Figure 4.
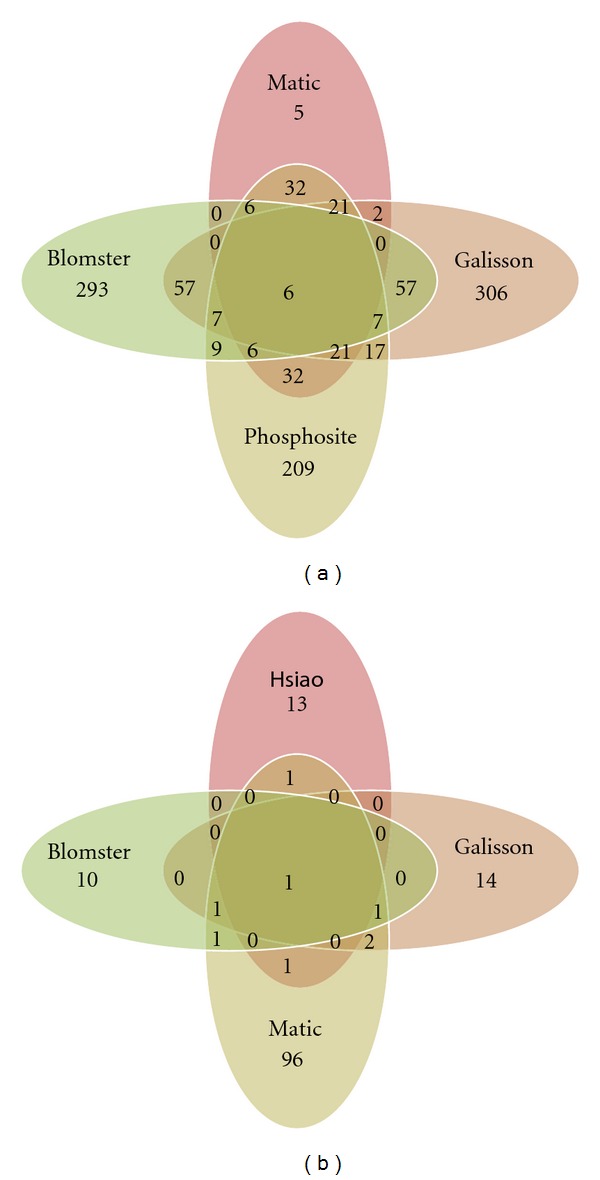
Comparative analysis of SUMO-modified proteins. (a) All proteins reported to be SUMOylated in the literature and at PhosphoSitePlus database (http://www.phosphosite.org/) were manually extracted and compared to those found by MS in 3 recent studies [39, 75, 76]. The protein list in the PhosphoSitePlus includes proteins for which the site of SUMO modification was not determined by MS. All protein names and accession numbers were first mapped to Uniprot accession numbers by using mapping data downloaded from ENSEMBL. Next, all Uniprot accession numbers were mapped to HGNC symbols and HGNC symbols for each study were uploaded to MySQL database. This means that all protein accessions that mapped to the same HGNC symbol were considered as redundant for the comparative analysis provided here. Finally, the necessary MySQL queries were made to define overlapping HGNC symbols between the different resources and the output used for creating the presented SUMO protein Venn diagram. List of proteins identified by other authors and confirmed by Matic et al.: PSMD12, TRIM24, CD3EAP, SART1, MYO1B, BRD4, SF3B1, LMNA, HNRNPC, PARP1, TOP1, KRT5, FOSL2, FLNA, MAP4, CANX, PML, STAT1, MKI67, RANGAP1, YLPM1, RBM25, RANBP2, VASP, HNRNPM, ADAR, ACTB, SUMO2, SUMO1, GTF2I, KHDRBS1, RLF, TRIM28, TCOF1, NAB1, SAFB2, NUMA1, IFI16, ZNF800, ARID4B, ZMYM1, ZMYM4, PTRF, PBRM1, CCAR1, RBM12B, FNBP4, ZBTB38, ZNF280C, KDM2B, GEMIN5, RREB1, SYMPK, ZBTB9, THOC1, ERBB2IP, RSF1, HNRNPUL1, PNN, BCLAF1, ACIN1, ZNF295, ZMYND8, TRIM33, ZBTB1, ZNF451, ACTG1, ACTB. Proteins considered in this analysis are included in the Supplementary Table 1. (b) Comparative analysis of SUMOylation sites. All peptide sequence reported with annotated SUMOylation sites based on mass spectrometry data from Matic et al. [39], Galisson et al. [76], Hsiao et al. [77], and Blomster et al. [78] were manually extracted. For each SUMO-modified site, six flanking amino acid residues on both sides were extracted. The resulting 13 amino acid residue sequences from each of the above mentioned studies were uploaded to an MySQL database and the necessary queries for comparing the peptides between studies were performed and used as input for the creation of the SUMO peptide Venn diagram.
