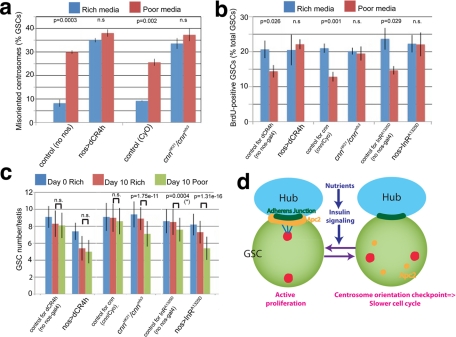
FIGURE 6:
An intact centrosome orientation checkpoint is required to slow the GSC cell cycle in poor media. (a) Centrosome misorientation frequency in GSCs mutant for cnn or expressing dominant-negative E-cadherin (dCR4h). n > 150/data point. p value (Student's t test, two-tailed) comparing centrosome misorientation in rich vs. poor media is shown. n.s., not statistically significant. (b) The S-phase index (BrdU incorporation frequency) in GSCs from checkpoint-defective mutants and those expressing InRA1325D. n > 450/data point. p value (Student's t test, two-tailed) comparing BrdU incorporation in rich vs. poor media is shown. n.s., not statistically significant. (c) Changes in GSC numbers in testes from control genotypes, cnn mutants, or those expressing dCR4h or InRA1325D after 10 d in rich vs. poor media. p value (Student's t test, two-tailed) comparing the GSC number in rich vs. poor media after 10 d is shown. n.s., not statistically significant. Asterisk indicates that a mild but statistically significant decrease in the GSC number was observed in this control group for unknown reasons. However, the degree of GSC loss was significantly (p < 0.01) more severe in InRA1325D-expressing testis compared with the control. n = 70–100 testes/data point. (d) Model of the regulation of the GSC division rate by nutrient availability and insulin signaling (see the text for details).