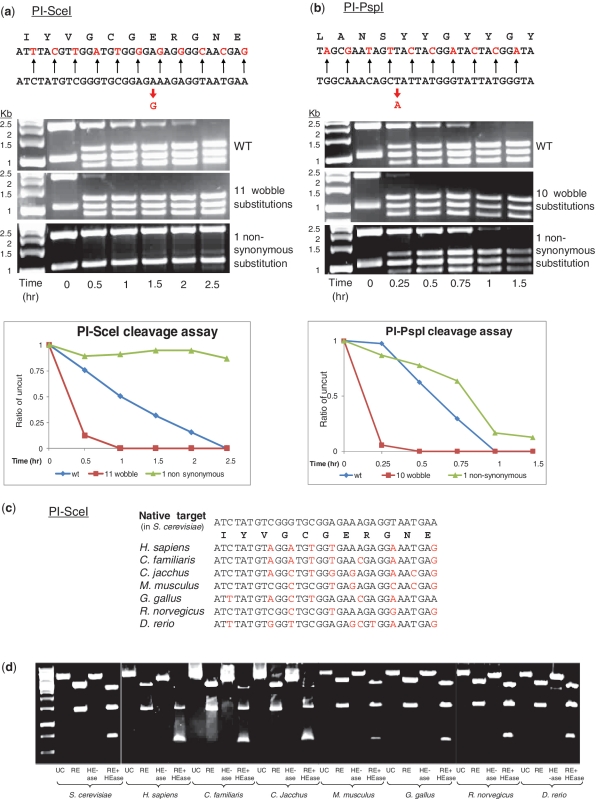
Figure 2.
Characterization of HEase plasticity in target recognition and its use for finding HEase targets in the human and other genomes. (a) and (b): HEase cleavage is tolerant of concomitant mutations at all wobble positions along the target site. The cleavage efficiency of the HEases PI-SceI (from S. cerevisiae) (a) and PI-PspI (from Pyrococcus species GB-D) (b) was assayed on different targets cloned on a bacterial plasmid. The cloned vectors were then fragmented by a restriction enzyme for the sake of visual clarity. Both HEases cleave targets mutated at all wobble positions (top DNA sequence) with higher efficiency than they cleave their native targets (bottom DNA sequence). Conversely, a single non-synonymous mutation (red arrow) is sometimes sufficient to abolish cleavage by PI-SceI and to reduce cleavage by PI-PspI. (c) and (d): PI-SceI can cleave its predicted targets from the human ATP6V1A1 gene and its homologs in the genomes of animal models. (c) Alignment of the native target of the PI-SceI HEase from S. cerevisiae with the predicted targets in the human ATP6V1A1 gene and its homologs in the genomes of animal models. (d) Results of an in vitro cleavage assay demonstrating that PI-SceI can cleave its predicted targets from the genomes of diverse organisms. UC, uncut; RE, cut by a restriction enzyme (XbaI); HEase, cut by an HEase, RE + HEase, cut by XbaI AND by PI-SceI. S. cerevisiae, Saccharomyces cerevisiae; H. sapiens, Homo sapiens; C. familiaris, Canis familiaris; C. jacchus, Callithrix jacchus; M. musculus, Mus musculus; G. gallus, Gallus Gallus; R. norvegicus, Rattus norvegicus; D. rerio, Danio rerio.