Abstract
Objective
To identify the genetic etiology in a family with autosomal dominant progressive sensorineural hearing loss.
Design
Prospective molecular genetic research study.
Setting
Academic genetic research laboratory.
Participants
Seventeen members of a family with dominant progressive nonsyndromic sensorineural hearing loss: 9 affected, 6 unaffected, and 2 spouses.
Interventions
Clinical data from questionnaires, interviews, serial audiograms, and medical records; genetic data from genome-wide linkage analysis and candidate gene mutation analysis.
Main Outcome Measures
Symptoms, age at onset, serial audiometric data, and the presence or absence of a deafness-associated mutation.
Results
Affected individuals in this family presented with autosomal dominant nonsyndromic high-frequency progressive sensorineural hearing loss, with age at onset ranging from 1 to 21 years. Genome-wide linkage analysis of single-nucleotide polymorphisms yielded evidence of linkage to an 18.9-Mb region on chromosome 1p34–p36, with a multipoint logarithm of odds score of 3.6. This interval contains a known deafness gene, KCNQ4, which underlies DNFA2 deafness. Sequencing of the 14 coding exons and intron-exon junctions of KCNQ4 revealed a novel heterozygous missense mutation, c.859G>C, p.Gly287Arg. The mutation disrupts the highly conserved GYG motif (glycine-tyrosine-glycine) of the phosphate- binding loop, hypothesized to be critical in maintaining pore structure and function. All 274 controls were negative for the mutation.
Conclusions
Autosomal dominant high-frequency hearing loss is genetically heterogeneous, and linkage analysis is an efficient means of identifying the etiology in larger families. Deafness in this family is caused by a novel mutation in KCNQ4.
In recent years, more than 50 genes responsible for hearing impairment have been identified.1 Whereas GJB2 mutations are the most common cause of recessive nonsyndromic deafness in the US population, dominant nonsyndromic deafness may result from mutations in a variety of genes, none of which predominate in the population. Although various clinical algorithms have been proposed, developing an efficient and cost-effective means to diagnose the genetic etiology of dominant deafness remains a challenge.
The DFNA2 (deafness, nonsyndromic, autosomal dominant 2) locus was originally mapped to chromosome 1p in 2 families with progressive high-frequency sensorineural hearing loss (SNHL).2 Ultimately, a potassium channel gene, KCNQ4 (OMIM NM_0047000), was identified as the responsible gene.3,4 We used genetic linkage analysis and candidate gene screening to identify the genetic cause of deafness in a family with dominant progressive high-frequency SNHL.
METHODS
This research was approved by the University of Michigan and the Baylor College of Medicine and Affiliated Hospitals institutional review boards, and informed consent was obtained from all the participants. A family (family 63) of European descent segregating high-frequency SNHL was ascertained through contact with the principal investigator (M.M.L.) (Figure 1). Participants completed questionnaires, underwent audiologic testing, and provided saliva or peripheral venous blood samples. Historical clinical and audiometric data were obtained as available. DNA was isolated from blood using the Gentra Puregene Cell Kit (Qiagen, Valencia, California) and from saliva using the Oragene DNA Kit (DNA Genoteck, Kanata, Ontario, Canada) according to the manufacturers’ instructions.
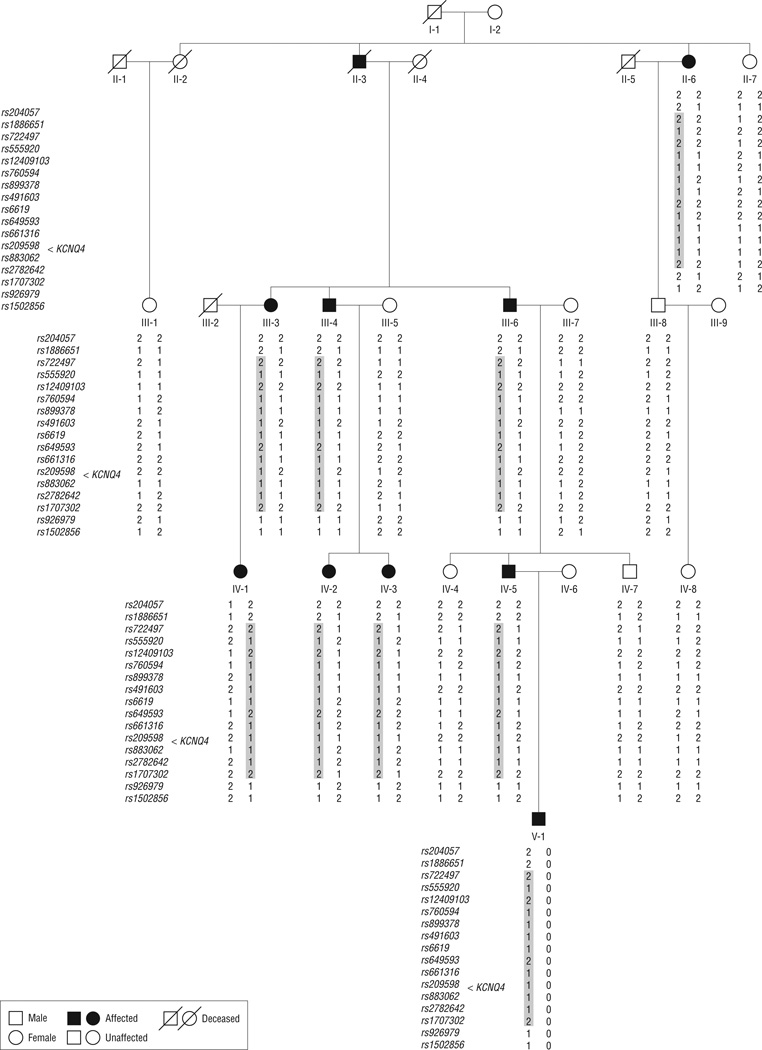
Figure 1.
Pedigree of family 63 segregating progressive high-frequency sensorineural hearing loss. Single-nucleotide polymorphisms are listed to the left of the individual’s genotype. The haplotype segregating with deafness is highlighted with a gray box. The interval is defined by recombination events at rs1886651 (centromeric) and rs926979 (telomeric). KCNQ4 maps between rs209598 and rs883062.
GENOTYPING AND LINKAGE ANALYSIS
A whole-genome scan was performed at the Center for Inherited Disease Research using the Illumina HumanLinkage-12 panel (Illumina Inc, San Diego, California), which contains 6090 single-nucleotide polymorphism (SNP) marker loci, for all 16 pedigree members from whom a DNA sample was available. Genotype data were screened for mendelian incompatibilities using PedCheck,5 and MERLIN (Multipoint Engine for Rapid Likelihood Inference)6 was used to assess the data for the occurrence of double recombination events over short genetic distances, which are most likely due to genotyping error. MLINK of the FASTLINK software package7 was used to perform 2-point linkage analysis, and Allegro1.2c8 was used for multipoint linkage analysis. An autosomal dominant mode of inheritance with complete penetrance and a disease allele frequency of 0.001 were used in the analysis. Marker allele frequencies were based on HapMap9 for the CEU (Centre d’Etude de Polmorphisme Humain) population. (CEU is a control population consisting of 180 Utah residents with Northern and Western European ancestry.) For the multipoint analysis, genetic map distances were based on the Build 36 version of the Rutgers combined linkage-physical map of the human genome.10 For SNP loci that were not included on the Rutgers map, the sequence-based physical map (Build 36) was used as a reference to determine the physical positions of SNP marker loci, and then interpolation was used to determine the genetic map distance. Haplotype reconstruction was performed using SimWalk2.11,12
SEQUENCING OF CANDIDATE GENES
Candidate genes mapping to the linkage region were screened by sequencing of all exons and intron-exon boundaries. Primer sets for polymerase chain reaction were designed using Primer3 Software13 (Table 1). Polymerase chain reaction was performed using the FailSafe PCR Kit according to the manufacturer’s instructions (Epicentre Biotechnologies, Madison, Wisconsin) with a 50-µL reaction volume, 200-ng template DNA, and 0.2-µM primer, with 32 cycles at 58°C annealing temperature and 1 minute 30 seconds of extension time. The polymerase chain reaction product was purified using Qiaquick columns (Qiagen) and was sequenced by the University of Michigan DNA Sequencing Core.
Table 1.
Polymerase Chain Reaction Primers Used for Amplification of KCNQ4 Exons
| Primer Name |
5′-3′ Sequence | Type | Exon Targeted |
|---|---|---|---|
| KCNQ4x1f | GGAAAGAGCAGCCCCAGT | Forward | 1 |
| KCNQ4x1r | GATCAGAAACCGGGCTTAGG | Reverse | 1 |
| KCNQ4x2f | CCAGGGAATTCCAATTCTGA | Forward | 2 |
| KCNQ4x2r | TTGCATAGAGGCAGGAGGTC | Reverse | 2 |
| KCNQ4x3f | CCCTAACCCAGGGACTCTTG | Forward | 3 |
| KCNQ4x3r | GATTGGGAGTAAAGGCGTCA | Reverse | 3 |
| KCNQ4x4f | GAGACCCGCACAGATTCAG | Forward | 4 |
| KCNQ4x4r | AGACAGCCCCTCTGACCTC | Reverse | 4 |
| KCNQ4x5f | CCTTTATCCCTTTCCCGTGT | Forward | 5 |
| KCNQ4x5r | TGAGTCAGGAGTCACGATGG | Reverse | 5 |
| KCNQ4x6f | CCTGTACCCCCAACAAGC | Forward | 6 |
| KCNQ4x6r | CAAATGTGTGACAGGGGTGA | Reverse | 6 |
| KCNQ4x7f | GACACCCTTGCAGCCTCTTA | Forward | 7 |
| KCNQ4x7r | AGCACACAGGGTTGACACAC | Reverse | 7 |
| KCNQ4x8f | CCAAGGACTGGGTTCTTTCA | Forward | 8 |
| KCNQ4x8r | TCTCAGCCACTTGTCCTCAA | Reverse | 8 |
| KCNQ4x9f | TCCACCCTGTCCTATTCTGG | Forward | 9 |
| KCNQ4x9r | GGCAGGTCTGAGAGAGGATG | Reverse | 9 |
| KCNQ4x10f | AGTCAGCTTTGTCCCCACTC | Forward | 10 |
| KCNQ4x10r | AGACGCGAGCCAGATCTTT | Reverse | 10 |
| KCNQ4x11f | CTGGTGGTTTGGCATACAAG | Forward | 11 |
| KCNQ4x11r | GGCTGGTCTCAAACTCCTGA | Reverse | 11 |
| KCNQ4x12f | TGGACAAGGGGATAGCAAAG | Forward | 12 |
| KCNQ4x12r | GGCCTCAGACTTCATTCAGG | Reverse | 12 |
| KCNQ4x13f | TTCTCCTTCATCAGGCAACC | Forward | 13 |
| KCNQ4x13r | GCATCCTCCCCATGTCAC | Reverse | 13 |
| KCNQ4x14f | CTAGCCAAGCTCCACCTTTC | Forward | 14 |
| KCNQ4x14r | CATAGGGCCTTGATGGAGTG | Reverse | 14 |
ANALYSIS OF VARIANTS
Sequence data were compared with the reference sequence (NM_004700) for KCNQ4, and variations were recorded. Novel variants were assayed in a panel of 275 controls (550 control chromosomes), of which 185 were from the Caucasian-American Panel and 90 were from the Polymorphism Discovery Resource (Coriell Cell Repositories, Camden, New Jersey), containing 24 European Americans, 24 African Americans, 24 Asian Americans, 12 Mexican Americans, and 6 Native Americans, according to the same protocols listed previously herein.
To assess the affect of amino acid substitutions, wild-type amino acid sequence was compared with mutant sequence using the SIFT (Sorting Intolerant from Tolerant) program.14 A SIFT score of 0.05 or lower is predicted to be deleterious, and a score higher than 0.05 predicts a tolerated substitution.
RESULTS
CLINICAL DATA
Of 17 participants, 9 were affected with hearing loss, 6 were unaffected, and 2 were spouses. Age at onset ranged from 1 to 21 years. Affected family members had sloping SNHL, with low-frequency thresholds less affected than high-frequency thresholds. In general, hearing was symmetrical. Hearing loss was noted to typically progress with age, with mild to moderate hearing loss identified early in life progressing to severe-profound loss by the seventh decade of life (Table 2). Several family members successfully used hearing aids; none were cochlear implant recipients. The phenotype in the family members affected with hearing loss was consistent with DFNA2 deafness.15
Table 2.
Audiologic Findings for Affected Members of Family 63
| Individual | Age When Tested, y |
Audiometric Pattern of Hearing Lossa |
|---|---|---|
| II-6 | 89 | Severe to profound SNHL |
| III-3 | 39 | Moderate to profound SNHL |
| III-4 | 36 | Mild sloping to moderately severe SNHL |
| III-4 | 44 | Moderate sloping to severe SNHL |
| III-4 | 51 | Moderate sloping to severe-profound SNHL |
| III-6 | 49 | Moderate to 4000-Hz sloping to profound SNHL |
| IV-1 | 7 | Mild sloping to moderately severe SNHL |
| IV-2 | 1.5 | Borderline normal sloping to mild SNHL in the better-hearing ear |
| IV-2 | 4 | Mild SNHL |
| IV-2 | 15 | Mild sloping to 4000-Hz sloping to moderately severe SNHL |
| IV-3 | 4 | Borderline normal sloping to mild SNHL |
| IV-3 | 5 | Mild to moderate SNHL |
| IV-3 | 10 | Mild to moderate SNHL |
| IV-3 | 18 | Mild to moderately severe SNHL |
| IV-5 | 7 | Mild sloping to moderate SNHL |
| IV-5 | 12 | Mild sloping to moderate SNHL |
| IV-5 | 24 | Mild sloping to moderately severe SNHL |
| V-1 | 3 | Moderate SNHL |
Abbreviation: SNHL, sensorineural hearing loss.
Hearing loss is bilateral unless otherwise specified.
GENETIC ANALYSIS
Results of the genome-wide SNP genotyping analysis showed significant evidence of linkage to chromosome 1p34-p36. Themaximum2-point logarithm of odds (LOD) score of 1.8 and the maximum multipoint LOD score of 3.6 both occurred at marker rs491603 (chromosome 1; 36.3 Mb) (Table 3). Haplotype analysis revealed recombination events that define an 18.9-Mb interval between rs1886651 (chromosome 1; 29.6 Mb) and rs926979 (chromosome 1; 48.5 Mb), containing the KCNQ4 gene (Figure 1). This region corresponds closely to the 3-unit support interval, which is slightly larger and extends from rs491603 to rs1242330 (chromosome 1; 53.2 Mb).
Table 3.
LOD Scores for the DFNA2 Locus Regiona
| Marker Name |
Physical Positionsb |
Genetic Positionsc |
MPT LODd |
2-Point LOD, Recombination Fraction | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 0.00 | 0.01 | 0.05 | 0.10 | 0.20 | 0.30 | 0.40 | ||||
| rs213006 | 21,551,187 | 46.050 | 1.25 | 0.34 | 0.34 | 0.33 | 0.32 | 0.26 | 0.18 | 0.10 |
| rs6426746 | 22,501,644 | 47.280 | −∞ | −∞ | 1.20 | 0.53 | 0.27 | 0.06 | 0.02 | 0.03 |
| rs742230 | 25,124,011 | 51.220 | −∞ | −∞ | 0.90 | 0.25 | 0.01 | 0.15 | 0.16 | 0.11 |
| rs6564 | 28,085,562 | 54.000 | 0.05 | 1.60 | 1.56 | 1.39 | 1.17 | 0.72 | 0.28 | 0.01 |
| rs1886651e,f | 29,579,999 | 55.000 | −∞ | −∞ | 0.15 | 0.35 | 0.40 | 0.19 | 0.07 | 0.12 |
| rs555920 | 30,832,451 | 57.011 | 2.91 | −0.09 | 0.03 | 0.13 | 0.21 | 0.21 | 0.13 | 0.04 |
| rs12409103 | 31,747,269 | 59.610 | 3.27 | 0.35 | 0.34 | 0.28 | 0.22 | 0.12 | 0.05 | 0.01 |
| rs760594 | 33,231,390 | 60.510 | 3.34 | 0.47 | 0.46 | 0.41 | 0.35 | 0.23 | 0.11 | 0.03 |
| rs899378 | 34,287,917 | 61.420 | 3.41 | 0.45 | 0.44 | 0.39 | 0.33 | 0.21 | 0.11 | 0.03 |
| rs491603 | 36,304,903 | 64.310 | 3.57 | 1.88 | 1.85 | 1.73 | 1.56 | 1.18 | 0.76 | 0.33 |
| rs6619 | 37,803,283 | 66.140 | 3.56 | 0.45 | 0.44 | 0.39 | 0.33 | 0.21 | 0.11 | 0.03 |
| rs649593 | 37,980,150 | 66.770 | 3.55 | 0.95 | 0.94 | 0.89 | 0.81 | 0.65 | 0.45 | 0.23 |
| rs661316 | 39,403,884 | 69.700 | 3.54 | 1.47 | 1.44 | 1.30 | 1.12 | 0.75 | 0.39 | 0.10 |
| rs209598g | 40,635,418 | 71.570 | 3.52 | 1.51 | 1.48 | 1.36 | 1.22 | 0.91 | 0.60 | 0.28 |
| rs883062g | 42,382,454 | 72.840 | 3.48 | −0.07 | 0.07 | 0.05 | 0.04 | 0.02 | 0.01 | 0.00 |
| rs2782642 | 43,658,908 | 74.310 | 3.43 | −0.04 | 0.04 | 0.03 | 0.02 | 0.01 | 0.00 | 0.00 |
| rs1707302 | 46,373,504 | 75.400 | 3.40 | 0.48 | 0.46 | 0.42 | 0.36 | 0.24 | 0.12 | 0.03 |
| rs926979e | 48,458,022 | 77.800 | 1.68 | −0.12 | 0.11 | 0.06 | 0.02 | 0.01 | 0.02 | 0.01 |
| rs1502856 | 50,644,434 | 78.260 | 1.52 | 0.50 | 0.49 | 0.44 | 0.37 | 0.24 | 0.12 | 0.03 |
| rs1242330f | 53,169,430 | 79.250 | 13.11 | −∞ | 0.90 | 0.28 | 0.08 | 0.04 | 0.04 | 0.01 |
Abbreviations: LOD, logarithm of odds; MPT, multipoint.
All the data are given for theta values of 0.0 to 0.4.
Physical map positions from Build 36 of the human reference sequence.
Genetic map positions from the Rutgers combined linkage-physical map of the human genome, Build 36.
Scores obtained using Allegro1.2c.
These markers flank the haplotype.
These markers delimit the 3-unit support nterval.
The gene KCNQ4 is located between the marker loci rs209598 and rs883062.
A heterozygous point mutation in exon 6 of KCNQ4, c.859G>C, was detected by sequencing DNA in affected family members (Figure 2). The c.859G>C nucleotide substitution results in a p.Gly287Arg amino acid substitution in the phosphate-binding loop (Ploop) region of KCNQ4. This mutation was not found in 275 controls (550 control chromosomes). SIFT analysis of the Gly287Arg amino acid substitution predicted a deleterious effect on protein function, with a score of 0.00.
Figure 2.
Sequencing chromatogram of KCNQ4 exon 6 in a representative affected individual demonstrating the heterozygous c.859G>C base substitution (arrow) that results in a p.Gly287Arg amino acid substitution.
COMMENT
The KCNQ4 gene encodes a tetrameric voltage-gated potassium protein with 6 transmembrane domains; the N and C termini of the protein are on the intracellular side of the membrane, and the fourth transmembrane domain is the voltage sensor portion of the protein. The fifth and sixth transmembrane domains of the protein form the pore region of the protein, with the peptides between these domains constituting the P-loop. In the complete protein tetramer, 4 of these P-loops form the selectivity filter of the protein.16 More than 15 mutations in KCNQ4 have been previously reported in association with DFNA2 deafness, including missense, frameshift, nonsense, and splicing mutations.17 Most mutations occur in the channel pore (P-loop) region.18 Surface channel expression studies in Xenopus oocytes suggest that missense mutations in the P-loop of KCNQ4 reduce potassium currents in a dominant negative manner.3,19 A dominant negative effect is also seen with mutations in the P-loop of a related gene, KCNQ1, which affect potassium currents and cause cardiac arrhythmias in long QT syndrome.20 This evidence supports the hypothesis that the conserved sequences of the P-loop region are critical to pore structure, function, and targeting.
More than 50 genes have been identified in which mutations cause nonsyndromic hereditary hearing impairment, but clinical genetic testing is not widely available in the United States for most of these genes. For certain types of hearing loss, there is 1 predominant gene in which mutations are commonly found, such as WFS1 mutations and autosomal dominant low-frequency SNHL.21 Forms of hearing loss associated with temporal bone anomalies are also associated with a limited number of genes.22 However, for dominant high-frequency SNHL, mutations may occur in any of many possible genes. Affected individuals in family 63 have mild low-frequency and moderate high-frequency SNHL early in life, progressing to moderate low-frequency and severe-profound high-frequency SNHL later in life, consistent with the phenotype of DFNA2 deafness associated with KCNQ4 mutations.2,23 For families with dominant nonsyndromic SNHL following this pattern, clinical genetic testing for KCNQ4 is reasonable to consider (for laboratory listings, see http://www.genetests.org).
High-throughput technologies, such as resequencing microarrays, have been developed to analyze multiple genes mutated in SNHL.24 However, these “gene chips” may be expensive, and genes such as KCNQ4 may be excluded. In addition, without a priori evidence of linkage to a particular chromosome, it is difficult to classify variants of unknown significance.
For families that are large enough to provide sufficient statistical power, genetic linkage analysis remains an efficient means of identifying deafness genes. New methods using SNP markers have greatly decreased the cost and time of whole-genome genotyping analyses. For dominant high-frequency hearing loss, phenocopies, such as individuals with hearing loss due to aging or noise exposure, are common, which may reduce the power to detect linkage. Although family 63 (with 9 affected individuals) may be at the borderline level of power to detect linkage for a dominant trait, we detected linkage with a significant multipoint LOD score of 3.6. The linkage region identified in family 63 also includes another reported hearing loss gene, GJB3.25 However, because the evidence implicating GJB3 mutations as a cause of dominant nonsyndromic deafness is unclear,17 GJB3 was not examined for mutations in this family.
In conclusion, we describe a novel mutation in KCNQ4 segregating in a family with autosomal dominant high-frequency SNHL. Linkage analysis may still have a role to play in facilitating identification of the molecular etiology of deafness in patients in the future.
Acknowledgments
Funding/Support: This work was supported by grants R21 DC010074 and R01 D007380 (Dr Lesperance) and R01DC03594(Dr Leal) from the National Institute on Deafness and Other Communication Disorders and by the Biomedical Research Council of the University of Michigan (Dr Lesperance). Genotyping services were provided by the Center for Inherited Disease Research through a fully funded federal contract (N01-HG-65403) from the National Institutes of Health to The Johns Hopkins University.
Footnotes
Author Contributions: All authors had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. Study concept and design: Lee, Leal, and Lesperance. Acquisition of data: Arnett, Emery, Kim, Boerst, Leal, and Lesperance. Analysis and interpretation of data: Arnett, Emery, Lee, Leal, and Lesperance. Drafting of the manuscript: Arnett, Emery, Boerst, Lee, Leal, and Lesperance. Critical revision of the manuscript for important intellectual content: Arnett, Kim, Lee, Leal, and Lesperance. Statistical analysis: Lee and Leal. Obtained funding: Leal and Lesperance. Administrative, technical, and material support: Arnett, Emery, Boerst, Lee, Leal, and Lesperance. Study supervision: Lesperance.
Financial Disclosure: None reported.
Previous Presentation: This study was presented at the 25th Annual Meeting of the American Society of Pediatric Otolaryngology; May 1, 2010; Las Vegas, Nevada.
Additional Contributions: We thank the family for their participation.
References
- 1.Brown SD, Hardisty-Hughes RE, Mburu P. Quiet as a mouse: dissecting the molecular and genetic basis of hearing. Nat Rev Genet. 2008;9(4):277–290. doi: 10.1038/nrg2309. [DOI] [PubMed] [Google Scholar]
- 2.Coucke P, Van Camp G, Djoyodiharjo B, et al. Linkage of autosomal dominant hearing loss to the short arm of chromosome 1 in two families. N Engl J Med. 1994;331(7):425–431. doi: 10.1056/NEJM199408183310702. [DOI] [PubMed] [Google Scholar]
- 3.Kubisch C, Schroeder BC, Friedrich T, et al. KCNQ4, a novel potassium channel expressed in sensory outer hair cells, is mutated in dominant deafness. Cell. 1999;96(3):437–446. doi: 10.1016/s0092-8674(00)80556-5. [DOI] [PubMed] [Google Scholar]
- 4.Coucke PJ, Van Hauwe P, Kelley PM, et al. Mutations in the KCNQ4 gene are responsible for autosomal dominant deafness in four DFNA2 families. Hum Mol Genet. 1999;8(7):1321–1328. doi: 10.1093/hmg/8.7.1321. [DOI] [PubMed] [Google Scholar]
- 5.O’Connell JR, Weeks DE. PedCheck: a program for identification of genotype incompatibilities in linkage analysis. Am J Hum Genet. 1998;63(1):259–266. doi: 10.1086/301904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Abecasis GR, Cherny SS, Cookson WO, Cardon LR. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet. 2002;30(1):97–101. doi: 10.1038/ng786. [DOI] [PubMed] [Google Scholar]
- 7.Cottingham RW, Jr, Idury RM, Schäffer AA. Faster sequential genetic linkage computations. Am J Hum Genet. 1993;53(1):252–263. [PMC free article] [PubMed] [Google Scholar]
- 8.Gudbjartsson DF, Jonasson K, Frigge ML, Kong A. Allegro, a new computer program for multipoint linkage analysis. Nat Genet. 2000;25(1):12–13. doi: 10.1038/75514. [DOI] [PubMed] [Google Scholar]
- 9.Frazer KA, Ballinger DG, Cox DR, et al. International HapMap Consortium. A second generation human haplotype map of over 3.1 million SNPs. Nature. 2007;449(7164):851–861. doi: 10.1038/nature06258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Matise TC, Chen F, Chen W, et al. A second-generation combined linkage physical map of the human genome. Genome Res. 2007;17(12):1783–1786. doi: 10.1101/gr.7156307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Weeks DE, Sobel E, O’Connell JR, Lange K. Computer programs for multilocus haplotyping of general pedigrees. Am J Hum Genet. 1995;56(6):1506–1507. [PMC free article] [PubMed] [Google Scholar]
- 12.Sobel E, Lange K. Descent graphs in pedigree analysis: applications to haplotyping, location scores, and marker-sharing statistics. Am J Hum Genet. 1996;58(6):1323–1337. [PMC free article] [PubMed] [Google Scholar]
- 13.Rozen S, Skaletsky H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol. 2000;132:365–386. doi: 10.1385/1-59259-192-2:365. [DOI] [PubMed] [Google Scholar]
- 14.Ng PC, Henikoff S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003;31(13):3812–3814. doi: 10.1093/nar/gkg509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Huizing EH, van den Wijngaart WS, Verschuure J. A follow-up study in a family with dominant progressive inner ear deafness. Acta Otolaryngol. 1983;95(5–6):620–626. doi: 10.3109/00016488309139453. [DOI] [PubMed] [Google Scholar]
- 16.Diven WF, Burckart GJ, Alper CM, Jaffe R, Evans RW, Doyle WJ. Expression of acute otitis media after receptor blockade of platelet activating factor, thromboxane, and leukotrienes in the chinchilla. Ann Otol Rhinol Laryngol. 1998;107(3):199–206. doi: 10.1177/000348949810700303. [DOI] [PubMed] [Google Scholar]
- 17.Smith RJ, Hildebrand M. DFNA2 nonsyndromic hearing loss. [Accessed June 15, 2010];Gene Reviews at Gene Tests: Medical Genetics Information Resources (database online) Updated April 4, 2008 http://www.genetests.org. [Google Scholar]
- 18.Van Hauwe P, Coucke PJ, Ensink RJ, Huygen P, Cremers CW, Van Camp G. Mutations in the KCNQ4 K+ channel gene, responsible for autosomal dominant hearing loss, cluster in the channel pore region. Am J Med Genet. 2000;93(3):184–187. doi: 10.1002/1096-8628(20000731)93:3<184::aid-ajmg4>3.0.co;2-5. [DOI] [PubMed] [Google Scholar]
- 19.Mencía A, González-Nieto D, Modamio-Høybjør S, et al. A novel KCNQ4 pore-region mutation (p.G296S) causes deafness by impairing cell-surface channel expression. Hum Genet. 2008;123(1):41–53. doi: 10.1007/s00439-007-0447-7. [DOI] [PubMed] [Google Scholar]
- 20.Wollnik B, Schroeder BC, Kubisch C, Esperer HD, Wieacker P, Jentsch TJ. Patho-physiological mechanisms of dominant and recessive KVLQT1 K+ channel mutations found in inherited cardiac arrhythmias. Hum Mol Genet. 1997;6(11):1943–1949. doi: 10.1093/hmg/6.11.1943. [DOI] [PubMed] [Google Scholar]
- 21.Bespalova IN, Van Camp G, Bom SJ, et al. Mutations in the Wolfram syndrome 1 gene (WFS1) are a common cause of low frequency sensorineural hearing loss. Hum Mol Genet. 2001;10(22):2501–2508. doi: 10.1093/hmg/10.22.2501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hilgert N, Smith RJ, Van Camp G. Forty-six genes causing nonsyndromic hearing impairment: which ones should be analyzed in DNA diagnostics? Mutat Res. 2009;681(2–3):189–196. doi: 10.1016/j.mrrev.2008.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hildebrand MS, Tack D, McMordie SJ, et al. Audioprofile-directed screening identifies novel mutations in KCNQ4 causing hearing loss at the DFNA2 locus. Genet Med. 2008;10(11):797–804. doi: 10.1097/GIM.0b013e318187e106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kothiyal P, Cox S, Ebert J, et al. High-throughput detection of mutations responsible for childhood hearing loss using resequencing microarrays. BMC Biotechnol. 2010;10:10. doi: 10.1186/1472-6750-10-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Xia JH, Liu CY, Tang BS, et al. Mutations in the gene encoding gap junction protein β-3 associated with autosomal dominant hearing impairment. Nat Genet. 1998;20(4):370–373. doi: 10.1038/3845. [DOI] [PubMed] [Google Scholar]