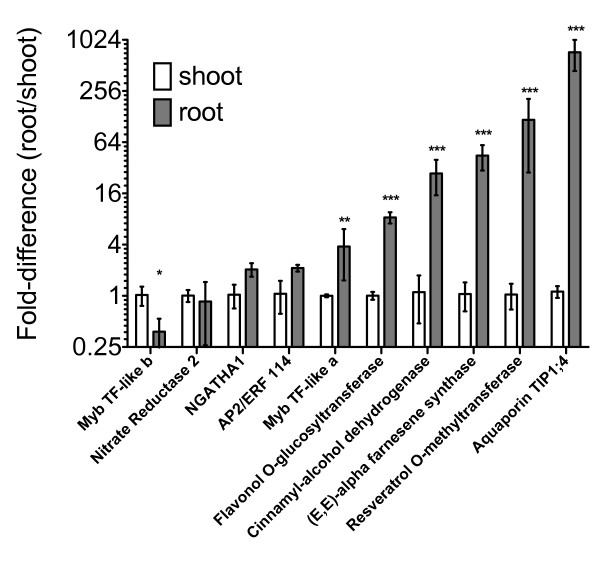
Figure 7.
Expression of candidate root-specific genes in roots and shoots of Cabernet Sauvignon. qRT-PCR analysis of ten selected transcripts in shoot (white bars) and root (gray bars) tissues. Transcript abundances derived from three biological replicates were normalized to an actin reference gene and fold differences were standardized to shoot expression values. Error bars represent standard error. Two-way ANOVA (gene, tissue) was performed followed by post-test Bonferroni-corrected t-statistics. Significant differences in gene expression (root compared with shoot) are indicated by asterisks. * denotes p < 0.05; ** denotes p < 0.01; *** denotes p < 0.001. Fold-differences are drawn on log scale. The tested genes are listed below in the order that they appear on the graph from left to right, with the number of root ESTs compared with non-root ESTs in parentheses. Myb family transcription factor-like b, (7 compared with 1); Nitrate reductase 2, (9 compared with 3); NGATHA1 transcription factor, (5 compared with 0); (AP2/ERF transcription factor, 6 compared with 0); Myb family transcription factor-like a, (5 compared with 0); Flavonol 3-O-glucosyltransferase, (10 compared with 1); Cinnamaldehyde dehydrogenase, (9 compared with 1); (E, E)-alpha-Farnesene synthase, (23 compared with 0); Resveratrol O-methyltransferase, (30 compared with 0); Aquaporin TIP1;4, (57 compared with 2).