Abstract
E. coli T7 bacteriophage gp2 protein is a potent inhibitor of host RNA polymerase (RNAP). Gp2 inhibits formation of open promoter complex by binding to the β′ jaw, an RNAP domain that interacts with downstream promoter DNA. Here, we used an engineered promoter with an optimized sequence to obtain and characterize a specific promoter complex containing RNAP and gp2. In this complex, localized melting of promoter DNA is initiated but does not propagate to include the point of the transcription start. As a result, the complex is transcriptionally inactive. Using a highly sensitive RNAP beacon assay we performed quantitative real-time measurements of specific binding of the RNAP-gp2 complex to promoter DNA and various promoter fragments. In this way, the effect of gp2 on RNAP interaction with promoters was dissected. As expected, gp2 greatly decreased RNAP affinity to downstream promoter duplex. However, gp2 also inhibited RNAP binding to promoter fragments that lacked downstream promoter DNA that interacts with the β′ jaw. The inhibition was caused by gp2-mediated decrease of the RNAP binding affinity to template and non-template strand segments of the transcription bubble downstream of the −10 promoter element. The inhibition of RNAP interactions with single-stranded segments of the transcription bubble by gp2 is a novel effect, which may occur via allosteric mechanism that is set in motion by the gp2 binding to the β′ jaw.
Keywords: RNA polymerase, transcription inhibition, T7 bacteriophage, open promoter complex, protein beacon assay
Introduction
RNA polymerase (RNAP) is amongst the most highly regulated enzymes in the bacterial cell. Bacteriophages have developed ingenious mechanisms to modify host RNAP to make it serve viral needs. Interactions between phage-encoded factors and host RNAPs have long served as a rich source of paradigms for transcriptional regulation (reviewed in1,2). One such example is provided by E. coli bacteriophage T7. The transcription of T7 phage genome proceeds in three stages. Early T7 genes are transcribed from several very strong promoters by E. coli RNAP containing primary σ70 subunit (Eσ70), whereas middle and late T7 genes are transcribed by phage-encoded RNAP3. The product of the middle T7 gene 2, a 7 kDa protein called gp2, binds to the β′ jaw domain of the host RNAP and inhibits it4,5. Only one gp2 molecule participates in tight interaction with an RNAP molecule6 and point mutations in as well as deletions of the β′ jaw abolish this binding5. Gp2 was originally thought to simply switch phage gene transcription from host to viral RNAP. However, its essential biological role is to prevent transcription by host RNAP into those parts of T7 genome that are normally transcribed by viral RNAP. The slow-moving E. coli RNAP slows down the fast-moving T7 RNAP, causing the latter enzyme to pause, which initiates aberrant processing of T7 replicative intermediates, leading to accumulation of improperly packaged, defective phage particles7,8.
The RNAP site of gp2 binding, the β′ jaw, is part of a trough that accepts double-stranded DNA downstream of the transcription start point9. An RNAP mutant lacking the β′ jaw (RNAP-β′Δ jaw) is active but forms unstable transcription initiation complexes9. Binding of gp2 to the β′ jaw prevents formation of detectable promoter complexes by Eσ70 (5,10). Gp2 has no effect on (and does not bind to) preformed open promoter complexes (RPo) or transcription elongation complexes5. Thus, the β′ jaw interactions with downstream DNA and gp2 appear to be mutually exclusive, suggesting that the mechanism of inhibition is based on steric blockade of the β′ jaw interactions with DNA by bound gp2.
It is commonly accepted that RPo formation is initiated when Eσ70 contacts with DNA upstream of the transcription start point are established. Since gp2 binds the β′ jaw, it is expected to inhibit a late stage of RPo formation. Thus, gp2 is predicted to trap an intermediate complex en route to RPo10. However, promoter complexes containing gp2, if they exist, are unstable and standard methods such as footprinting failed to reveal the interaction of the preformed complex of Eσ70 and gp2 (Eσ70-gp2) with promoter10. Here, we used an engineered promoter with optimized sequence to obtain and partially characterize a specific transcriptionally inactive promoter complex containing Eσ70 and gp2. DNase I footprinting and KMnO4 probing of this complex reveal that gp2 prevents establishment of downstream DNA contacts and interferes with DNA melting near the transcription start site but has no effect on RNAP interactions with DNA at and upstream the −10 promoter element. Using a recently developed fluorescence RNAP beacon assay11 we performed quantitative real-time measurements of specific binding of the Eσ70-gp2 complex to promoter DNA and various promoter fragments. In this way, the effect of gp2 on RNAP interaction with promoters was dissected. The data show that gp2 not only greatly decreases RNAP affinity to downstream promoter duplex but also reduces RNAP affinity to non-template strand segment of the transcription bubble located between the −10 element and the transcription start site (called “discriminator” on some promoters). Further, gp2 appears to impede the entry of the template strand into the active center cavity. The inhibition of RNAP-discriminator interactions by gp2 is a novel effect, which may be either direct or indirect, i.e., due to allosteric changes caused by gp2 binding to the β′ jaw. We propose that promoter complexes trapped by gp2 correspond to an intermediate of promoter complex formation. Availability of such complexes, which are very transient in the absence of gp2, opens way for their functional and structural characterization.
Results
Experimental set-up: use of the RNAP beacon assay to monitor the inhibition of E. coli Eσ70 interactions with promoter DNA by T7 gp2
T7 gp2 binding to the jaw domain of the E. coli RNAP β′ subunit decreases Eσ70 affinity to promoter DNA and prevents formation of catalytically competent RPo5,10. Eσ70 binding to promoter DNA and various promoter fragments can be measured using fluorescence protein beacon assay11. The beacon assay relies on the detection of fluorescence signal from RNAP holoenzyme containing the σ70 subunit with fluorescent label site-specifically incorporated in region 2.3, close to the part of σ that recognizes the −10 promoter element. The ground level fluorescence of such RNAP beacon is low because aromatic σ amino acids quench the fluorescence of the label. When RNAP beacon binds promoter DNA or a promoter fragment specific interactions between σ region 2.3 and DNA are established. Since aromatic amino acids that quench the label participate in interactions with promoter DNA, the fluorescent probe becomes “unquenched” leading to increased fluorescence.
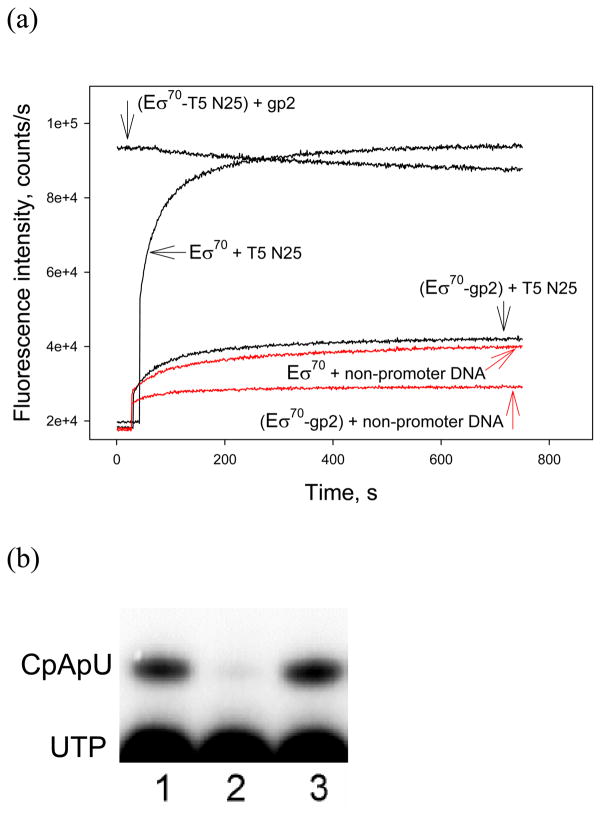
As can be seen from Fig. 2a, the fluorescence signal generated upon the addition of T5 N25 promoter-containing DNA fragment to the RNAP beacon preincubated with excess gp2 is considerably lower than that observed in the absence of gp2. The gp2 effect was insignificant when gp2 was added to preformed complex of RNAP beacon and T5 N25 promoter DNA (Fig. 2a). The results are consistent with standard abortive transcript synthesis assay (Fig. 2b): transcription from T5 N25 was strongly inhibited when gp2 was added to RNAP before the addition of promoter DNA (lane 2), but no effect was observed when gp2 was added to preformed open complexes (lane 3). Thus, the beacon assay allows one to follow the inhibitory action of T7 gp2 on RNAP interactions with promoter.
Figure 2. T7 gp2 inhibits RNAP interaction with the T5 N25 promoter.
(a) Time dependence of the change in fluorescence upon mixing 1 nM [211Cys-TMR] σ70 RNAP holoenzyme with 2 nM −65/+35 T5N25 (black) or a non-promoter DNA fragment (red) in the presence or in the absence of gp2. The upper curve (labeled as “(RNAP+T5N25)+gp2” was obtained in experiment where gp2 was added to the RNAP beacon preincubated with 2 nM T5N25 promoter fragment for 30 min. In other experiments, gp2 was added to preformed RNAP-T5N25 or RNAP-non-promoter DNA complexes.
(b) Inhibition of abortive transcript synthesis from T5N25 promoter fragment by gp2. Reactions contained Eσ70 alone (lane 1) or Eσ70 with gp2 added before (lane 2) or after (lane 3) open complex formation. An autoradiograph of denaturing polyacrylamide gel is presented.
A stable ternary complex containing Eσ70, gp2, and promoter DNA can form when RNAP interactions with promoter are optimized
Gp2 also decreased to the binding of RNAP beacon to control non-promoter DNA fragment (Fig. 1) that contained multiple substitutions in the −35 and −10 regions and lacked transcription activity in vitro. Curiously, the signal observed upon the addition of T5 N25 DNA to Eσ70-gp2 was higher than that observed upon the addition of non-promoter DNA (Fig. 2a). We interpreted this result by hypothesizing that some specific interactions between Eσ70-gp2 complex and promoter are established. However, we could not detect specific binding of T5 N25 DNA to RNAP in the presence of gp2 using KMnO4 probing or DNase I footprinting techniques (data not shown). We hypothesized that this negative result may reflect insufficient stability of ternary (RNAP-gp2-T5 N25) complexes. To improve the putative ternary complex stability we prepared a derivative of −65 to +35 T5 N25 DNA fragment containing consensus −35 element, an optimized UP element12, and the extended −10 element (Fig. 1). We expected that RNAP binding to this promoter, which we call N25cons, would be strengthened due to improved interactions upstream of the −10 promoter element. As expected, the DNA fragment containing N25cons generated significant fluorescence signal when combined with RNAP beacon in the presence of gp2 (Fig. S1a), indicating that specific complex between Eσ70-gp2 and N25cons forms on this promoter. In the absence of gp2, Eσ70 was highly active on N25cons in the abortive transcription initiation assay (Fig. 1b, lane 1), yet, despite the considerable fluorescence signal observed in the beacon assay, transcription from N25cons was highly sensitive to the addition of gp2 prior to promoter complex formation (Fig. S1b, lane 2). Gp2 had no effect on preformed open complexes on N25cons (Fig. S1b, lane 3).
Figure 1. Structures of DNA probes used.
The probe names used in the text are in red. In fork junction probe names, numbers in left and right parenthesis correspond to borders of upper and bottom strands of fork junctions, with respect to the transcription start position located at +1 (underlined), “SP” stands for the SP18 linker that joins non-template strand oligos with template strand oligos in short hairpin fork junctions (shown as a vertical line in corresponding probe structures). Asterisks above the [−50/+14] probe indicate positions where the fragments used in experiment shown in Fig. 8 were truncated.
Eσ70–gp2 formed complexes with the N25cons promoter that were readily detectable and clearly distinct from complexes formed by Eσ70 alone or when gp2 was added after promoter complex formation (Fig. 3). First, DNase I footprinting demonstrated that while Eσ70 strongly protected DNA up to +18 (with weaker protection extending approximately up to position +25), no protection downstream of position +8 was observed in complexes formed by Eσ70–gp2 on either DNA strand (Fig. 3, compare lanes 2 and 3, and 11 and 12). In contrast, the upstream portion of the footprint was identical in all complexes. Second, Exo III treatment of promoter complexes revealed that the downstream boundary of Eσ70 promoter complex, located at position +18 in the absence of gp2, was mostly shifted to around position +3 in the case of Eσ70–gp2 complex (Fig. S2). The upstream boundary, located approximately at position −42, was slightly shifted downstream in the presence of gp2, whether added before or after DNA (Fig. S2, compare lanes 6, 7, and 8). While this observation suggests that gp2 may modify RNAP interactions upstream of the −35 promoter element, this effect is weak, as experiment shown in Fig. 7 (see below) demonstrates. KMnO4 probing revealed that in the absence of gp2, thymine bases at positions from −11 to +3 on the template strand are highly accessible to permanganate, with thymine at position +4 and cytidine at position −14 exhibiting weak sensitivity to the modifying reagent (Fig. 3). In complexes formed by Eσ70–gp2, the reactivity of bases located from the upstream edge of the transcription bubble up to −6 was altered insignificantly, while intensity of KMnO4-sensitive bands further downstream was decreased (Fig. 3, compare lanes 15 and 16). On the non-template strand, strong reactivity at and around the transcription start point was observed in the absence of gp2 (Fig. 3, lane 6). This reactivity was strongly decreased in Eσ70–gp2 complexes (Fig. 3, lane 7). We explain the residual KMnO4 reactivity at and around the transcription start site seen in reactions containing Eσ70–gp2 complexes by the presence of a small fraction of Eσ70–promoter complexes in the reactions. The presence of such complexes is also evident from Exo III footprinting (a minor fraction of exonuclease stops at position of +18, Fig. S2, lane 3) and residual abortive initiation activity (Fig. S1b, lane 5). We therefore suspect that in Eσ70–gp2 complexes per se, the transcription start site is closed explaining why these complexes are transcriptionally inactive. The KMnO4 reactivity decreased insignificantly upon a 2-min challenge of the Eσ70–gp2-N25cons complex with 50 μg/ml of DNA competitor heparin (data not shown).
Figure 3. Eσ70 forms complexes on the N25cons promoter in the presence of gp2.

Reactions containing indicated proteins and N25cons promoter DNA fragment terminally labeled at the template (“t”) or non-template (“nt”) strands were footprinted with DNase I or probed with KMnO4 as described in Materials and Methods. Gp2 was added prior to (lanes 3, 7, 12, 16) or after (lanes 4, 8, 13, 17) open complex formation. Reaction products were resolved by denaturing PAGE and revealed using Phosphorimager.
Figure 7. The effect of T7 gp2 and T4 AsiA on Eσ70 binding to [−59/−12] probe.
2 nM of [−59/−12] probe was combined with RNAP beacon (1 nM) preincubated for 15 min with or without 0.2 μM T7 gp2 or T4 AsiA and increase in fluorescence was monitored.
Overall, the results indicate that when interactions with the upstream promoter part are strong, RNAP forms a specific, stable, yet transcriptionally inactive promoter complex in the presence of gp2. The complex is only partially open and formation of contacts in the downstream portion of the transcription bubble, as well as contacts with double-stranded DNA downstream of the transcription initiation start point appears to be compromised. In the ternary complex, the DNA trajectory downstream from the −10 element must differ considerably from that in RPo. A similar complex may form on the wild-type T5N25 promoter, however it is too transient to be detected by footprinting.
Quantitative dissection of the gp2 inhibitory effect using model promoter fragments
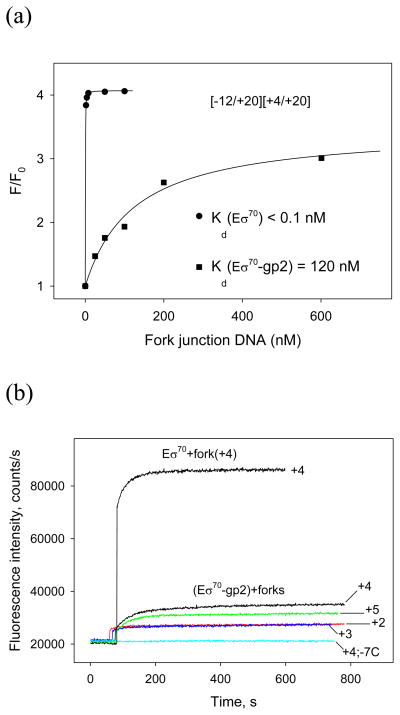
Eσ70 specifically binds upstream and downstream fork junctions--partially double stranded DNA containing short non-template strand overhangs with the −10 promoter element sequence--and this interaction can be monitored by the beacon method11,13,14. By “upstream” and “downstream” we here mean the location of double-stranded segments relative to the non-template strand overhangs. Gp2 inhibits Eσ70 binding to downstream fork junction probes, whereas RNAP binding to upstream fork junction probe [−38/−11][ −38/−12] (shown in Fig. 1) containing a short single-stranded extension corresponding to the −11A base is not affected14. These results indicate that downstream fork junctions contain promoter segments whose recognition by RNAP is targeted by gp2 and, therefore, these probes can be used to study the gp2 inhibition mechanism in more detail. We decided to quantitatively measure Eσ70 binding to various downstream fork junctions and some other probes (shown in Fig. 1) in the presence of gp2. To reach binding saturation, much higher concentrations of downstream fork junction probes were required in the presence of gp2 than in its absence (Fig. 4a, see also14). However, the binding observed in the presence of gp2 was specific, since control probes bearing a C to T substitution at the highly conserved −7 position did not generate any noticeable signal (see, for example, Fig. 4b). The equilibrium dissociation constant (Kd) for Eσ70–gp2 binding to [−12/+20][+4/+20] downstream fork junction probe (with junction point positioned at +4) was calculated by fitting the experimental dependence of the fluorescence signal amplitude on probe concentration to a chemical equilibrium equation (e.g. by titration assay11) (Fig. 4a). The resulting value was 120 nM. Free Eσ70 bound the same probe with a Kd of less than 0.1 nM (Fig. 4a). Thus, gp2 decreases RNAP affinity to the downstream fork junction promoter fragments by more than 1000-fold.
Figure 4. Binding of downstream fork junction probes to the Eσ70-gp2 complex.
(a) Titration of Eσ70 and Eσ70-gp2 with [−12/+20][+4/+20] downstream fork junction. The Kd values were determined by fitting the dependence of relative fluorescence signal amplitude (F/Fo) on probe concentration to a chemical equilibrium equation11. (b) The effect of fork junction point position on Eσ70-gp2 interaction with downstream fork junction probes. Time dependence of the increase in fluorescence upon mixing the (RNAP beacon-gp2) complex with 50 nM of [−12/+20][+2/+20], [−12/+20][+3/+20], [−12/+20][+4/+20], [−12/+20][+5/+20], and [−12/+20; −7C][+4/+20] probes abbreviated as +2, +3, +4, +5, and +4; −7C, correspondingly. The upper curve corresponds to RNAP beacon binding to 50 nM [−12/+20][+4/+20] probe in the absence of gp2.
As shown in Fig. 4b, probes with junction points at +2 and +3 bound the Eσ70-gp2 less efficiently than the [−12/+20][+4/+20] probe. The apparent dependence of downstream fork junctions affinity to Eσ70–gp2 on the junction position is somewhat surprising, since negligible difference in affinity of these probes to Eσ70 was detected14.
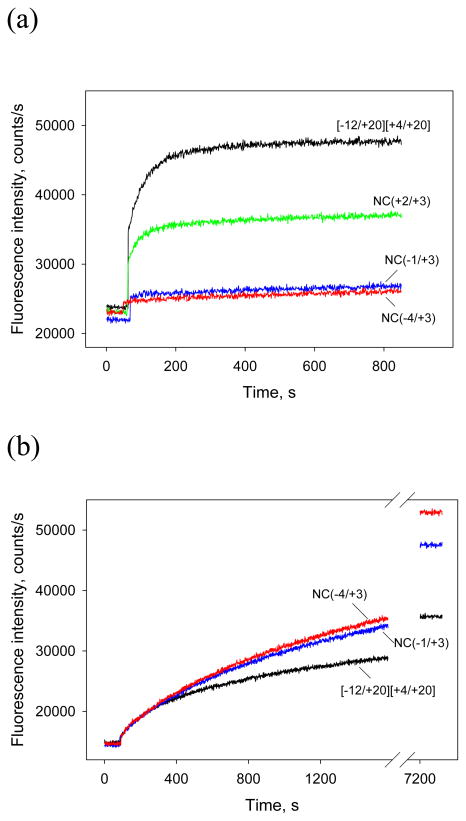
The effect of gp2 on RNAP interaction with a segment of template strand adjacent to the transcription start point was estimated by comparing the binding of [−12/+20][+4/+20] or [−12/+9][+4/+9]SP, a short hairpin downstream fork junction, and their derivatives bearing 2–7 nt-long non-complementary extensions of template strand. Fluorescence signal changes observed upon Eσ70 -gp2 interaction with these probes are shown in Fig. 5a and Fig. S4. The kinetic curves in Fig. 5a indicate that much higher fluorescence levels are observed when Eσ70 -gp2 is mixed with parent [−12/+20][+4/+20] probe than with derivative probes containing template strand overhangs. We determined the Kd value for RNAP-gp2 binding to one such probe, NC(−3/+4). Low signal level made it impossible to determine the Kd value for this probe by titration assay. Therefore, an estimate of RNAP-gp2 binding to NC(−3/+4) (Kd > 0.6 μM) was obtained from a competition binding assay11 by measuring RNAP-gp2 binding to −12/+3 oligo with and without 0.2 μM NC(−3/+4) (Fig. S3). in contrast, the mutant β′Δ jaw RNAP has higher affinity to downstream fork junction probes bearing template strand extensions than to parent probes (Fig. 5b, Table 1). Thus, the inhibitory effect of single-stranded template strand extension on binding of the [−12/+20][+4/+20] derivatives is specifically caused by the gp2 binding to RNAP.
Figure 5. Binding of downstream fork junction probes with template strand overhangs to the Eσ70-gp2 complex and Δβ(1149–1190)Eσ70.
(a) Time dependence of the fluorescence intensity in samples containing 1 nM [211Cys-TMR] σ70 holoenzyme, 200 nM gp2 and 50 nM of indicated downstream fork junction probes.
(b) Time dependence of the fluorescence intensity upon the addition of 3 nM of indicated downstream fork junction probes to 1 nM [211Cys-TMR] σ70 holoenzyme bearing the Δβ′(1149–1190) deletion.
Table 1.
Dissociation constants for the binding of promoter fragments to RNAP beacon, RNAP beacon complex with gp2 and Δβ′(1149–1190) RNAP beacon.
| DNA probe | Kd, RNAP, μM | Kd, (RNAP- gp2), μM | Kd, (RNAP- β′Δjaw), μM |
|---|---|---|---|
| [−12/+20][+4/+20] | <10−4 | 0.12 | 3.4–10−3 |
| NC(−4/+3) | <10−4 | >0.6 | 1.0–10−3 |
| −12/+2 | 0.16 | 4.4 | 0.46 |
| −12/−3 | 1.8 | 17 | |
| −12/−6 | 80 | 87 |
Upon incubation with Eσ70-gp2, [−12/+9][+4/+9]SP-based downstream fork junction with template strand extension generated saturated signal similar to that observed with parental [−12/+9][+4/+9]SP; however, the kinetics of signal increase was significantly slower with the probe bearing the extension (Fig. S4). These data are in marked contrast with the situation observed in the absence of gp2, when an overhang from +3 to −4 actually improves the affinity of short fork junctions to Eσ70 by ~40-fold (Fig. S5). These results indicate that the interactions of template strand overhangs with RNAP are affected by gp2 in the Eσ70-gp2 complexes with downstream fork junctions.
If gp2 inhibited transcription solely by targeting interactions with the double-stranded DNA downstream of the transcription start point, then no effects on the interaction with single-stranded DNA in the transcription bubble should be expected. To determine if gp2 effect of transcription bubble formation is direct or indirect, we studied the effect of gp2 on the interaction of RNAP with oligonucleotides containing sequences matching the −10 promoter element consensus sequence and corresponding to single-stranded non-template extensions of downstream fork junction probes. Eσ70 specifically binds such oligonucleotide probes15 and the interaction can be measured by the RNAP beacon assay11. We found that gp2 inhibits binding of −12/+2 oligo 27-fold (Fig. 6A, Table 1). The effect was specific since i) only very small decrease in RNAP binding to −12/+2 was observed in the presence of R58E gp2 mutant having impaired affinity to RNAP10, and ii) no inhibition of −12/+2 oligo binding was observed upon gp2 addition to RNAP beacon containing RNAP-β′Δjaw14 (data not shown).
Figure 6. Binding of non-template oligos containing the −10 promoter element consensus sequence to free Eσ70 and the Eσ70-gp2 complex.
Titration of [211Cys-TMR] σ70 holoenzyme and the gp2-[211Cys-TMR] σ70 holoenzyme complex with oligonucleotide probes. Solid lines correspond to nonlinear regression fit of the data.
Eσ70 binding to −12/−3 oligo was inhibited by gp2 ~9-fold; the binding of −12/−6 oligo was not affected (Figs. 6b, 6c). The results suggest that gp2 hinders Eσ70 interactions with the −5/+2 segment of the non-template strand. Consistent with this conclusion, gp2 also decreased binding of upstream fork junction probes bearing longer (−11/+2 and −11/−3) non-template strand overhangs, whereas the binding of an upstream fork bearing a short −11/−7 overhang was not affected (Fig. S6).
Comparison of Kd values of Eσ70-gp2 binding to −12/+2 oligo and [−12/+20][+4/+20] fork junction shows that the +3/+20 segment increases affinity 37-fold. In contrast, the +3/+20 segment increased Eσ70 affinity 9000-fold14. Thus, gp2 greatly decreased RNAP affinity to downstream promoter duplex. We also determined Kd values for binding of RNAP beacon containing RNAP-β′Δ jaw to −12/+2 oligo and [−12/+20][+4/+20] probe (Table 1). In agreement with previous results, RNAP-β′Δ jaw binds the downstream fork junction probe much weaker than wild-type RNAP14; however, the mutant RNAP affinity is ~30-fold higher than that of Eσ70-gp2. The data also show that unlike the addition of gp2, deletion of the β′ jaw has only a slight effect on the −12/+2 oligo binding (Table 1).
As noted above, the Exo III footprinting data suggested that gp2 might affect RNAP-promoter interactions at around position −42. Moreover, this effect must be distinct from the “normal” function of gp2 as it is observed whether gp2 is added before of after promoter complex formation (Fig. S2). To determine the significance of this new effect, we measured the consequences of gp2 addition on RNAP binding to [−59/−12] probe. A control, T4 AsiA protein, which is known to inhibit RNAP interaction with the −35 element16, strongly inhibited RNAP binding to this probe (Fig. 7). In contrast, gp2 had very little or no effect on the binding of this probe to Eσ70 (Fig. 7). We conclude that gp2 does not noticeably affect RNAP-promoter interactions upstream of the −10 promoter element.
Effect of gp2 on RNAP interaction with double-stranded promoter fragments truncated downstream of the transcription start point
The DNase I footprinting of the ternary Eσ70-gp2-N25cons complex suggests that gp2 blocks RNAP-promoter contacts downstream from approximately +8 position. A set of double-stranded probes was used to assess the influence of promoter segments downstream of the +6 position on promoter interactions with the RNAP-gp2 complex. The parent probe was a derivative of the [−50/+14] fragment of the T5 N25 promoter containing consensus −35 element and the extended −10 element (Fig. 1). The binding of [−50/+14] and three other derivative probes truncated at +6, +8, and +12 was assessed by measuring RNAP beacon signals upon the addition of i) gp2 to preformed Eσ70 complexes with each probe or ii) DNA probes to the Eσ70-gp2 complex. Upon reaching equilibrium binding, signal amplitudes should be the same independent of the order of DNA and gp2 addition. As can be seen in Fig. 6a, the binding curves were strikingly different when gp2 was added to preformed Eσ70-DNA complexes. The fluorescence intensity decrease observed with the [−50/+14] and [−50/+12] probes was very slow. In contrast, signal amplitudes rapidly reached equilibrium values upon the addition of gp2 to Eσ70 complexed with [−50/+6] or [−50/+8]) probes. Moreover, abortive transcript synthesis from [−50/+6] was inhibited when gp2 was added either prior or after this probe (Fig. 6b, compare lanes 2 and 3). Combined with DNase I footprinting and structural modeling10 data, these results suggest that Eσ70 interactions with duplex DNA downstream of ca. position +9 and with gp2 are mutually exclusive. On the other hand, Eσ70-DNA contacts upstream of ca. position +9 are compatible with gp2 binding, however, the interaction with gp2 leads to formation of an inactive Eσ70-gp2-promoter complex.
Discussion
The T7 gp2 is a powerful inhibitor of promoter complex formation on most promoters. RNAP interactions with gp2 and promoter were thought to be mutually exclusive and promoters that are resistant to gp2, such as the galP1 promoter5, escape inhibition by displacing gp2 from RNAP via optimized interactions of downstream DNA with the enzyme9. The principal result of this work is the demonstration that at conditions when RNAP contacts with promoter elements are strong, a complex between RNAP σ70 holoenzyme, T7 gp2, and promoter can form. This complex is transcriptionally inactive, however, based on DNase I and ExoIII footprints and KMnO4 probing, it resembles “normal” RPo at and upstream of the −10 promoter element. Downstream of the −10 element, the complex containing gp2 differs from RPo: the transcription start point is not melted and contacts with DNA downstream of position +8 are lacking (in RPo, strong downstream protection extends to position +18). Thus, it appears that in the presence of gp2, melting of promoter is initiated normally but does not proceed to the transcription initiation start point. The inhibition of promoter melting near the transcription start site is consistent with a proposal based on a structural model of the gp2-RNAP complex10. While promoter complex we describe here was obtained on a specially designed promoter, we surmise that similar complexes may be formed on other promoters. However, these complexes are too transient to be detected by standard biochemical assays.
It could have been expected that gp2 prevents RPo formation simply by sterically blocking loading of downstream DNA into the enzyme trough formed by theβ′ jaw and several other RNAP domains9,10. However, the fact that gp2 inhibits transcription from open complexes formed on a model template, which lacks DNA downstream from the +6 position (Fig. 6b), suggests that the mechanism of gp2 inhibition is more complex. In fact, the absence of promoter opening at and immediately upstream of transcription start point may be caused by gp2 binding per se, rather than by the absence of downstream DNA loading. Indeed, gp2 inhibits RNAP binding to various promoter fragments lacking distal part of the downstream promoter duplex segment, including oligonucleotides and upstream fork junctions probes which bear no DNA bases downstream from the transcription start site. Since these effects occur in the absence of β′ jaw-DNA contacts, they, therefore, indicate that gp2 inhibits RNAP interaction with downstream part of the transcription bubble between positions −6/+2. Since the site of the binding of gp2, a small protein, is located away from RNAP sites interacting with single-stranded promoter DNA, we believe that this inhibition may be of allosteric nature. For example, the binding of gp2 to the β′ jaw may interfere with movements of the RNAP β′ clamp domain that may be required for establishment of interactions with single-stranded DNA segments17,18. In the RNAP holoenzyme, σ70 subunit region 1.1 is positioned inside the RNAP active-site channel from where it is displaced by the downstream promoter segment upon RPo formation19,20. Recent results suggest that efficient inhibition of Eσ70 by gp2 requires the σ70 region 1.16. Thus, the inhibition of RNAP interactions with single-stranded DNA by gp2 may also involve the σ70 region 1.1. Additional biophysical experiments will be required to test these ideas.
The high affinity for RNAP and the two-pronged mechanism of inhibition of open promoter complex formation pathway, first, by affecting promoter opening, and second, by interfering with the β′ jaw contacts with downstream DNA, make gp2 a very efficient inhibitor. This is a biologically relevant property, since any leaky transcription from early phage promoters, particularly the A3 promoter8, lead to transcriptional interference between the slow-moving host RNAP and fast-moving T7 RNAP, causing abnormal progeny phage genome processing and packaging.
Melting of the transcription start site is an obviously critical step during the formation of the catalytically-competent open promoter complex. Apparently, different factors and/or changes in the RNAP structure can affect promoter melting near the start site, whithout having an effect on the nucleation of promoter melting further upstream. The complex formed in the presence of gp2 appears to be similar to previously described complex formed by RNAP lacking the β lobe21. However, the mutant RNAP complexes establish proper downstream contacts in the presence of transcription substrates and are transcriptionally active. An antibiotic myxopyronin binds to a pocket inside the RNAP β clamp domain and blocks promoter melting propagation towards the active site downstream from the −3 position18,22, again resulting in a complex that may be similar to the gp2-containing complex. Some promoters having GC-rich discriminator sequences are known to form unstable open complexes, a property that is relevant for their specific inhibition during aminoacid starvation2,23,24. It is possible that the stringent control dependent destabilization of complexes formed on these promoters is due to the same mechanism that is used by gp2. Future work will have to establish how such partially opened promoter complexes are related to intermediate complexes on the pathway to RPo formation25–27.
Our work demonstrates that protein beacon assay is a powerful approach to uncover mechanistic details of transcription factor action. The high sensitivity and the ability to probe RNAP interactions with upstream, downstream, and central (corresponding to the transcription bubble) regions of a promoter allows the use of the beacon assay to study even such potent transcription initiation inhibitors as T7 gp2. In addition, because the beacon assay signal relies on unquenching of σ70 region 2.3 aromatic residues with promoter DNA, it allows one to study the local environment of the sigma 2 aromatic cluster in response to different inputs. Other transcription regulators and inhibitors affecting RNAP-promoter interactions, including numerous factors encoded by bacteriophages, can be informatively analyzed by this technique.
Materials and Methods
Proteins
RNA polymerase holoenzyme containing σ70 derivative labeled at position 211 with fluorescent label 5-TMR ([211Cys-TMR] σ70), was prepared as in11. T7 phage gp2 wild type and R58E mutant, E. coli core RNAP containing a deletion (1149–1190) in theβ′ subunit, and T4 phage AsiA protein were prepared as described previously5,10,16.
DNA probes
DNA oligonucleotides and modified oligonucleotides containing the SP18 spacer were synthesized by Integrated DNA Technologies. Fork junction and double stranded DNA probes were prepared as in11,14. A control non-promoter DNA fragment consisted of mutated −65+35 T5 N25 strands in which −10 and −35 promoter element sequences were changed to CGCGGC and CCACTT, respectively.
Fluorometric assays
Fluorescence measurements were performed using a Quanta-Master QM4 spectrofluorometer (PTI) in transcription buffer (40 mM Tris-HCl, pH 8.0, 100 mM NaCl, 5% glycerol, 1 mM DTT and 10 mM MgCl2) containing 0.02% Tween 20 at 25°C. Final assay mixtures (800 μl) contained 1 nM labeled RNAP holoenzyme and DNA probes at various concentrations. The TMR fluorescence intensities were recorded with an excitation wavelength of 550 nm and an emission wavelength of 578 nm. Time-dependent fluorescence changes were monitored after manual-mixing of RNAP beacon (800 μl) and a DNA probe (<20 μl) in a cuvette; the mixing dead-time was 15 s.
To obtain equilibrium dissociation constants (Kd) of the RNAP beacon with oligonucleotides and fork junctions, the experimental dependence of the fluorescent signal amplitude on DNA probe concentration was fitted to a chemical equilibrium equation (i.e. titration assay) as described in11, unless otherwise noted.
The complex of Eσ70 with gp2 (Eσ70-gp2) was prepared by mixing 1 nM Eσ70 beacon with 200 nM gp2 followed by 10 min incubation at room temperature. Increasing gp2 concentration up to 600 nM did not influence binding of DNA probes to Eσ70-gp2 under conditions of the measurements.
In vitro abortive initiation assay
Abortive transcription reactions were performed in a final volume of 10 μl and contained 200 nM of E. coli σ70-holoenzyme (Epicentre), 500 nM of gp2 (where indicated), and 50 nM of various DNA templates prepared as described above in transcription buffer. Reactions were mixed as indicated, incubated for 10 minutes at 37 °C, followed by the addition of CpA RNA dinucleotide primer (200 μM), cold UTP (20 μM) and [α-32P] UTP (3000 Ci/mmol). The reactions were incubated for a further 10 minutes at 37 °C and then terminated by the addition of an equal volume of urea-formamide loading buffer. The reaction products were resolved on a 20% (w/v) polyacrylamide denaturing gel and visualized using a PhosphorImager.
Footprints
For in vitro footprinting experiments, the double-stranded DNA promoter fragments were prepared as in11 using radioactively 5′ end-labeled with [32P]-γATP oligonucleotides corresponding to either non-template or template strand. Next, the labeled promoter fragments were purified on Micro Bio-spin 6 columns (BioRad) and used for the assays.
DNase I footprinting
Promoter complexes were formed in a final volume of 10 μl and contained 200 nM of E. coli σ70-holoenzyme (Epicentre), 2 μM of gp2 (where indicated), and 50 nM of a DNA template in transcription buffer. Reactions were mixed with the order of addition of particular components as indicated, incubated for 10 minutes at 37 °C, followed by the addition of 0.2 units of DNase I (New England Biolabs). The reactions were proceeded for 30 s at 37 °C and were terminated by the addition of 20 μl of stop buffer (10 μg of calf thymus DNA and water) followed by ethanol precipitation. Samples were dissolved in 8 μl of urea-formamide loading buffer and resolved in 7% polyacrylamide denaturing gel.
KMnO4 probing
Reactions were set up as described for DNase I footprinting. Promoter complexes were treated with 1 mM KMnO4 at room temperature for 30 s. Reactions were terminated by the addition of 300 mM β–mercaptoethanol followed by ethanol precipitation and 20-min treatment with 10% piperidine at 95 °C. Reaction products were next treated with chloroform to remove piperidine, ethanol precipitated, dissolved in 8 μl of urea-formamide loading buffer and resolved in 7% polyacrylamide denaturing gel.
Exo III footrpinting
Reactions were set up as described above. Promoter complexes were treated with 0.5 units of Exo III (New England Biolabs) for 2 minutes at 37 °C. Reactions were terminated by the addition of 20 μl of stop buffer (10 μg of calf thymus DNA and water), and precipitated with ethanol. Samples were dissolved in 8 μl of urea-formamide loading buffer and resolved in 7% polyacrylamide denaturing gel.
Supplementary Material
Figure 8. The effect of gp2 on Eσ70 interaction with double-stranded promoter fragments truncated at various positions downstream of the transcription start point.
(a) Time dependence of fluorescence intensity in samples containing 1 nM [211Cys-TMR] σ70 holoenzyme, 200 nM gp2 and 2 nM of either of [−50/+14], [−50/+12], [−50/+8], or [−50/+6] fragments. Upper curves correspond to experiments in which gp2 was added to the RNAP beacon preincubated with a DNA probe for 30 min; bottom curves show fluorescence intensity changes upon addition of the DNA probes to the RNAP beacon preincubated with gp2. The addition of gp2 or DNA probe is indicated by vertical arrows. (b) Inhibition of abortive transcript synthesis from the [−50/+6] probe by gp2. Reactions contained Eσ70 alone (lane 1) or RNAP with gp2 added before (lane 2) or after (lane 3) open complex formation.
Acknowledgments
this work was supported by NIH grants GM59295 and GM64530, and the Russian Academy of Sciences Presidium Molecular and Cellular Biology Program grant to KS, SW is the recipient of a BBSRC David Phillips Fellowship (BB/E023703) and CS of a BBSRC DTG studentship.
Abbreviations used
- RNAP
RNA polymerase
- Eσ70
RNAP holoenzyme containing primary σ70 subunit
- RPo
open promoter complex
- Kd
equilibrium dissociation constant
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Nechaev S, Severinov K. Bacteriophage-induced modifications of host RNA polymerase. Annu Rev Microbiol. 2003;57:301–322. doi: 10.1146/annurev.micro.57.030502.090942. [DOI] [PubMed] [Google Scholar]
- 2.Haugen SP, Ross W, Gourse RL. Advances in bacterial promoter recognition and its control by factors that do not bind DNA. Nat Rev Microbiol. 2008;6:507–519. doi: 10.1038/nrmicro1912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Molineux IJ. The T7 group. In: Calendar RL, editor. The Bacteriophages. 2. Oxford University Press; New York: 2005. pp. 277–301. [Google Scholar]
- 4.Hessellbach B, Nakada D. Inactive complex formation between E. coli RNA polymerase and inhibitor protein purified from T7 phage infected cells. Nature. 1975;258:354–357. doi: 10.1038/258354a0. [DOI] [PubMed] [Google Scholar]
- 5.Nechaev S, Severinov K. Inhibition of Escherichia coli RNA polymerase by bacteriophage T7 gene 2 protein. J Mol Biol. 1999;289:815–826. doi: 10.1006/jmbi.1999.2782. [DOI] [PubMed] [Google Scholar]
- 6.Sheppard C, Cámara B, Shadrin A, Akulenko N, Liu M, Baldwin G, et al. Inhibition of Escherichia coli RNAp by T7 Gp2 protein: role of negatively charged strip of amino acid residues in Gp2. J Mol Biol. 2011;407:623–632. doi: 10.1016/j.jmb.2011.02.013. [DOI] [PubMed] [Google Scholar]
- 7.Qimron U, Kulczyk A, Hamdan S, Tabor S, Richardson C. Inadequate inhibition of host RNApolymerase restricts T7 bacteriophage growth on hosts overexpressing udk. Mol Microbiol. 2007;67:448–457. doi: 10.1111/j.1365-2958.2007.06058.x. [DOI] [PubMed] [Google Scholar]
- 8.Savalia D, Robins W, Nechaev S, Molineux I, Severinov K. The role of the T7 Gp2 inhibitor of host RNA polymerase in phage development. J Mol Biol. 2010;402:118–126. doi: 10.1016/j.jmb.2010.07.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ederth J, Artsimovitch I, Isaksson LA, Landick R. The downstream DNA jaw of bacterial RNA polymerase facilitates both transcriptional initiation and pausing. J Biol Chem. 2002;277:37456–37463. doi: 10.1074/jbc.M207038200. [DOI] [PubMed] [Google Scholar]
- 10.Camara B, Liu M, Reynolds J, Shadrin A, Liu B, Kwok K, et al. T7 phage protein Gp2 inhibits the Escherichia coli RNA polymerase by antagonizing stable DNA strand separation near the transcription start site. Proc Natl Acad Sci USA. 2010;107:2247–2252. doi: 10.1073/pnas.0907908107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mekler V, Pavlova O, Severinov K. The interaction of Escherichia coli RNA polymerase σ70 subunit with promoter elements in the context of free σ70, RNA polymerase holoenzyme, and the β′–σ70 complex. J Biol Chem. 2011;286:270–279. doi: 10.1074/jbc.M110.174102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Estrem ST, Gaal T, Ross W, Gourse RL. Identification of an UP element consensus sequence for bacterial promoters. Proc Natl Acad Sci USA. 1998;95:9761–9766. doi: 10.1073/pnas.95.17.9761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Guo J, Gralla JD. Promoter opening via a DNA fork junction binding activity. Proc Natl Acad Sci USA. 1998;95:11655–11660. doi: 10.1073/pnas.95.20.11655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mekler V, Minakhin L, Severinov K. A critical role of downstream RNA polymerase-promoter interactions in the formation of initiation complex. J Biol Chem. 2011;286:22600–22608. doi: 10.1074/jbc.M111.247080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Marr MT, Roberts JW. Promoter recognition as measured by binding of polymerase to nontemplate strand oligonucleotide. Science. 1997;276:1258–1260. doi: 10.1126/science.276.5316.1258. [DOI] [PubMed] [Google Scholar]
- 16.Severinova E, Severinov K, Fenyö D, Marr M, Brody EN, Roberts JW, Chait BT, Darst SA. Domain organization of the Escherichia coli RNA polymerase sigma 70 subunit. J Mol Biol. 1996;263:637–647. doi: 10.1006/jmbi.1996.0604. [DOI] [PubMed] [Google Scholar]
- 17.Darst SA, Opalka N, Chacon P, Polyakov A, Richter C, Zhang G, Wriggers W. Conformational flexibility of bacterial RNA polymerase. Proc Natl Acad Sci USA. 2002;99:4296–4301. doi: 10.1073/pnas.052054099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mukhopadhyay J, Das K, Ismail S, Koppstein D, Jang M, Hudson B, et al. The RNA polymerase “switch region” is a target for inhibitors. Cell. 2008;135:295–307. doi: 10.1016/j.cell.2008.09.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mekler V, Kortkhonjia E, Mukhopadhyay J, Knight J, Revyakin A, Kapanidis AN, et al. Structural organization of bacterial RNA polymerase holoenzyme and the RNA polymerase promoter open complex. Cell. 2002;108:599–614. doi: 10.1016/s0092-8674(02)00667-0. [DOI] [PubMed] [Google Scholar]
- 20.Murakami KS, Masuda S, Darst SA. Structural basis of transcription initiation: an RNA polymerase holoenzyme initiation: RNA polymerase holoenzyme at 4 Å resolution. Science. 2002;296:1280–1284. doi: 10.1126/science.1069594. [DOI] [PubMed] [Google Scholar]
- 21.Severinov K, Darst SA. A mutant RNA polymerase that forms unusual open promoter complexes. Proc Natl Acad Sci USA. 1997;94:13481–13486. doi: 10.1073/pnas.94.25.13481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Belogurov GA, Vassylyeva MN, Sevostyanova A, Appleman JR, Xiang AX, Lira R, et al. Transcription inactivation through local refolding of the RNA polymerase structure. Nature. 2009;457:332–335. doi: 10.1038/nature07510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Paul BJ, Ross W, Gaal T, Gourse RL. rRNA transcription in Escherichia coli. Annu Rev Genet. 2004;38:749–770. doi: 10.1146/annurev.genet.38.072902.091347. [DOI] [PubMed] [Google Scholar]
- 24.Rutherford ST, Villers CL, Lee JH, Ross W, Gourse RL. Allosteric control of Escherichia coli rRNA promoter complexes by DksA. Genes Dev. 2009;23:236–248. doi: 10.1101/gad.1745409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Saecker RM, Tsodikov OV, McQuade KL, Schlax PE, Jr, Capp MW, Record MT., Jr Kinetic studies and structural models of the association of E. coli σ70 RNA polymerase with the lambdaP(R) promoter: large scale conformational changes in forming the kinetically significant intermediates. J Mol Biol. 2002;319:649–671. doi: 10.1016/S0022-2836(02)00293-0. [DOI] [PubMed] [Google Scholar]
- 26.Sclavi B, Zaychikov E, Rogozina A, Walther F, Buckle M, Heumann H. Real-time characterization of intermediates in the pathway to open complex formation by Escherichia coli RNA polymerase at the T7A1 promoter. Proc Natl Acad Sci USA. 2005;102:4706–4711. doi: 10.1073/pnas.0408218102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Saecker RM, Record MT, Jr, Dehaseth PL. Mechanism of bacterial transcription initiation: RNA polymerase - promoter binding, isomerization to initiation-competent open complexes, and initiation of RNA synthesis. J Mol Biol. 2011 doi: 10.1016/j.jmb.2011.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.