Abstract
Pitipeptolides A (1) and B (2) are cyclic depsipeptides isolated from the marine cyanobacterium Lyngbya majuscula from Piti Bomb Holes, Guam. Additional analogues have now been isolated by revisiting larger collections of the same cyanobacterium. The four newly identified analogues, pitipeptolides C–F (3–6), are the tetrahydro analogue (3), an analogue with a lower degree of methylation (4) as well as two homologues (5 and 6) of pitipeptolide A. The structures were elucidated using 2D NMR experiments, chiral HPLC analysis and comparison with pitipeptolide A. The newly identified analogues showed weaker cytotoxic activities compared to the two major parent compounds, pitipeptolides A (1) and B (2), against HT-29 colon adenocarcinoma and MCF7 breast cancer cells. On the other hand, pitipeptolide F (6) was the most potent pitipeptolide in a disc diffusion assay against Mycobacterium tuberculosis. The latter finding suggests that the structure of pitipeptolides could be optimized for selective antibacterial activity.
Keywords: Marine cyanobacteria, Lyngbya majuscula, pitipeptolides, cytotoxin, antimycobacterial activity
1. Introduction
The marine environment is a largely unexplored natural resource that covers most of the earth’s surface. The first Census of Marine Life (CoML 2000-2010) has just completed a decade-long inventory that exposed some of the ocean’s secrets and reflected its uncharted massive biodiversity (O’Dor et al., 2010). From a pharmaceutical perspective, this resource yields several classes of secondary metabolites that cover biologically relevant chemical space. In particular, marine microbes are important producers of these bioactive secondary metabolites (Bhatnagar and Kim, 2010). Among marine microbes, cyanobacteria are one of the most promising groups (Burja et al., 2001); and within the cyanobacteria, the genus Lyngbya is the most prolific producer of bioactive compounds, including cytotoxic, antimicrobial and antifungal cyclic peptides (Tan, 2007; Liu and Rein, 2010).
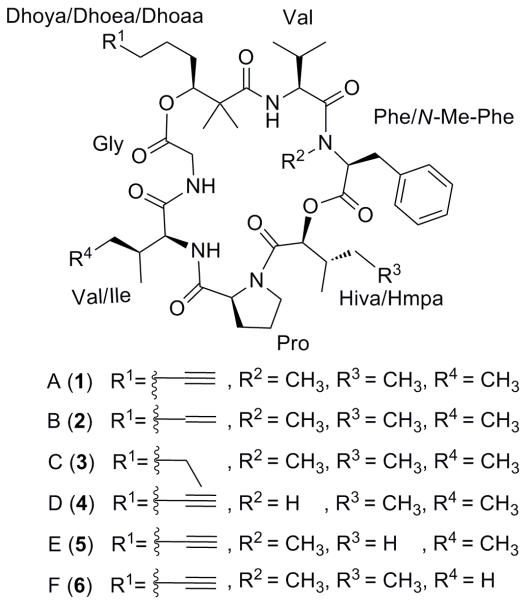
Pitipeptolides A (1) and B (2) are cyclic depsipeptides isolated from a Guamanian sample of the marine cyanobacterium Lyngbya majuscula (Luesch et al., 2001). Both compounds contain standard proteinogenic as well as N-methylated amino acids, an α-hydroxy acid, and are characterized by the presence of the unique β-hydroxy fatty acid units 2,2-dimethyl-3-hydroxy-7-octynoic (Dhoya) or 2,2-dimethyl-3-hydroxy-7-octenoic (Dhoea) acids (Figure 1). Both compounds were reported to possess weak cytotoxic and moderate antimycobacterial activities. Moreover, the total synthesis of pitipeptolide A was achieved, which confirmed its structure and the assigned configuration (Peng et al., 2005). Revisiting the cyanobacterium on a larger scale led to the identification of the recently reported proline-rich cyclic depsipeptide pitiprolamide (Montaser et al., 2010) and now four more pitipeptolide analogues named pitipeptolides C–F (3–6). Here we report the isolation, structure determination as well as the biological characterization of compounds 3–6.
Figure 1.
Structures of pitipeptolides A–F (1–6).
2. Results and Discussion
The cyanobacterial sample was extracted three times with EtOAc–MeOH mixtures. This organic extract was subjected to solvent partitioning steps, yielding 7.2 g of a semi-polar n-BuOH fraction. The n-BuOH fraction was fractionated by silica gel chromatography. The presence of pitipeptolides in one fraction was identified by 1H NMR, and subsequent purification of this fraction by reversed-phase HPLC yielded pitipeptolides 3–6.
The 1H NMR spectrum of pitipeptolide C (3) indicated close structural similarity to pitipeptolide A (1). HRESIMS suggested a molecular formula of C44H70N5O9 (m/z 812.5178 for [M + H]+), which is four mass units higher than that of 1. Furthermore, the acetylenic carbons were absent in the 13C NMR for this analogue, and two carbon resonances for an additional methylene as well as a methyl carbon were present instead. All this data indicated that 3 is a tetrahydro- analogue of pitipeptolide A (1), containing the completely saturated fatty acid derived unit 2,2-dimethyl-3-hydroxy octanoic acid (Dhoaa).
Pitipeptolide D (4) had the molecular formula C43H63N5O9 as suggested by the HRESIMS data (m/z 816.4524 for [M + Na]+). This molecular formula suggested a pitipeptolide A analogue with a lower degree of methylation. Further analysis of 1H and 13C NMR spectra showed that the Phe residue in 4 lacks the N-Me modification. The additional amide proton resonates as a doublet at δH 6.21 ppm (Table 1). No ROESY correlation was detected between the two α-protons in the Val (δH 4.17 ppm) and Phe (δH 4.25 ppm) units, while a ROESY correlation was evident between the α-proton in Val (δH 4.17 ppm) and the amide proton in Phe (δH 6.21 ppm), indicating a trans peptide bond. The absence of the N-Me group did not influence the amide linkage’s conformation; the ROESY correlation between the two α-protons in the Val and Phe units was also missing in compound 3, and a strong ROESY correlation was obvious between the α-proton in Val (δH 4.71 ppm) and the N-methyl group (δH 2.78 ppm) in N-Me Phe, which points to a trans conformation for the amide linkage between both units.
Table 1.
NMR spectroscopic data for pitipeptolides C (3) and D (4) in CDCl3 (δ in ppm, J in Hz) at 600 MHz
| Unit | C/H No. | Pitipeptolide C (3) | Pitipeptolide D (4) | ||||
|---|---|---|---|---|---|---|---|
| δ C | δH (J) | HMBC a | δ C | δH (J) | HMBC a | ||
| Dhoaab/Dhoyac | 1 | 175.7 qC | – | 3, 2 (Val), 9, 10, NH (Val) | 175.9 qC | – | 3, 2 (Val), 9, 10, NH (Val) |
| 2 | 45.7 qC | – | 3, 9, 10 | 46.0 qC | – | 3, 9, 10 | |
| 3 | 77.6 CH | 4.92 dd (9.3, 3.2) | 9, 10 | 77.3 CH | 5.11 dd (10.2, 2.3) | 4, 5, 9, 10 | |
| 4 | 30.3 CH2 | 1.56 m 1.45 m |
– | 28.4 CH2 | 1.79 m 1.53 m |
2, 5, 6 | |
| 5 | 25.7 CH2 | 1.23 m | 3, 4 | 24.4 CH2 | 1.43 m | 3, 4, 6 | |
| 6 | 31.9 CH2 | 1.24 m | 4, 7, 8 | 18.0 CH2 | 2.27 m 2.19 m |
3, 4, 5 | |
| 7 | 22.5 CH2 | 1.29 m | 6, 8 | 83.6 qC | – | 5, 6, 8 | |
| 8 | 14.4 CH3 | 0.88 t (7.0) | 7 | 69.3 CH | 1.96 t (2.6) | 6 | |
| 9 | 19.8 CH3 | 1.28 s | 3, 10 | 18.5 CH3 | 1.25 s | 3, 10 | |
| 10 | 23.1 CH3 | 1.13 s | 9 | 23.8 CH3 | 1.16 s | 3, 9 | |
| Val | 1 | 172.1 qC | – | 2, 3 (Dhoaa), 10 (N-Me- Phe) |
171.5 qC | – | 2, 3, NH, NH (Phe) |
| 2 | 53.4 CH | 4.71 dd (9.4 , 1.8) | 5, NH | 57.6 CH | 4.17 dd (8.6, 4.2) | 3, 4, 5, NH | |
| 3 | 29.8 CH | 1.75 m | 2, 5 | 30.6 CH | 1.89 m | 2, 4, 5 | |
| 4 | 16.1 CH3 | 1.00 d (6.8) | 2, 3, 5 | 17.0 CH3 | 0.79 d (6.7) | 2, 3, 5 | |
| 5 | 20.5 CH3 | 0.89 d (6.7) | 2, 3, 4 | 19.7 CH3 | 0.78 d (6.7) | 2, 3, 4 | |
| NH | – | 6.11 d (9.1) | – | 5.90 d (8.7) | – | ||
| N-Me-Pheb/Phec | 1 | 172.7 qC | – | 2 | 172.6 qC | – | 2, 3, 2 (Hmpa), NH |
| 2 | 65.9 CH | 3.84 dd (11.3, 3.9) | 3, 10 | 55.0 CH | 4.25 ddd (8.2, 6.6, 4.1) | 3, NH | |
| 3 | 34.1 CH2 | 3.20 dd (14.3, 3.7) 3.12 dd (14.3, 11.3) |
2, 5/9 | 35.3 CH2 | 3.27 dd (14.5, 4.1) 3.11 dd (14.5, 10.8) |
2, 5/6 | |
| 4 | 137.8 qC | – | 2, 3, 6/8 | 137.3 qC | – | 2, 3, 6/8 | |
| 5/9 | 129.2 CH | 7.11 d (7.1) | 3, 6/8, 7 | 129.4 CH | 7.14 d (7.2) | 3, 6/8 | |
| 6/8 | 128.9 CH | 7.28 m | 5/9, 7 | 128.8 CH | 7.29 m | 5/6, 7 | |
| 7 | 127.2 CH | 7.24 m | 5/9 | 127.1 CH | 7.24 m | 5/9, 6/8 | |
| 10 | 39.4 CH3 | 2.78 s | 2 | – | – | – | |
| NH | – | – | – | – | 6.21 d (6.6) | – | |
| Hmpa | 1 | 169.8 qC | – | 2 | 169.3 qC | – | 2 |
| 2 | 78.4 CH | 4.94 d (7.3) | 6 | 78.3 CH | 4.91 d (6.5) | 3, 4, 6 | |
| 3 | 37.4 CH | 1.81 m | 2, 4, 5, 6 | 37.3 CH | 1.84 m | 2, 4, 5, 6 | |
| 4 | 25.2 CH2 | 1.58 m 1.15 m |
2, 5, 6 | 25.5 CH2 | 1.21 m 1.66 m |
2, 5, 6 | |
| 5 | 11.9 CH3 | 0.89 t (6.7) | 4 | 11.0 CH3 | 0.90 t (7.4) | 3, 4 | |
| 6 | 14.6 CH3 | 0.93 d (6.9) | 2, 4 | 14.7 CH3 | 0.97 d (7.2) | 3, 4 | |
| Pro | 1 | 170.4 qC | – | 2 | 170.4 qC | – | 2, NH (Ile) |
| 2 | 61.4 CH | 4.63 d (6.9) | 3, 4 | 61.2 CH | 4.59 d (7.5) | 3, 4 | |
| 3 | 31.4 CH2 | 2.67 m 1.93 m |
2, 4, 5 | 31.5 CH2 | 2.60 m 1.99 m |
2, 5 | |
| 4 | 21.9 CH2 | 1.97 m 1.76 m |
2, 3, 5 | 21.8 CH2 | 2.00 m 1.77 m |
2, 5, 6 | |
| 5 | 46.5 CH2 | 3.70 m 3.54 dd (10.3, 9.1) |
2, 3, 4 | 46.6 CH2 | 3.70 m 3.58 dd (11.5, 9.8) |
2, 3 | |
| Ile | 1 | 171.9 qC | – | 2, 2 (Gly), NH (Gly) | 171.4 qC | – | 2, 2 (Gly), NH (Gly) |
| 2 | 61.1 CH | 4.22 dd (8.7, 8.6) | 3, 6, NH | 60.5 CH | 4.25 dd (8.6, 8.1) | 6 | |
| 3 | 35.3 CH | 2.05 m | 2 | 35.6 CH | 2.05 m | 2, 4, 5, 6 | |
| 4 | 25.9 CH2 | 1.58 m 1.23 m |
2, 6 | 25.5 CH2 | 1.58 m 1.21 m |
2, 5, 6 | |
| 5 | 11.1 CH3 | 0.87 t (7.0) | 4 | 11.0 CH3 | 0.89 t (7.5) | 4 | |
| 6 | 16.1 CH3 | 1.00 d (6.8) | 2, 4 | 14.8 CH3 | 0.98 d (7.2) | 4 | |
| NH | – | 7.97 d (8.4) | – | – | 7.64 d (8.1) | – | |
| Gly | 1 | 170.2 qC | – | 2, 3 (Dhoaa), NH (Ile) | 169.9 qC | – | 2, 3 (Dhoya) |
| 2 | 41.3 CH2 | 4.60 dd (17.8, 9.3) 3.97 d (17.8) |
– | 41.4 CH2 | 4.34 dd (18.0, 8.6) 3.95 dd (18.0, 2.0) |
– | |
| NH | – | 6.38 d (8.9) | – | – | 6.50 d (7.4) | – | |
Protons showing long-range correlation to indicated carbon
Refers to compound 3
Refers to compound 4
The two constitutional isomers pitipeptolides E (5) and F (6) co-eluted during the first round of reversed-phase HPLC purification of the silica gel chromatography fraction containing pitipeptolides. Both compounds showed 1H NMR spectra very similar to each other and to 1. Moreover, HRESIMS (m/z 816.4521 for [M + Na]+ for 5; m/z 816.4518 for [M + Na]+ for 6) showed that both compounds have the same molecular formula as pitipeptolide D (4) and are additional pitipeptolide A analogues with one less methylene group. Further analysis of the NMR spectra showed that 5 had Hmpa→Hiva displacement, while 6 had Ile→Val displacement compared to 1. The structures of compounds 3–6 were further confirmed by analyzing their MS/MS fragmentation patterns (Figure 2).
Figure 2.
ESI-MS/MS fragmentation patterns of pitipeptolides C–F (3–6).
The absolute configurations of the amino and α-hydroxy acids in 3–6 were determined using chiral HPLC after acid hydrolysis. The configurations of the fatty acid derived units were determined by comparing NMR and optical rotation data with those of pitipeptolide A (1). The closely matching NMR chemical shifts and optical rotations of all the analogues indicated the same relative and absolute configuration. All chiral centers in 3–6 had S configuration.
Pitipeptolides A (1) and B (2) have been previously shown to possess weak cytotoxicity towards cancer cells as well as moderate antimycobacterial activities (Luesch et al., 2001). It is unclear if the relevant targets in eukaryotic and prokaryotic cells are related or the same. However, compounds 3–6 appeared to be less cytotoxic against cancer cells than 1 and 2 in HT-29 colon adenocarcinoma and MCF7 breast cancer cell lines (Table 3). The same structural features required for cytotoxic activity against cancer cells do not appear to be critical for the antimycobacterial effect, since compounds 3 and 5 showed similar antimycobacterial activities compared to 1 and 2, and strikingly, compound 6 showed the highest potency in the disc diffusion assay against Mycobacterium tuberculosis. Notably, 4 lacked significant activities against both mammalian and bacterial cells. The above findings lead to the following structure-activity relationship conclusions: (1) N-methylation in the Phe unit is important for both cytotoxic and antibacterial activities; (2) the π system in the fatty acid unit is one of the essential features for the cytotoxic activity in mammalian cells, but it is not essential for the antibacterial activity; (3) decreasing the hydrophobicity of certain units (Hmpa→Hiva and Ile→Val) decreased the anticancer activity (5 and 6), while on the other hand, particularly 6 possessed increased antibacterial potency. This indicates that the cytotoxic activity in cancer cells could be separated from the antimycobacterial activity, and the selectivity of the antibacterial compounds could be increased through structural modifications. Pitipeptolides did not show any significant antibacterial activities against either the Gram negative bacterium Pseudomonas aeruginosa or the Gram positive bacteria Bacillus cereus and Staphylococcus aureus.
Table 3.
Cytotoxicity in cancer cells and antimycobacterial activities of pitipeptolides A–F (1–6)
| Antimycobacterial Activity | Cytotoxicity IC50( μM)a | ||||
|---|---|---|---|---|---|
| Diameter of Zone of Inhibition (mm)a | HT-29 | MCF7 | |||
| 100 μg | 50 μg | 10 μg | |||
| Pitipeptolide A (1) | 28 | 23 | 9 | 13 | 13 |
| Pitipeptolide B (2) | 30 | 24 | 14 | 13 | 11 |
| Pitipeptolide C (3) | 26 | 21 | 18 | 67 | 73 |
| Pitipeptolide D (4) | 10 | 0 | 0 | > 100 | > 100 |
| Pitipeptolide E (5) | 21 | 15 | 0 | 75 | > 100 |
| Pitipeptolide F (6) | 40 | 30 | 10 | 87 | 83 |
| 10 μg | 5 μg | 1 μg | |||
| Streptomycin | 40 | 30 | 0 | ||
| Paclitaxel | 0.007 | 0.006 | |||
n = 2
The structures of pitipeptolides suggest they are biosynthesized by a mixed polyketide synthase/nonribosomal peptide synthetase (PKS/NRPS) pathway. N-methylation is one of the modifications characterizing non-ribosomally synthesized peptides. This modification plays an important role in protecting the peptide bond against proteolytic cleavage, and thus influences the biological activity of the peptide (Schaller, 1997; Marahiel et al., 1997). Thus, cellular stability might be one explanation for the much reduced activities in all our assays of pitipeptolide D (4) which lacks this modification. Non-ribosomal peptide synthetases are also characterized by moderate substrate specificity, which results in structural diversity. This specificity can even be lost when it comes to discrimination between two similar substrates (Marahiel et al., 1997; Magarvey et al., 2006), which may be the reason for the simultaneous production of the two analogues 5 and 6 that differ from 1 by only one methylene group.
The abundance of pitipeptolides in this sample suggests that this class of cyclic depsipeptides may play an important ecological role. Pitipeptolide A (1) is the major depsipeptide in this series (378 mg, 0.14% of total dry weight) followed by pitipeptolide B (2) (70.5 mg, 0.024% of total dry weight). The production of pitipeptolides C, E and F (3, 5 and 6) (23 mg, 30 mg and 26 mg, respectively, ≈ 0.009% of total dry weight each) is about 16 times lower than that of 1, and only 1.1 mg (≈ 0.0004% of total dry weight) of pitipeptolide D (4) was isolated, possibly indicating that 1 is the most beneficial analogue for this organism. In accordance with this conclusion, pitipeptolide A was previously reported as a defensive secondary metabolite that acts as a feeding deterrent at natural concentrations against a range of marine grazers (Cruz-Rivera and Paul, 2007).
Thacker and Paul (2004) have shown that despite the isolation of a plethora of novel secondary metabolites from different globally distributed collections of L. majuscula, these chemical compounds could not be strictly associated to specific geographical regions. In accordance, we noticed a close chemical relationship between this population of L. majuscula from Guam and another collection from Madagascar that yielded the cyclodepsipeptides antanapeptins (Nogle and Gerwick, 2002). The collection from Madagascar has yielded dolastatin 16 as well as antanapeptins A–D. Similarly, this collection from Guam has yielded the dolastatin 16 analogue pitiprolamide and pitipeptolides A–F (1–6); the latter show structural homology to antanapeptins. Moreover, the structural differences between antanapeptin A–D are the same as between pitipeptolides A–C (1–3) and E (5), summarized as degrees of unsaturation in the fatty acid units and an Ile→Val displacement. The structural homology between dolastatin 16 and pitiprolamide has been noted before (Montaser et al, 2010). For simplicity, we compared the structural differences between antanapeptin A (7) and pitipeptolide A (1) (Figure 3), which could be summarized as follows: (1) while both compounds have α,β-substituted octynoic acid units at the same position, 7 has a mono-methyl fatty acid derivative (Hmoya) instead of a dimethyl-fatty acid unit (Dhoya) in 1; (2) 7 incorporates a Hiva unit instead of Hmpa in 1 which is a common displacement as mentioned before, and therefore this particular difference probably does not require a genetic variation, (3) the biosynthetic gene cluster responsible for 7 is expected to have an additional N-methyltransferase domain associated with the Ile module, (4) 7 lacks the Gly unit, giving a hexa-cyclodepsipeptide instead of the hepta-cyclodepsipeptide as in 1. Because of these differences between both groups of depsipeptides, antanapeptins may have failed to show any toxicity against brine shrimp or any antibacterial activities (Nogle and Gerwick, 2002).
Figure 3.
Structural comparison between pitipeptolide A (1) and antanapeptin A (7). Subunit sequences are shown below the structures; dotted lines indicate connected residues. The absolute configuration of the Hmoya unit in 7 is still unknown.
The recently reported marine cyanobacterial depsipeptides cocosamides A and B (Gunasekera et al., 2011) represent another example of cyanobacterial metabolites that share some structural features with pitipeptolides. Cocosamides were isolated from the marine cyanobacterium Lyngbya majuscula from Cocos Lagoon,Guam, and showed weaker cytotoxic activities against HT-29 and MCF7 cancer cell lines than pitipeptolides.
3. Conclusion
A population of the cyanobacterium Lyngbya majuscula from Piti Bomb Holes in Guam produces a class of bioactive cyclodepsipeptides named pitipeptolides. In addition to the previously reported major analogues pitipeptolides A (1) and B (2), the cyanobacterium contained more analogues, four of which are reported here (3–6). All pitipeptolide analogues have minor structural differences. However, the structural variations gave a better insight into the contribution of some structural features to cancer cytotoxic and antimycobacterial activities.
Pitipeptolides might prove to be privileged structures that could interact with different biological targets. Consequently, those compounds showed several biological activities besides their ecological role. Despite different environmental and geographical regions, chemical similarity was detected between this Guamanian sample and another L. majuscula sample from Madagascar. The structural similarity between the isolated secondary metabolites from both samples suggests genetic similarity, including the putative gene cluster encoding biosynthetic enzymes, and pitipeptolides could be considered analogues of antanapeptins. Notably, those structural modifications uniquely granted the pitipeptolides cytotoxic and antibacterial activities that were not found in the antanapeptins. Therefore, we could consider pitipeptolides as lead compounds and we expect further structural optimization to selectively enhance their cytotoxic or antibacterial activities.
4. Experimental
4.1 General experimental procedures
Optical rotation was measured on a Perkin-Elmer 341 polarimeter. UV and optical density were measured on a SpectraMax M5 (Molecular Devices), and IR data were obtained on a Perkin-Elmer Spectrum One FT-IR Spectrometer. 13C NMR spectrum was recorded on a Bruker 500 MHz spectrometer operating at 125 MHz. 1H and 2D NMR spectra were recorded on a Bruker Avance II 600 MHz spectrometer. All spectra were obtained in CDCl3 using residual solvent signals (δH 7.26, δC 77.16 ppm) as internal standards. HSQC and HMBC experiments were optimized for 1JCH = 145 and 1JCH = 7 Hz, respectively. HRMS data was recorded on an Agilent LC-TOF mass spectrometer equipped with an APCI/ESI multimode ion source detector in positive ion mode. LC-MS data were obtained using an API 3200 triple quadrupole MS (Applied Biosystems) equipped with a Shimadzu LC system. ESIMS fragmentation data were recorded on an API 3200 by direct injection with a syringe driver. 2-Hydroxy isovaleric acid (Hiva) standards were purchased from Sigma-Aldrich. 2-Hydroxy 3-methyl pentanoic acid (Hmpa) standards were synthesized from isoleucine (Van Draanen et al., 1991).
4.2 Marine cyanobacterial sample
The sample of the marine cyanobacterium Lyngbya majuscula (recollection of UOG strain VP627) was collected at Piti Bomb Holes, Guam, in February 2000. A voucher sample has been preserved at the Smithsonian Marine Station at Fort Pierce, FL.
4.3 Extraction and isolation
The freeze-dried organism was extracted three times with EtOAc–MeOH (1:1) to afford the crude organic extract (35.5 g). The resulting extract was partitioned between hexanes and 80% aqueous MeOH; the methanolic phase was concentrated to dryness, and the residue was further partitioned between n-BuOH and H2O. After concentrating the n-BuOH extract in vacuo, the resulting residue (7.2 g) was subjected to flash chromatography over silica gel, eluting with CH2Cl2 followed by increasing gradients of i-PrOH in CH2Cl2, and finally with MeOH. The fraction eluting with 4% i-PrOH in CH2Cl2 was fractionated on a semipreparative reversed-phase HPLC column (YMC-Pack ODS-AQ, 250 × 10 mm, 5 μm, 2 mL/min; UV detection at 220/254 nm) using a MeOH/H2O linear gradient (75% to 100% over 30 min, and then 100% MeOH for 10 min) to yield 10 collected fractions. Pitipeptolides A (1) (378 mg) and B (2) (70.5 mg) eluted at tR 21.3 min (fraction 4) and tR 23.8 min (fraction 7), respectively. Pitipeptolide C (3) (23 mg) eluted at tR 25 min as a single peak (fraction 9). Further purification of fractions 2 and 3 gave compounds 4, 5 and 6 (see below).
Pitipeptolide D (4) was further purified from fraction 2 (eluting between tR 19.2 to 20.2 min using the conditions mentioned above) using semipreparative reversed-phase HPLC (Luna C18, 250 × 10 mm, 5 μm, 2.0 mL/min; UV detection at 200/220 nm) with a MeOH/H2O linear gradient (75% to 100% aqueous MeOH over 20 minutes then 100% MeOH for 10 min). The peak eluting at tR 19.9 min was subjected to further purification on another semipreparative reversed-phase HPLC column (Phenomenex Phenylhexyl, 250 × 10 mm, 5 μm, 2.0 mL/min; UV detection at 220/200 nm) using a MeOH/H2O linear gradient (85% to 100% aqueous MeOH over 35 min, and then 100% MeOH for 10 min) to afford 1.1 mg of the pure compound at tR 20.1 min.
Pitipeptolides E (5) and F (6) co-eluted at tR 20.6 min (fraction 3 from the first HPLC run mentioned above on the YMC-Pack ODS-AQ column) to give a total of 68 mg of an impure fraction. 5 mg of this mixture was further purified several times on an analytical reversed-phase HPLC column (Allure Restec C18, 250 × 4.6 mm, 5 μm, 1.0 mL/min; UV detection at 220/200 nm) using a MeOH/H2O linear gradient (75% to 100% aqueous MeOH over 35 min, and then 100% MeOH for 10 min) to yield compound 5 at tR 12.2 min (2.2 mg), and compound 6 at tR 12.7 min (1.9 mg). Extrapolating those values suggests a total of about 30 mg of 5 and 26 mg of 6 in this sample.
4.4 Pitipeptolide C (3)
Colorless, amorphous solid; [α]20D –121° (c 0.11, MeOH); UV (MeOH) λmax (log ε) 202 (4.57) nm; IR (film) νmax 3403, 2961, 2932, 2874, 1727, 1653, 1511, 1464, 1415, 1371, 1188, 1030, 736, 702 cm−1 ; 1H NMR, 13C NMR and HMBC data, see Table 1; HRESI/APCIMS m/z 812.5178 for [M + H]+ (calcd for C44H70N5O9 812.5168).
4.5 Pitipeptolide D (4)
Colorless, amorphous solid; [α]20D –112° (c 0.12, MeOH); UV (MeOH) λmax (log ε) 202 (4.29) nm; IR (film) νmax 3410, 2966, 2935, 2876, 1735, 1654, 1513, 1455, 1370, 1180, 1039, 735, 701 cm−1 ; 1H NMR, 13C NMR and HMBC data, see Table 1; HRESI/APCIMS m/z 816.4524 for [M + Na]+ (calcd for C43H63N5O9 816.4518).
4.6 Pitipeptolide E (5)
Colorless, amorphous solid; [α]20D –105° (c 0.13, MeOH); UV (MeOH) λmax (log ε) 202 (4.34) nm; IR (film) νmax 2967, 2372, 2343, 2297, 1986, 1655, 1512, 1175, 804, 702 cm−1 ; 1H NMR, 13C NMR and HMBC data, see Table 2; HRESI/APCIMS m/z 816.4521 for [M + Na]+ (calcd for C43H63N5O9Na 816.4518).
Table 2.
NMR spectroscopic data for pitipeptolides E (5) and F (6) in CDCl3 (δ in ppm, J in Hz) at 600 MHz
| Unit | C/H No. | Pitipeptolide E (5) | Pitipeptolide F (6) | ||||
|---|---|---|---|---|---|---|---|
| δ C | δH (J) | HMBC a | δ C | δH (J) | HMBC a | ||
| Dhoya | 1 | 175.2 qC | – | 2 (Val), 3, 9, 10, NH (Val) |
175.3 qC | – | 2 (Val), 3, 9, 19, NH (Val) |
| 2 | 45.2 qC | – | 3, 4, 9, 10 | 45.3 qC | – | 9, 10 | |
| 3 | 77.1 CH | 4.97 dd (9.3, 2.9) | 5, 9 | 77.2 CH | 4.95 dd (9.6, 2.0) | 1 (Gly), 1, 2, 4, 9, 10 | |
| 4 | 28.8 CH2 | 1.82 m 1.58 m |
3, 5 | 28.9 CH2 | 1.83 m 1.48 m |
3, 5, 6 | |
| 5 | 24.4 CH2 | 1.43 m | 3, 4, 6 | 24.4 CH2 | 1.43 m | 3, 4, 6 | |
| 6 | 18.1 CH2 | 2.22 m | 3, 5, 7 | 18.1 CH2 | 2.22 m | 5 | |
| 7 | 83.6 qC | – | 5, 6, 8 | 83.4 qC | – | 5, 6, 8 | |
| 8 | 69.3 CH | 1.97 t (2.6) | 6 | 69.4 CH | 1.96 t (2.6) | 6, 7 | |
| 9 | 19.4 CH3 | 1.30 s | 10 | 19.4 CH3 | 1.25 s | 3, 10 | |
| 10 | 23.0 CH3 | 1.16 s | 3, 5, 9 | 23.0 CH3 | 1.16 s | 3, 9 | |
| Val | 1 | 171.6 qC | – | 2, 3, NH, 10 (N-Me- Phe) |
171.7 qC | – | 2, 3, NH, 10 (N-Me- Phe) |
| 2 | 53.4 CH | 4.69 dd (9.6 , 1.8) | 3, 4, 5, NH | 53.5 CH | 4.70 dd (9.0, 1.4) | 4, 5, NH | |
| 3 | 29.5 CH | 1.76 m | 2, 4, 5 | 29.5 CH | 1.76 m | 2, 4, 5 | |
| 4 | 16.0 CH3 | 0.90 d* | 2, 3, 5 | 16.0 CH3 | 0.90 d (7.1) | 2, 3, 5 | |
| 5 | 20.3 CH3 | 0.89 d* | 2, 3, 4 | 20.5 CH3 | 0.89 d (6.9) | 2, 3, 4 | |
| NH | – | 6.06 d (8.9) | – | – | 6.09 d (9.0) | – | |
| N-Me-Phe | 1 | 172.3 qC | – | 2, 3 | 171.8 qC | – | 2 |
| 2 | 65.9 CH | 3.86 dd (11.6, 3.2) | 3, 10 | 65.9 CH | 3.85 dd (11.2, 3.4) | 3, 10 | |
| 3 | 33.9 CH2 | 3.21 dd (14.2, 3.5) 3.12 dd (14.2, 11.2) |
2, 5/9 | 34.0 CH2 | 3.21 dd (14.4, 3.3) 3.11 dd (14.4, 11.4) |
1, 2, 5/9 | |
| 4 | 137.3 qC | – | 2, 3, 6/8 | 137.3 qC | – | 2, 3, 6/8 | |
| 5/9 | 129.2 CH | 7.12 d (7.5) | 6/8, 7 | 129.4 CH | 7.10 d (7.2) | 6/8 | |
| 6/8 | 128.9 CH | 7.29 m | 5/9, 7 | 129.0 CH | 7.28 m | 5/6, 7 | |
| 7 | 127.2 CH | 7.24 m | 5/9, 6/8 | 127.3 CH | 7.24 m | 5/9, 6/8 | |
| 10 | 39.3 CH3 | 2.80 s | 2 | 39.3 CH3 | 2.78 s | 2 | |
| Hivab/Hmpac | 1 | 169.2 qC | – | 2 | 172.3 qC | – | 2 |
| 2 | 78.5 CH | 4.90 d (6.7) | 3, 4, 5 | 78.4 CH | 4.92 d (6.8) | 4, 6 | |
| 3 | 30.7 CH | 2.05 m | 2, 4, 5 | 37.3 CH | 1.81 m | 2, 5, 6 | |
| 4 | 18.2 CH2 | 0.98 d* | 2, 3, 5 | 24.9 CH2 | 1.15 m 1.58 m |
6 | |
| 5 | 18.2 CH2 | 0.98 d* | 2, 3, 4 | 11.7 CH3 | 0.89 t (8.1) | 4 | |
| 6 | – | – | – | 14.6 CH3 | 0.93 d (7.2) | 2, 3 | |
| Pro | 1 | 170.0 qC | – | 2, 3, NH (Ile) | 170.4 qC | – | 2, NH (Ile) |
| 2 | 61.2 CH | 4.62 d (7.8) | 3 | 61.3 CH | 4.62 d (6.5) | 3, 4, 5 | |
| 3 | 31.4 CH2 | 2.65 m 1.96 m |
2, 4, 5 | 31.4 CH2 | 2.64 m 1.97 m |
2, 4, 5 | |
| 4 | 21.8 CH2 | 1.97 m 1.76 m |
2, 3, 5 | 21.8 CH2 | 1.98 m 1.77 m |
2, 3, 5 | |
| 5 | 46.5 CH2 | 3.70 m 3.55 dd (10.4, 10.2) |
2, 3, 4 | 46.5 CH2 | 3.70 m 3.56 dd (12.4, 9.7) |
2, 3, 4 | |
| Ileb/Valc | 1 | 171.4 qC | – | 2, 2 (Gly), NH (Gly) | 171.4 qC | – | 2, 2 (Gly), NH (Gly) |
| 2 | 61.1 CH | 4.20 dd (9.3, 8.5) | 3, 6, NH | 62.4 CH | 4.13 dd (9.8, 8.5) | 3, 4, 5, NH | |
| 3 | 35.2 CH | 2.04 m | 2, 4, 5, 6, NH | 29.2 CH | 2.24 m | 2, 4, 5 | |
| 4 | 25.9 CH2 | 1.58 m 1.24 m |
2, 5, 6 | 19.5 CH3 | 1.01 d (6.9) | 3, 5 | |
| 5 | 10.8 CH3 | 0.87 t (7.5) | 4 | 19.7 CH3 | 1.03 d (7.0) | 3, 4 | |
| 6 | 15.8 CH3 | 1.00 d (6.8) | 2, 3, 4 | – | – | – | |
| NH | – | 7.91 d (8.2) | – | – | 7.88 d (8.3) | – | |
| Gly | 1 | 169.9 qC | – | 2, 3 (Dhoya), NH | 169.9 qC | – | 2, 3 (Dhoya), NH |
| 2 | 41.0 CH2 | 4.62 dd (17.9, 8.7) 3.99 d (17.9) |
– | 41.2 CH2 | 4.62 dd (18.0, 9.4) 4.01 d (18.0) |
– | |
| NH | – | 6.39 d (9.3) | – | – | 6.40 d (9.0) | – | |
Protons showing long-range correlation to indicated carbon
Refers to compound 5
Refers to compound 6
Could not deduce coupling constants due to overlapping peaks
4.7 Pitipeptolide F (6)
Colorless, amorphous solid; [α]20D –101° (c 0.10, MeOH); UV (MeOH) λmax (log ε) 202 (4.34) nm; IR (film) νmax 3404, 2964, 1726, 1646, 1506, 1413, 1370, 1275, 1177, 1094, 1032, 967, 805, 733, 701 cm−1; 1H NMR, 13C NMR and HMBC data, see Table 2; HRESI/APCIMS m/z 816.4518 for [M + Na]+ (calcd for C43H63N5O9Na 816.4518).
4.8 Acid Hydrolysis and Enantioselective Analysis
A sample of compounds 3–6 (0.1 mg each) was hydrolyzed with 6N HCl (0.5 mL) at 110 °C for 24 h. The hydrolysate was concentrated to dryness, resuspended in 100 μL H2O, and then analyzed by chiral HPLC. Amino acid units were analyzed by HPLC/MS chiral analysis [column: Chirobiotic TAG (250 × 4.6 mm), Supelco; solvent: MeOH/10 mM NH4OAc (40:60, pH 5.6); flow rate 0.5 mL/min; detection by ESIMS in positive ion mode (MRM scan)]. The retention times (tR min; MRM ion pair, parent→ product) of the authentic amino acids were as follows: l-Pro (12.3; 116→70), d-Pro (32), l-Val (7.7; 118→72), d-Val (17.0), l-Ile (8.2; 132→86.1), d-Ile (20.5), l-Phe (15.1; 166.2→120.2), d-Phe (22.5), N-Me-l-Phe (21.2; 180→134.2), N-Me-d-Phe (37.0). The hydrolysates showed peaks corresponding to l-amino acids at tR 12.3, 7.7, 8.2, 15.1 and 21.2 min. The MS parameters used were as follows: DP 32 , EP 4 , CE 21.8 , CXP 2.8 , CUR 30, CAD medium, IS 4500, TEM 700, GS1 65, and GS2 65. The absolute configurations of the α-hydroxy acid units were analyzed by chiral HPLC analysis under different conditions; [column: CHIRALPAK MA (+) (50 × 4.6 mm); solvent: CH3CN/2 mM CuSO4 (10:90); flow rate 1 mL/min; detection by UV (254 nm)]. The retention times (tR min) of the authentic standards were as follows: (2S,3S)-Hmpa (30.5), (2R,3R)-Hmpa (17.0), (2S,3R)-Hmpa (24.1), (2R,3S)-Hmpa (14.0), (2S)-Hiva (8.9), (2R)-Hiva (5.0). The retention times of the samples corresponded to (2S,3S)-Hmpa (30.5) in compounds 3, 4, and 6, and (2S)-Hiva (8.9) in 5.
4.9 Cell viability assay
Cells were propagated and maintained in Dulbecco’s modified Eagle medium (Invitrogen, Carlsbad, CA) supplemented with 10% fetal bovine serum (HyClone Laboratories, Logan, UT) at 37 °C humidified air and 5% CO2. Cells were seeded in 96-well plates (MCF7 10,500 cells/well; HT-29 13,000 cells/well). After 24 h, cells were treated with various concentrations of the test compound, or solvent control (1% EtOH). After 48 hours of incubation, cell viability was measured using MTT according to the manufacturer’s instructions (Promega, Madison, WI).
4.10 Disc diffusion assay
An inoculum of Mycobacterium tuberculosis [ATCC # 25177] was grown in a liquid shake culture (Middlebrook 7H9 broth with glycerol and Middlebrook ADC enrichment) for 12 days at 37 °C and adjusted to OD600 ≈ 0.1. This culture was used to inoculate the agar plates. The test compounds were dissolved in EtOH and three different amounts were loaded on sterile 6-mm filter paper discs. The discs were kept at room temperature for 10 minutes to dry, and then loaded on the inoculated agar plates. The plates were incubated in a humidified environment at 37 °C for 12 days, after which the diameter of zone of inhibition (mm) was measured. Streptomycin was used as positive control and the solvent (10 μL) was used as negative control. The experiment was done in duplicate.
Supplementary Material
Acknowledgments
This research was supported by the National Institutes of Health, NIGMS Grant P41GM086210. We thank J. R. Rocca for technical assistance with NMR data acquisition. This is contribution 852 from the Smithsonian Marine Station.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Appendix A. Supporting Information
Supplementary data associated with this article can be found in the online version, at doi:…
References
- Bhatnagar I, Kim SK. Immense essence of excellence: marine microbial bioactive compounds. Mar. Drugs. 2010;8:2673–2701. doi: 10.3390/md8102673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burja AM, Banaigs B, Abou-Mansour E, Burgess JG, Wright PC. Marine cyanobacteria– a prolific source of natural products. Tetrahedron. 2001;57:9347–9377. [Google Scholar]
- Cruz-Rivera E, Paul VJ. Chemical deterrence of a cyanobacterial metabolite against generalized and specialized grazers. J. Chem. Ecol. 2007;33:213–217. doi: 10.1007/s10886-006-9212-y. [DOI] [PubMed] [Google Scholar]
- Gunasekera SP, Owle CS, Montaser R, Luesch H, Paul VJ. Malyngamide 3 and cocosamides A and B from the marine cyanobacterium Lyngbya majuscula from Cocos Lagoon. Guam. J. Nat. Prod. 2011;74:871–876. doi: 10.1021/np1008015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L, Rein K,S. New peptides isolated from Lyngbya species: a review. Mar. Drugs. 2010;8:1817–1837. doi: 10.3390/md8061817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luesch H, Pangilinan R, Yoshida WY, Moore RE, Paul VJ. Pitipeptolides A and B, new cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod. 2001;64:304–307. doi: 10.1021/np000456u. [DOI] [PubMed] [Google Scholar]
- Magarvey NA, Beck ZQ, Golakoti T, Ding Y, Huber U, Hemscheidt TK, Abelson D, Moore RE, Sherman DH. Biosynthetic characterization and chemoenzymatic assembly of the cryptophycins. Potent anticancer agents from cyanobionts. ACS Chem. Biol. 2006;1:766–779. doi: 10.1021/cb6004307. [DOI] [PubMed] [Google Scholar]
- Marahiel MA, Stachelhaus T, Mootz HD. Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem. Rev. 1997;97:2651–2673. doi: 10.1021/cr960029e. [DOI] [PubMed] [Google Scholar]
- Montaser R, Abboud KA, Paul VJ, Luesch H. Pitiprolamide, a proline-rich dolastatin 16 analogue from the marine cyanobacterium Lyngbya majuscula from Guam. J. Nat. Prod. 2011;74:109–11. doi: 10.1021/np1006839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nogle LM, Gerwick WH. Isolation of four new cyclic depsipeptides, antanapeptins A–D, and dolastatin 16 from a Madagascan collection of Lyngbya majuscula. J. Nat. Prod. 2002;65:21–24. doi: 10.1021/np010348n. [DOI] [PubMed] [Google Scholar]
- O’Dor R, Miloslavich P, Yarincik K. Marine biodiversity and biogeography – regional comparisons of global Issues, an introduction. PLoS One. 2010;5:e11871. doi: 10.1371/journal.pone.0011871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peng Y, Pang H, Xu Z, Ye T. The total synthesis of pitipeptolide A. Lett. Org. Chem. 2005;2:703–706. [Google Scholar]
- Schaller A. Action of proteolysis-resistant systemin analogues in wound signaling. Phytochemistry. 1997;47:605–612. doi: 10.1016/s0031-9422(97)00523-2. [DOI] [PubMed] [Google Scholar]
- Tan LT. Bioactive natural products from marine cyanobacteria for drug discovery. Phytochemistry. 2007;68:954–979. doi: 10.1016/j.phytochem.2007.01.012. [DOI] [PubMed] [Google Scholar]
- Thacker RW, Paul VJ. Morphological, chemical and genetic diversity of tropical marine cyanobacteria Lyngbya spp. and Symploca spp. (Oscillatoriales) Appl. Environ. Microbiol. 2004;70:3305–3312. doi: 10.1128/AEM.70.6.3305-3312.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Draanen NA, Arseniyadis S, Crimmins MT, Heathcock CH. Protocols for the preparation of each of the four possible stereoisomeric α-alkyl-β-hydroxy carboxylic acids from a single chiral aldol reagent. J. Org. Chem. 1991;56:2499–2506. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.