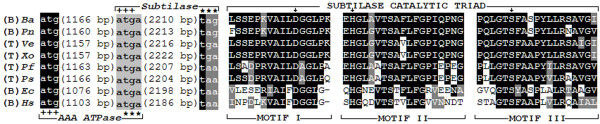
Figure 7.
Comparison of 8 members of the NCBI protein cluster CLSK923564. Ba; Burkholderia ambifaria AMMD [GenBank:YP_773860], Pn; Polaromonas naphthalenivorans CJ2 [GenBank:YP_982671], Ve; Verminephrobacter eiseniae EF01-2 [GenBank:YP_998470], Xo; Xanthomonas oryzae pv. oryzae PXO99A [GenBank:YP_001913386], Pf; P. fluorescens Pf0-1 [GenBank:YP_351424], Ps; P. syringae pv. phaseolicola 1448A [GenBank:YP_272486], Ec; E. coli O127:H6 str. E2348/69 [GenBank:YP_002329331], Hs; H. somni strain 2336 [GenBank:YP_001785203]. Gene features are shown on the left side: Start and stop codons of ORFs encoding ATPase and sutbilase are marked with plus and asterisk signs, respectively. The length of each ORF is measured from the first base after the start codon to the last base before the stop codon. The 4 bp gene overlap is box-shaded light gray. (B) denotes genes found within prophage-like sequences and (T) denotes genes associated with transposons. ClustalW-BOXSHADE multiple sequence alignment of subtilase motifs containing the catalytic triad is shown on the right side: Homologous regions are box-shaded black (identical amino acid residues) and gray (conserved amino acid substitutions). Arrows point to the conserved residues Asp-His-Ser of subtilases of the D-H-S family.