Figure 7.
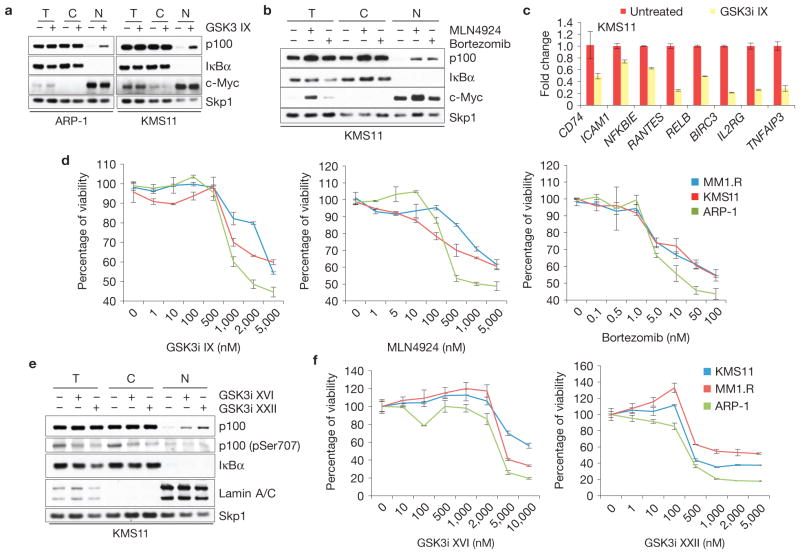
Pharmacologic inhibition of GSK3, Cullin–RING ligases or the proteasome impairs the growth of multiple myeloma cells. (a) GSK3 inhibition induces nuclear accumulation of p100 in HMMCLs. KMS11 and ARP-1 cells were treated with GSK3i IX (2 μM for 8 h). Total (T), cytoplasmic (C) and nuclear (N) fractions were isolated and analysed by immunoblotting as indicated. Cytoplasmic and nuclear markers, IκBα and c-Myc, respectively, are also shown. (b) Proteasome or Cullin–RING ligase inhibitors induce nuclear accumulation of p100 in HMMCLs. KMS11 cells were treated with either bortezomib or MLN4924 for 4 h. Total, cytoplasmic and nuclear fractions were analysed by immunoblotting as indicated. Cytoplasmic and nuclear markers, IκBα and c-Myc, respectively, are also shown. (c) NF-κB-dependent transcription is impaired in HMMCLs treated with a GSK3 inhibitor. KMS11 cells were treated with GSK3i IX and the steady-state levels of the indicated mRNAs were analysed by real-time PCR (±s.d., n = 3). The value for the amount of PCR product present in DMSO-treated cells was set as 1. (d) Inhibition of GSK3, Cullin–RING ligases or the proteasome decreases the viability of HMMCLs. KMS11, MM1.R and ARP-1 cells were treated with increasing concentrations of GSK3i IX, MLN4924 or bortezomib. Cell viability was measured by MTS assay at 72 h after treatment and individually normalized to untreated cells (0 nM), arbitrarily set as 100%. Error bars represent s.d., n = 4. (e) Next-generation GSK3 inhibitors induce accumulation of nuclear p100 in HMMCLs. KMS11 cells were treated with GSK3i XVI or GSK3i XXII (5 μM and 1 μM for 8 h, respectively). Total, cytoplasmic and nuclear fractions were isolated and analysed by immunoblotting as indicated. Cytoplasmic and nuclear markers, IκBα and Lamin A/C, respectively, are also shown. (f) Next-generation GSK3 inhibitors decrease the viability of HMMCLs. KMS11, MM1.R and ARP-1 cells were treated with increasing concentrations of GSK3i XVI or GSK3i XXII. Cell viability was measured by MTS assay at 72 h after treatment. Each value was individually normalized on the DMSO-treated cells, arbitrarily set as 100%. Error bars represent s.d., n = 4. Uncropped images of blots are shown in Supplementary Fig. S8.