubMed] [Google Scholar]
- 19.Rainey FA. Class II. Clostridia class nov. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 3, Springer-Verlag, New York, 2009, p. 736. [Google Scholar]
- 20.Wiegel J. 2009. Order III. Thermoanaerobacterales ord. nov. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 3, Springer-Verlag, New York, p. 1224. [Google Scholar]
- 21.Wiegel J. 2009. Family I. Thermoanaerobacteraceae fam. nov. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 3, Springer-Verlag, New York, p. 1225. [Google Scholar]
- 22.BAuA. Classification of Bacteria and Archaea in risk groups. TRBA 2010; 466:123 [Google Scholar]
- 23.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene Ontology: tool for the unification of biology. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Klenk HP, Göker M. En route to a genome-based classification of Archaea and Bacteria? Syst Appl Microbiol 2010; 33:175-182 10.1016/j.syapm.2010.03.003 [DOI] [PubMed] [Google Scholar]
- 25.Wu D, Hugenholtz P, Mavromatis K, Pukall R, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ, et al. A phylogeny-driven genomic encyclopaedia of Bacteria and Archaea. Nature 2009; 462:1056-1060 10.1038/nature08656 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.List of growth media used at DSMZ: http://www.dsmz.de/microorganisms/media_list.php
- 27.Gemeinholzer B, Dröge G, Zetzsche H, Haszprunar G, Klenk HP, Güntsch A, Berendsohn WG, Wägele JW. The DNA Bank Network: the start from a German initiative. Biopreservation and Biobanking 2011; 9:51-55 10.1089/bio.2010.0029 [DOI] [PubMed] [Google Scholar]
- 28.JGI website. http://www.jgi.doe.gov/
- 29.The Phred/Phrap/Consed software package. http://www.phrap.com
- 30.Zerbino DR, Birney E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 2008; 18:821-829 10.1101/gr.074492.107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Han C, Chain P. Finishing repeat regions automatically with Dupfinisher. In: Proceeding of the 2006 international conference on bioinformatics & computational biology. Arabnia HR, Valafar H (eds), CSREA Press. June 26-29, 2006: 141-146. [Google Scholar]
- 32.Lapidus A, LaButti K, Foster B, Lowry S, Trong S, Goltsman E. POLISHER: An effective tool for using ultra short reads in microbial genome assembly and finishing. AGBT, Marco Island, FL, 2008 [Google Scholar]
- 33.Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 2010; 11:119 10.1186/1471-2105-11-119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pati A, Ivanova NN, Mikhailova N, Ovchinnikova G, Hooper SD, Lykidis A, Kyrpides NC. GenePRIMP: a gene prediction improvement pipeline for prokaryotic genomes. Nat Methods 2010; 7:455-457 10.1038/nmeth.1457 [DOI] [PubMed] [Google Scholar]
- 35.Markowitz VM, Ivanova NN, Chen IMA, Chu K, Kyrpides NC. IMG ER: a system for microbial genome annotation expert review and curation. Bioinformatics 2009; 25:2271-2278 10.1093/bioinformatics/btp393 [DOI] [PubMed] [Google Scholar]
- 36.Auch AF, von Jan M, Klenk HP, Göker M. Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci 2010; 2:117-134 10.4056/sigs.531120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Auch AF, Klenk HP, Göker M. Standard operating procedure for calculating genome-to-genome distances based on high-scoring segment pairs. Stand Genomic Sci 2010; 2:142-148 10.4056/sigs.541628 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Uffen RL. Xylan degradation: a glimpse at microbial diversity. J Ind Microbiol Biotechnol 1997; 19:1-6 10.1038/sj.jim.2900417 [DOI] [Google Scholar]
- 39.Navdaeva V, Zurbriggen A, Waltersperger S, Schneider P, Oberholzer AE, Bähler P, Bächler C, Grieder A, Baumann U, Erni B. Phosphoenolpyruvate: Sugar phosphotransferase system from the hyperthermophilic Thermoanaerobacter tengcongensis. Biochemistry 2011; 50:1184-1193 10.1021/bi101721f [DOI] [PubMed] [Google Scholar]