Figure 1.
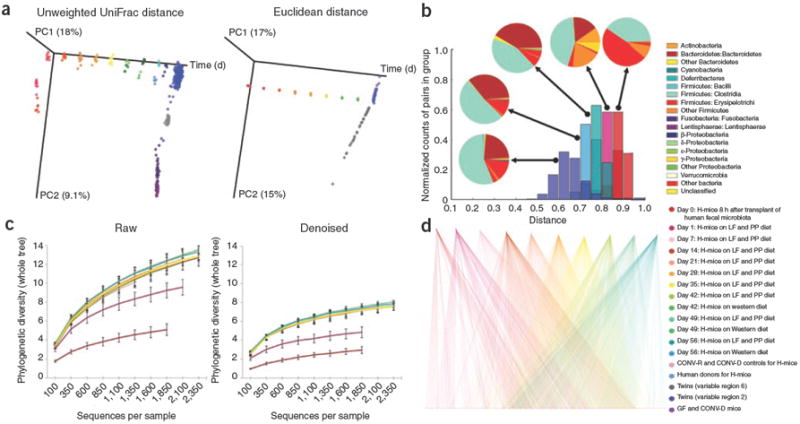
QIIME analyses of the distal gut microbiotas of conventionally raised and conventionalized mice, gnotobiotic mice colonized with a human fecal gut microbiota (H-mice), and human adult mono- and dizygotic twins. (a) Principal coordinates analysis plots for mice, H-mice and twins. Colors correspond to separate samples by species and time point, and are consistent throughout the panels. (b) Unweighted UniFrac distance histograms between the data for fecal microbiota of human twins; human donors for the H-mice study; day 56 post-transplant H-mice on a low-fat (LF) and plant polysaccharide–rich (PP) diet; day 1 H-mice (LF and PP diet); and day 0 H-mice. Taxonomic classifications are presented at the class level. (c) Alpha diversity rarefaction plots of phylogenetic diversity for the H-mice samples. (d) OTU network connectivity of H-mice time series data. CONV-D, conventionalized mice; CONV-R, conventionally raised mice; and GF, germ-free mice.
