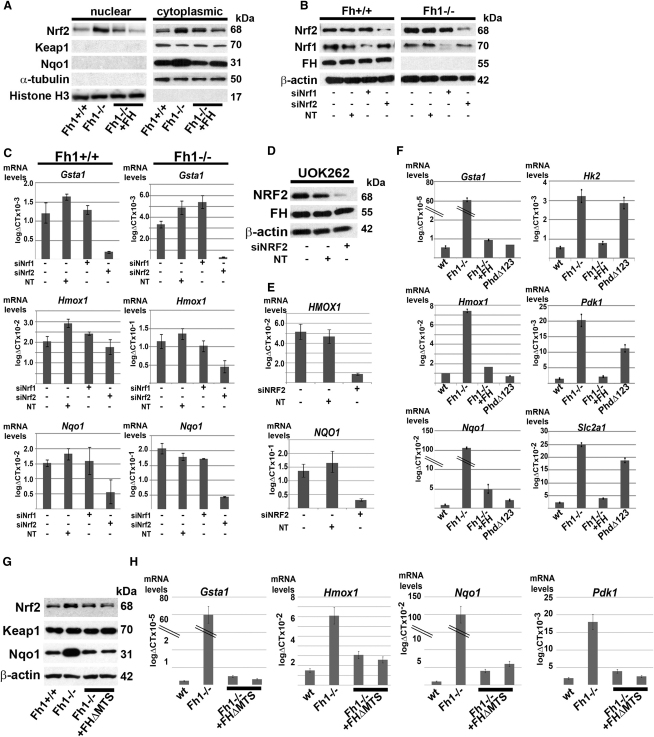
Figure 5.
Upregulation of the Antioxidant Pathway in FH-Deficient Cells Is NRF2-Dependent and HIF/PHD-Independent
(A) Immunoblot of MEF lysates from Fh1+/+, Fh1−/−, and two independent clones of Fh1−/− reconstituted with wild-type FH (Fh1−/−+FH) shows increased levels of Nrf2 in both the nuclear and cytosolic fractions of Fh1−/− cells. Protein levels of Nqo1 are also increased in Fh1−/− MEFS. Protein loading for the nuclear and cytoplasmic fractions is indicated by histone H3 and α-tubulin, respectively.
(B) Immunoblot of Fh1+/+ and Fh1−/− MEFs following siRNA knockdown of Nrf1, Nrf2 and a nontargeting (NT) control. Protein loading is indicated by β-actin.
(C) Q-PCR analysis following siRNA knockdown of Nrf1 or Nrf2 in Fh1+/+ and Fh1−/− MEFs. Gsta1, Hmox1, and Nqo1 are significantly reduced by depletion of Nrf2 (p < 0.02), but not by either Nrf1 knockdown, or the nontargeting control (NT).
(D) Immunoblot of UOK 262 cells for NRF2 and FH following siRNA knockdown. Protein loading is indicated by β-actin.
(E) Q-PCR analysis in UOK 262 cells shows a significant reduction of HMOX1 and NQO1 expression following siRNA knockdown of NRF2 (p < 0.05), but not in cells treated with a non-targeting (NT) control.
(F) Q-PCR analysis of Gsta1, Hmox1, Nqo1, Hk2, Pdk1, and Slc2a1 in Fh1+/+, Fh1−/−, Fh1−/−+FH, and PhdΔ123 MEFs. Fh1−/− MEFs have significantly elevated levels of antioxidant response- and Hif-target genes, whereas PhdΔ123 MEFs upregulate Hif-target genes, but not antioxidant response genes.
(G) Immunoblot of MEF lysate from Fh1+/+, Fh1−/− and two independent clones of Fh1−/− reconstituted with extramitochondrial wild-type FH (Fh1−/−+FHΔMTS) shows increased levels of Nrf2 and Nqo1 in the Fh1−/− cells while Keap1 is equivalent in all the lines. Protein loading is indicated by β-actin.
(H) Q-PCR analysis of Gsta1, Hmox1, and Nqo1 and Pdk1 in Fh1+/+, Fh1−/−, and Fh1−/−+FHΔMTS MEFs. Fh1−/− MEFs have significantly elevated levels of antioxidant response and Hif-target genes, which are ameliorated by extramitochondrial FH expression (O'Flaherty et al., 2010).
All error bars indicate ± 1 SD calculated from three biological replicates, each assayed in duplicate.
See also Figure S1.