Figure 2.
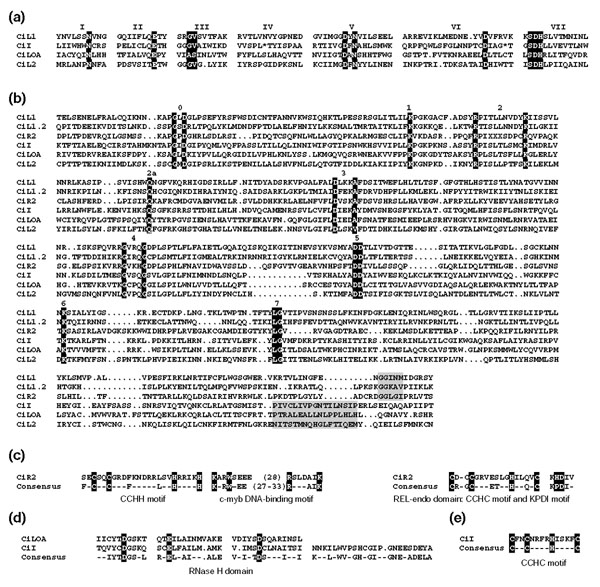
Multiple-sequence alignments of the Ciona non-LTR retrotransposons. (a) The APE region. Only blocks of highly similar residues are shown. The Roman numerals above the alignments correspond to those defined by Tu et al. [17]. Highly conserved residues, as convenient landmarks, are shaded. (b) Reverse transcriptase sequences. Numbers above the sequences and the black-shaded residues refer to the conserved peaks described by Malik et al. [11]. Gray-shaded residues correspond to the CRE/R2/R4/L1/RTE and Tad/R1/LOA/Jockey/CR1/I subgroups described in [11]. (c) CiR2 domains. The CCHH and c-Myb DNA-binding motifs are shown in the amino-terminal domain and the REL-endo domain with the CCHC and KPDI motifs in the carboxy-terminal domain [18]. (d) CiLOA and CiI RNH domains. Only the highly conserved regions of the RNase H domain of the blocks defined by Tu et al. [17] are depicted. The three amino-acid residues identified in the active site of E. coli RNase H are shaded. (e) CCHC motif of the putative CiI-ORF1 as defined by Fawcett et al. [32].
