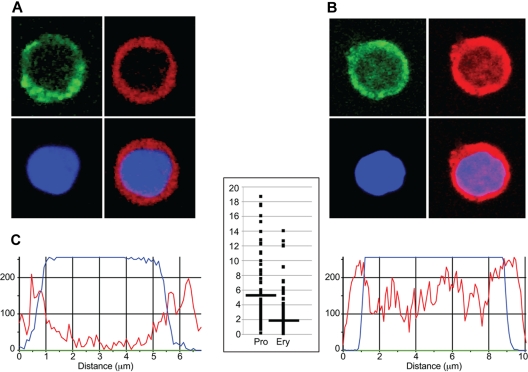
Figure 3.
Immunohistochemical analysis of EKLF in progenitor cell and erythroblast nuclei. Day E13.5 fetal liver cells from HA-tagged EKLF mice were FACS sorted into populations of erythroid progenitor cells (A; CD71+ TER119−) and erythroblasts (B; CD71+ TER119+). The cells were fixed and stained with DAPI (DNA) and an AlexaFluor-594–conjugated anti-HA antibody to identify EKLF in individual cells. (A-B) The top left images represent CD71 staining (green); top right images, EKLF (red); bottom left images, DNA (blue); and bottom right images, the EKLF and DNA images merged. Magenta represents nuclear EKLF. (C) Quantification of EKLF distribution in individual nuclei. The blue tracings represent the relative amount of DAPI signal, and the red tracings represent the relative amount of AlexaFluor-594 signal (EKLF) across an individual progenitor cell (left) or erythroblast (right). The y-axis on the graph in the center represents the ratio of the EKLF-to-DNA signal in the peripheral 20% of the nucleus (10% each side) to the ratio of the EKLF-to-DNA signal in the central 20% of the nucleus for progenitor cells (Pro) and erythroblasts (Ery). A total of 8 peripheral and 8 central data points from 30 nuclei were analyzed for a total of 240 points in each cell type. Images were acquired at room temperature using a Zeiss LSM 510 NLO Meta system mounted on a Zeiss Axiovert 200M microscope with an oil immersion Plan-Apochromat 63×/1.4 DIC objective lens. Excitation wavelengths of 488 nm (3%), 561 nm (6%), and 770 nm (3%) were used to detect anti-CD71 FITC, anti-HA–AlexaFluor-594, and DAPI, respectively. Fluorescent emissions were collected in a BP 500- to 550-nm IR blocked filter, BP 575- to 615-nm IR blocked filter, BP 641- to 705-nm custom filter, and a BP 390- to 465-nm IR blocked filter, respectively. All pinholes were set with a range from 1.11 to 1.33 Airy units, corresponding to an optical slice of 1.0 μm (excluding the DAPI channel where a multiphoton laser was used). All confocal images were of frame size 512 × 512 pixels, scan zoom of 3, and line averaged 8 times. Confocal images were postprocessed using MediaCybernetics' Image-Pro Plus Version 7.0 software package. Each image was processed using a built-in Line Profile tool. A line was drawn through the middle of each cell to determine the density of red (EKLF) and blue (DAPI) signals in the middle 20% of the nucleus and on both edges (10% each side).