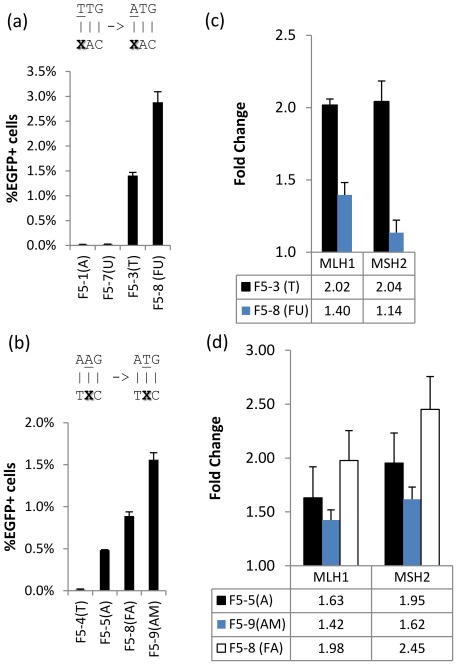
Figure 2. Chemically-modified base analogs.
(a–b)Modified base-containing oligos complementary to the non-transcribed strand’s (a)first potential start codon TTG and (b) second potential start codon AAG, where each mismatched base X in the targeting oligo is shown in parenthesis. (c–d) RNAi targeting key mismatch repair proteins MLH1 and MSH2 for the (a)TTG and (b)AAG start codon targeted by oligos containing modified bases. Data was normalized relative to scr shRNA, thus in (c) each MMR component silencing produced a 2-fold improvement for the natural T base, while the improvement for FU was reduced. This is seen to a lesser degree in (d) comparing A and AM, while FA was further improved, suggesting it is more strongly recognized by MMR. n = 4.