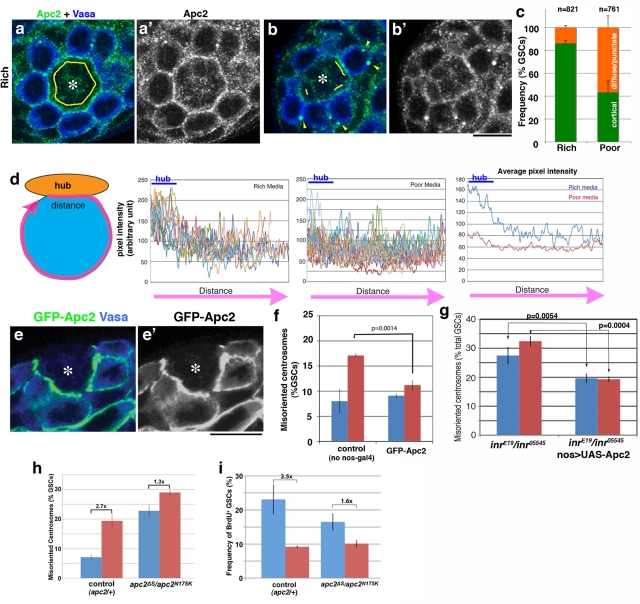
FIGURE 4:
Apc2 mediates centrosome orientation in response to nutrients. (a, b) Representative Apc2 staining in apical testes tips from flies raised in rich (a) or poor (b) media. Cortical Apc2 localization is indicated with yellow lines. Cytoplasmic punctae of Apc2 are indicated with yellow arrowheads. Green, Apc2; blue, Vasa. (c) Quantification of Apc2 localization in rich or poor media. “Cortical” indicates Apc2 protein at the hub-GSC junction; “diffuse/puncta” indicates Apc2 in the cytoplasm or occasionally at the GSC cortex outside the hub-GSC interface. n = GSCs scored. (d) Pixel intensity analysis of Apc2 protein around the GSC cortex. Circumference of GSC was traced and the pixel intensity was analyzed (see Materials and Methods for detail). Fourteen GSCs from the rich media and 19 GSCs from the poor media were analyzed. Average pixel intensity for rich vs. poor media is shown at the far right. (e) Localization of GFP-Apc2 to the hub-GSC interface following mild expression (nos-gal4>UAS-GFP-Apc2; at 18°C) in poor media. Green, GFP-Apc2; blue, Vasa. (f) The frequency of GSC centrosome misorientation upon mild expression of GFP-Apc2. n > 300/data point. (g) The frequency of GSC centrosome misorientation in the InR mutant that expresses Apc2. n > 300/data point. (h) The frequency of GSC centrosome misorientation in the apc2 mutant. n > 300/data point. (i) The S-phase index (BrdU incorporation) of apc2 mutant GSCs. n > 300/data point.