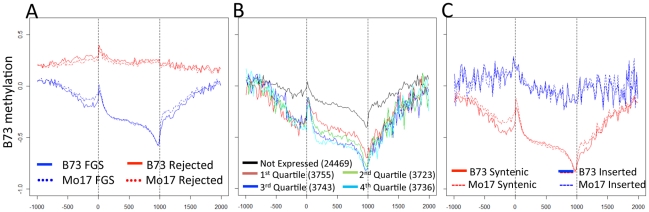
Figure 2. Gene body methylation and expression levels in maize genes.
(A) The relative methylation levels (log2(IP/input)) were assessed for probes within 1000 bp of the transcription start and termination site. The relative position for probes within the gene was normalized to a scale of 1000. The genes in the filtered gene set (FGS) exhibit a lower methylation and a more dynamic pattern across the length of the gene than the rejected genes. The vertical dashed lines indicated the beginning and end of transcription for each gene. (B) The relative expression level for all FGS genes was assessed using published RNAseq data from B73 leaf tissue [56] and genes were assigned as not expressed or quartile 1–4 based on their expression level. In general, the genes show similar patterns of methylation but the higher expressed genes exhibit lower levels of methylation within and around the gene. (C) Each of the FGS genes was also classified according to whether it was located in a syntenic position relative to sorghum and/or rice or in a non-syntenic position. The syntenic genes exhibit much lower levels of methylation than the non-syntenic genes. This difference between syntenic and non-syntenic genes can also be seen in the regions immediately surrounding the gene.