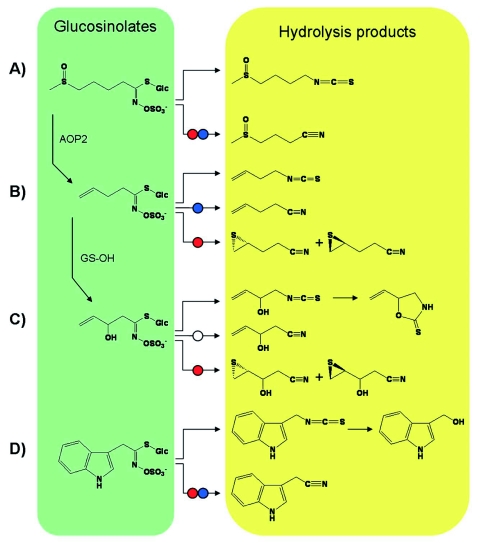
Figure 2.
Chemical diversity of glucosinolate breakdown products formed upon tissue damage.
Structures of intact glucosinolates (green box) and their potential breakdown products (yellow box) in Arabidopsis are depicted for A) 4-methylsulfinylbutylglucosinolate, B) 3-butenylglucosinolate, C) 2-hydroxy-3-butenylglucosinolate, and D) indol-3-ylmethylglucosinolate. Vertical arrows and enzyme names indicate the biosynthetic link between the glucosinolates in A)-C). Arabidopsis AOP2 corresponds to GSL-ALK in Brassica species. Hydrolysis in the absence of specifier proteins (horizontal arrow, no circle) results in isothiocyanate (R-N=C=S) formation. Non-enzymatic cyclization of the isothiocyanate derived from 2-hydroxy-3-butenylglucosinolate yields goitrin (5-ethenyl-1,3-oxazolidine-2-thione, C). Indol-3-ylmethylisothiocyanate is known to further react to indole-3-carbinol, D). Red circles indicate activity of epithiospecifier protein (ESP) on alkenylglucosinolates (B, D) and on glucosinolates with other aliphatic or indolic side chains (A, D). Blue circles indicate activity of nitrile-specifier proteins (NSPs) (A, B, D). The formation of the simple nitrile from 2-hydroxy-3-butenylglucosinolate by NSPs (white circle) has not yet been demonstrated experimentally.