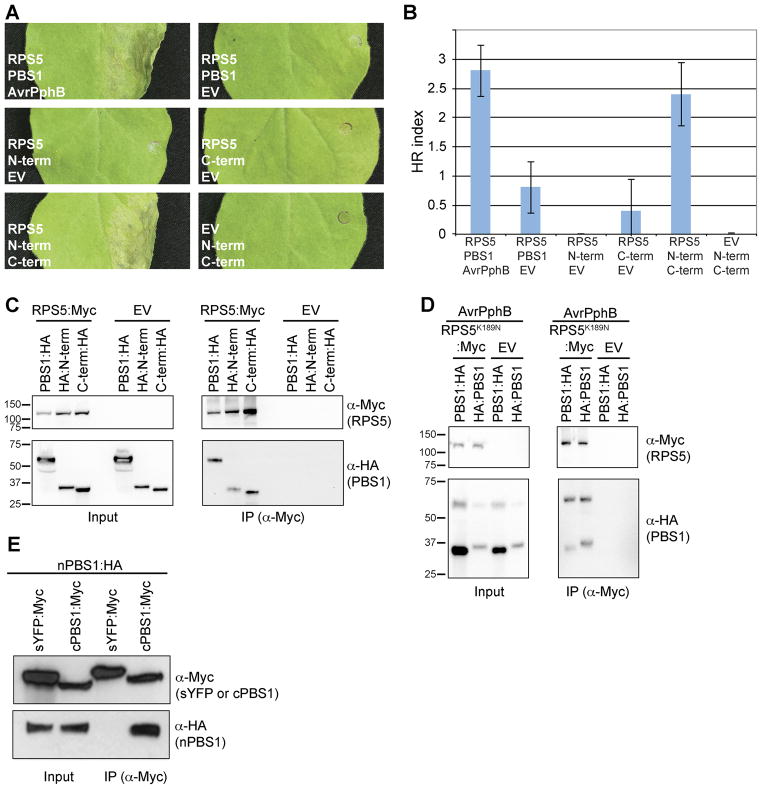
Fig. 1. Both PBS1 cleavage products are necessary and sufficient for activating RPS5 and associate with RPS5.
A. Activation of RPS5 by co-expression with both PBS1 cleavage products. The indicated proteins were transiently co-expressed in N. benthamiana. Leaves were photographed 24 h after transgene induction. N-term, PBS1 N-terminal engineered cleavage product; C-term, PBS1 C-terminal engineered cleavage product; EV, empty vector.
B. Quantification of HRs induced by co-expression of RPS5 and PBS1 cleavage products. The HR was quantified as described in Experimental Procedures, with sample sizes of 5 injections. Error bars indicate standard error.
C. Co-immunoprecipitation of PBS1 cleavage products with RPS5. The indicated PBS1 constructs were transiently co-expressed with RPS5:Myc or empty vector.
D. AvrPphB and RPS5K189N:Myc in N. benthamiana. Proteins were immunoprecipitated with anti-cMyc (IP). As a control for expression, a portion of each sample was taken prior to immunoprecipitation (input). Immunoblots were performed with the antibodies indicated on the right. The slower migrating band in panel C detected in the anti-HA blot corresponds to full-length PBS1 due to incomplete cleavage by AvrPphB.
E. Co-IP of engineered PBS1 cleavage products with each other. The indicated constructs were transiently co-expressed in N. benthamiana and immunoprecipitated with anti-cMyc. sYFP is used as a negative control and is targeted to the plasma membrane by an N-terminal fusion with the first 20 amino acids of RPS5 (Qi et al., 2012).