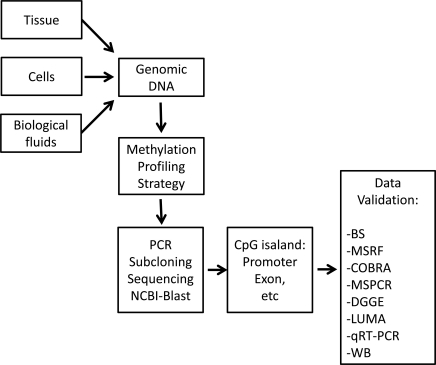
Fig. (3).
Step-by-step approach from the discovery to validation of the CpG methylation. Genomic DNA could be extracted from different types of biological specimens. Based on the investigators need methylation study could begin using a variety of methylation profiling tools and could be followed by PCR and subcloning into a vector, sequencing, aligning into NCBI-Blast data base. After proper identification of the gene or sequence of interest or candidate CpG islands, and designing specific primers, final data validation assays is necessary to confirm the relationship between methylation level and gene or protein expression. This will be performed by different methods listed in the flow chart. BS, Bisulfite sequencing; MSRF, methylation-sensitive restriction fingerprinting; COBRA, Combined bisulfite restriction analysis; MSPCR, methylation-sensitive PCR; DGGE, denaturing gel electrophoresis; LUMA, luminometric methylation assay; qRT-PCR, quantitative real-time PCR; WB, western blotting.