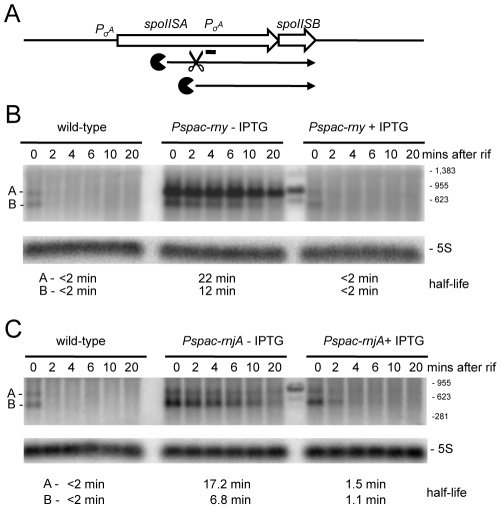
Figure 5. Degradation of the spoIISAB mRNA depends on both RNase Y and J1.
(A) Structure and predicted degradation pathway(s) of the spoIISAB transcript. ORFs are shown as large white arrows, transcripts as thin black arrows. Scissors indicate cleavage by RNase Y, ‘Pacman’ symbols represent 5′-3′ degradation by RNase J1. A short thick line indicates the position of the probe used. PσA indicates the approximate promoter position and relevant sigma factor mapped in [26]. (B) Northern blot of total mRNA isolated at times after addition of rifampicin (rif) from wild-type and RNase Y depleted (Pspac-rny−IPTG) and RNase Y induced (Pspac-rny+IPTG) cells. The blot was probed with 5′-labeled oligo CCB826 (Table S6) and reprobed with oligo HP246 against 5S rRNA. The half-lives of the different RNA species (A, B) from the spoIISAB are given under the Northern blot. Migration positions of RNA markers are shown to the right of the blot. Some cross-hybridization to the marker is visible in the lane between the (−) and (+) IPTG samples. (C) Northern blot of total mRNA isolated at times after addition of rifampicin (rif) from wild-type and RNase J1 depleted (Pspac-rnjA−IPTG) and RNase J1 induced (Pspac-rnjA+IPTG) cells. Description as in panel (B).