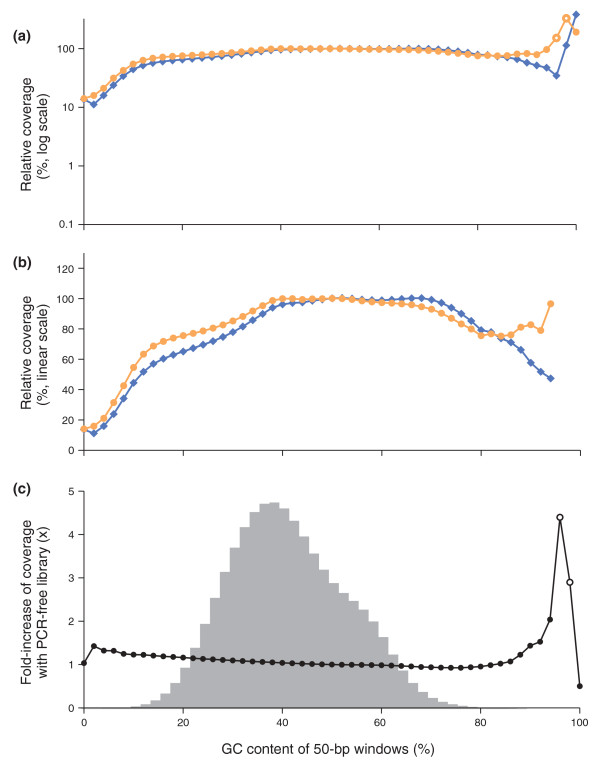
Figure 7.
Sequencing bias with PCR-amplified and PCR-free libraries. (a,b) Shown is the mean normalized coverage of 50-bp windows in the human genome having the GC-content indicated on the x-axis for a PCR-free (orange dots) and a PCR-amplified (blue diamonds) Illumina sequencing library. Both fragment libraries had approximately 180-bp inserts. The PCR amplification was performed with AccuPrime Taq HiFi (long denat., primer extension at 65°C). The coverage was plotted on a log10 (a) and a linear scale (b). The data points at extremely high GC, where the reads from the PCR-free library had a mean base quality of less than Q20 (open symbols), were omitted in the middle panel (b). (c) The ratios of the two curves in (a,b), that is, the fold-increase in mean coverage by sequencing a PCR-free library instead of a PCR-amplified library. The shaded histogram is the %GC distribution of 50-bp windows in the human genome. More than 99.9% of all 50-bp windows in the genome contain 8% to 88% GC and received a less than 1.25-fold increase in coverage. Less than 0.01% of all 50-bp windows contain 90% or more GC. The open circles at 96% and 98% GC denote data for which the mean base quality of the reads from the PCR-free library was below Q20.