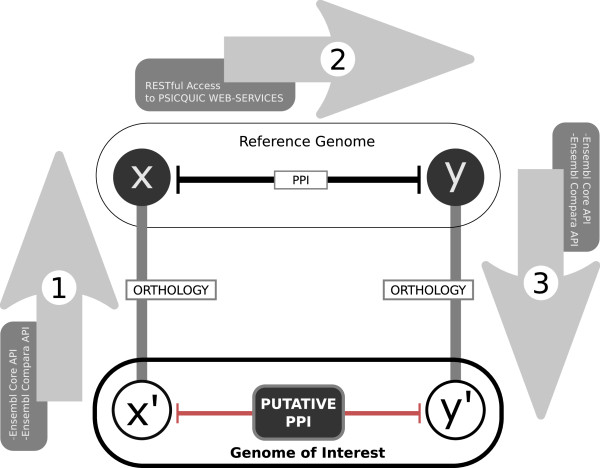
Figure 1.
The Bio::Homology::InterologWalk concept. Schematics illustrating the principle behind interolog mapping. Proteins x and y are known to interact in a reference genome. If they have orthologues x' and y' in the genome of interest, under certain conditions the existence of a putative x' - y' interaction can be assumed. Bio::Homology::InterologWalk implements this in a three-steps algorithm. 1. get orthologues of genes of interest in reference genome(s). queries the initial gene list against one or more Ensembl databases and retrieves their orthologues. Options can be set to specify stringency of retrieved hits. Ancillary data fields are computed. 2. get interactions in reference genome(s). queries the orthology list built in (1.) against PSICQUIC-enabled PPI databases using REST. This step will enrich the dataset built in (1.) with the interactors of those orthologues, if any, plus ancillary data -- including parameters describing the nature and origin of the annotated interaction. 3. get orthologues from reference genome(s) back in genome of interest. In this step the interactor list built in (2.) is queried against one or more Ensembl databases (again using the Ensembl Perl API) to find orthologues back in the original genome of interest. As in (1.), a number of supplementary information fields are computed.