Figure 6.
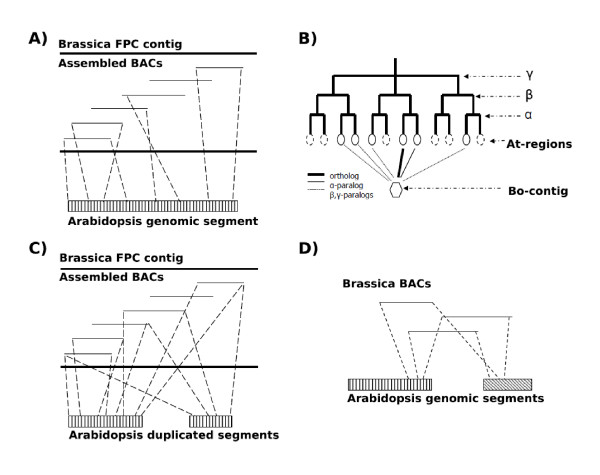
Comparative mapping of Brassica FPC contigs onto the Arabidopsis genome. In subfigures (cartoons, not based on real data) A, C and D, Brassica contigs are displayed with assembled BAC clones (depicted by overlapping lines), and interspecific chromosomal synteny is shown in dashed lines. A). Interspecific chromosomal synteny inference. B). A Brassica contig (shown with a hexagon shape) is expected to be linked to multiple homologous regions in Arabidopsis (shown with circles), at most one ortholog, one α-paralog, two β-paralogs, and eight γ-paralogs. DNA losses may have removed some of them (shown with dashed-lined circles). C). A Brassica contig is linked to Arabidopsis duplicated regions. Unbalanced synteny often permits one to distinguish between orthology and paralogy, or reveals differential gene losses among paralogous regions. D). Inference of synteny discontinuity is shown for a Brassica contig against two Arabidopsis regions, which may indicate a chromosomal breakpoint during the diversification of the two species.