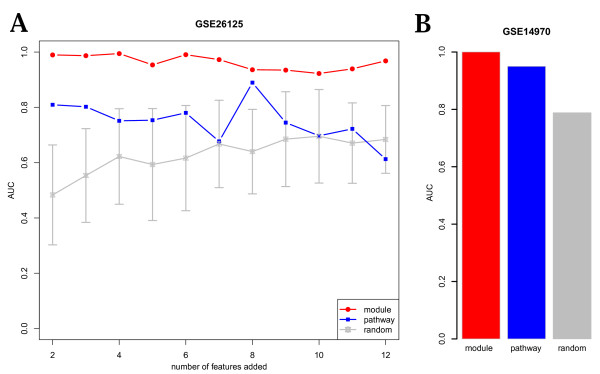
Figure 4.
Within- and cross-validation performance comparison of modules and pathways. (A) Within-dataset classification evaluation. X-axis corresponds to the number of features added to the logistic classifier. Assuming that there are K features (modules/pathways/random gene sets), then for each k ≤K, select the first k th modules to train the classifier. The final classification performance was reported as the AUC on the testing set using the classifier optimized from the validation set in a five-fold cross-validation. Modules were ordered in decreasing significance of MI, pathways were ordered in decreasing significance of enrichment, and random gene sets were in the order as their compared modules. (B) In cross-dataset classification evaluation, all 12 features were trained on GSE26125 and validated on 12 disease samples from GSE14970 combined with 5 controls from GSE26125.