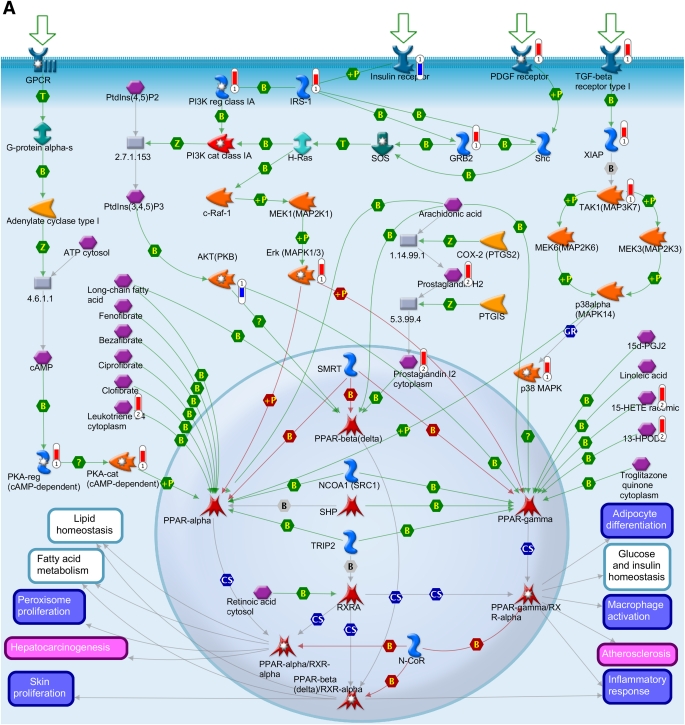
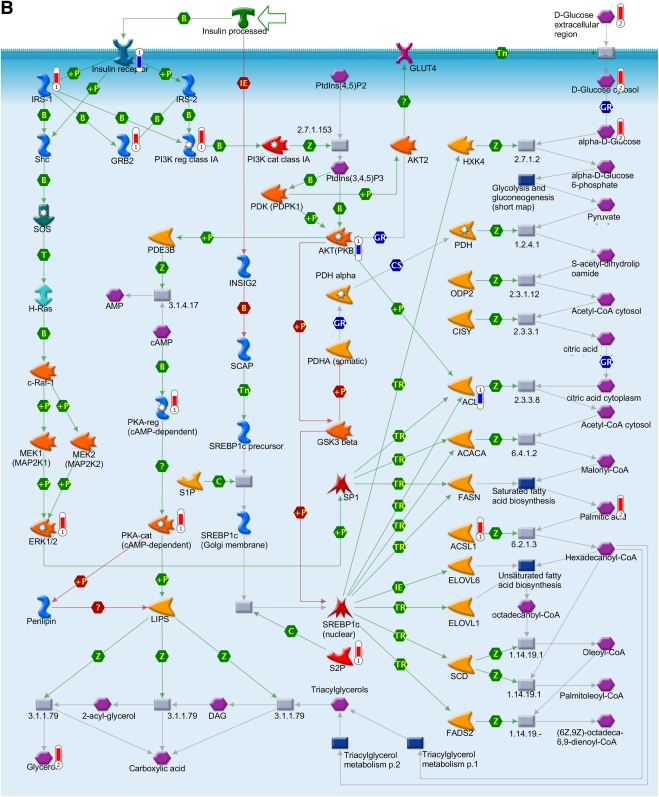
FIG. 4.
Integration of the metabolomics with transcriptomics data and their superimposition to build metabolic networks. A: Metabolic network of PPAR transcription pathway, which is connected to other metabolic processes such as lipid homeostasis, glucose, fatty acid metabolism, and inflammatory response. B: Network model of downstream of insulin-signaling pathways. The metabolites and the gene names shown in red are upregulated, and the same shown in blue are downregulated during insulin deficiency. B, binding; C, cleavage; CoA, coenzyme A; Erk, extracellular signal–related kinase; HPODE, hydroperoxylinoleic acid; IE, influence on expression; MAP, mitogen-activated protein; MAPK, MAP kinase; PDGF, platelet-derived growth factor; PI3K, phosphatidylinositol 3-kinase; PKA, cAMP-dependent protein kinase; PKB, protein kinase B; P-, dephosphorylation; RXR, retinoid X receptor; SREBP1c, sterol regulatory element–binding protein 1c; T, transformation; TGF, transforming growth factor; TR, transcription regulation; +P, phosphorylation; Z, catalysis; GPCR, G protein-coupled receptor; 15d-PGJ2, deoxy-delta prostaglandin J2; PDK/PDK1, 3-phosphoinositide-dependent protein kinase -1; ACACA, acetyl-CoA carboxylase; ACSL, acyl-CoA synthetase long-chain family members; ACLY, ATP citrate lyase; BCAA, branch chain amino acid; CISY, citrate synthase; DAG, diacylglycerol; ELOVL, elongation-of-very-long-chain-fatty acids; EMT, epithelial-mesenchymal transition; BEH, ethylene-bridged hybrid; 4E-BP1, eukaryotic translation initiation factor 4E binding protein 1; FADS1, fatty acid desaturase 1; FASN, fatty acid synthase; GSK3β, glycogen synthase kinase 3; GNAS, G protein αs- dependent adenylate cyclase; GRB2, growth factor receptor-bound protein 2; H-Ras, Harvey rat sarcoma viral oncogene homolog; HGF, hepatocyte growth factor; HXK, hexokinase; HSS, high-strength silica; HODE, hydroxyoctadecadienoic acid; INSIG2, insulin-induced gene 2; IRS-1 and IRS-2, insulin receptor substrates-1 and -2; TRIP, mediator complex subunit 1; MEK/MAP1, mitogen-activated protein kinase kinase 1; NCOA1, nuclear receptor coactivator 1; NRC1/SRC1, nuclear receptor coactivator 1; N-CoR, nuclear receptor corepressor; SMRT, nuclear receptor corepressors; NUDT1, nudix (nucleoside diphosphate-linked moiety X)-type motif 1; PtdIns(3,4,5)P3, phosphatidylinositol 3,4,5-triphosphate; P13K, phospatidylinositol 3-kinase; PtdIns(4,5)P2, phosphatidylinositol 4,5-biphosphate; PGE, prostaglandin; PTGIS, prostaglandin I2 (prostacyclin) synthase; PDGHS, prostaglandin-endoperoxide synthase 2 prostaglandin G/H synthase; COX2, cyclooxygenase 2; PDHA, pyruvate dehydrogenase (lipoamide) α1; QCs, quality controls; RARs, retinoic acid receptors; RXRA, retinoid X nuclear receptor (α; SHC, Src homology 2 domain containing transforming protein 1; SHP, small heterodimer partner; SOS, son of sevenless protein homologs 1 and 2; c-Raf-1, gene homolog 1; XIAP, X-linked inhibitor of apoptosis.