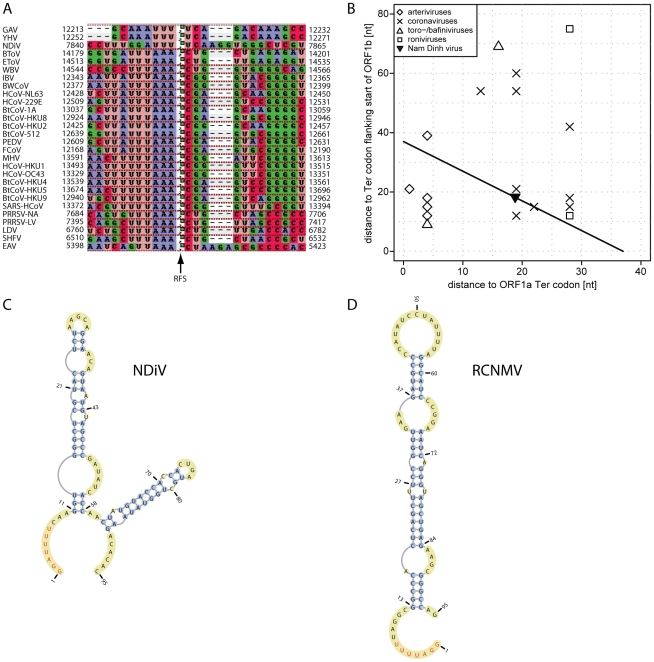
Figure 4. ORF1a/ORF1b ribosomal frameshifting in the NDiV genome.
(A) The nucleotide alignment around the open reading frame (ORF) 1a/1b ribosomal −1 frameshift (RFS) in 28 nidovirus species. The alignment column marked with an arrow indicates the RFS position and precedes a nucleotide which is read twice, as the third nucleotide of an ORF1a codon and the first coding nucleotide of ORF1b, respectively. In NDiV this residue is predicted to be located at genome position 7851. Numbers flanking the alignment indicate genomic positions. (B) RFS position in the ORF1a/ORF1b overlap of nidoviruses. The RFS position in selected nidoviruses is plotted as distances between RFS and termination codons flanking ORF1b and ORF1a from upstream and downstream, respectively. In NDiV, the predicted RFS (filled triangle) and all other candidate positions are located on black line in the 40 nt ORF1a/ORF1b overlap. (C and D) RNA secondary structure of sequence fragments around of the ORF1a/ORF1b overlap in NDiV and Red clover necrotic mosaic virus (RCNMV) as predicted by pknotsRG [30]. The putative slippery sequences are in red (see also A). The predicted stem loop in RCNMV closely matches the one presented in [31].