Table 1. Comparative performance of FEL and MEME on simulated data where  varies along phylogenetic lineages.
varies along phylogenetic lineages.
| Japanese encephalitis virus env | Vertebrate rhodopsin | Camelid VHH | ||||||||
| ω− | q + | ω+ = 4 | ω+ = 12 | ω+ = 36 | ω+ = 4 | ω+ = 12 | ω+ = 36 | ω+ = 4 | ω+ = 12 | ω+ = 36 |
| 0 | 0.1 | 0.00 0.06 | 0.01 0.25 | 0.03 0.50 | 0.00 0.21 | 0.00 0.53 | 0.02 0.81 | 0.00 0.53 | 0.00 0.95 | 0.04 0.99 |
| 0 | 0.25 | 0.01 0.12 | 0.06 0.32 | 0.12 0.51 | 0.01 0.30 | 0.04 0.68 | 0.15 0.88 | 0.00 0.66 | 0.14 0.98 | 0.56 1.00 |
| 0 | 0.5 | 0.06 0.12 | 0.19 0.29 | 0.34 0.45 | 0.09 0.28 | 0.34 0.59 | 0.54 0.82 | 0.23 0.77 | 0.85 0.98 | 0.96 0.98 |
| 0.2 | 0.1 | 0.00 0.05 | 0.01 0.21 | 0.02 0.41 | 0.00 0.09 | 0.01 0.35 | 0.02 0.67 | 0.00 0.16 | 0.01 0.87 | 0.04 0.98 |
| 0.2 | 0.25 | 0.02 0.08 | 0.07 0.27 | 0.14 0.48 | 0.03 0.17 | 0.09 0.55 | 0.17 0.84 | 0.01 0.42 | 0.27 0.96 | 0.62 0.99 |
| 0.2 | 0.5 | 0.05 0.11 | 0.18 0.29 | 0.36 0.49 | 0.13 0.25 | 0.36 0.60 | 0.55 0.76 | 0.30 0.72 | 0.84 0.99 | 0.90 0.99 |
| 0.4 | 0.1 | 0.00 0.04 | 0.01 0.15 | 0.03 0.37 | 0.01 0.07 | 0.02 0.30 | 0.03 0.57 | 0.01 0.10 | 0.04 0.78 | 0.10 0.97 |
| 0.4 | 0.25 | 0.02 0.06 | 0.09 0.27 | 0.15 0.45 | 0.04 0.16 | 0.09 0.49 | 0.21 0.78 | 0.03 0.32 | 0.33 0.97 | 0.63 0.99 |
| 0.4 | 0.5 | 0.07 0.10 | 0.17 0.26 | 0.33 0.46 | 0.17 0.28 | 0.39 0.58 | 0.51 0.76 | 0.40 0.62 | 0.82 0.94 | 0.96 1.00 |
Power to detect sites under selection ( ) are reported for FEL and MEME (in boldface) for each unique combination of negative selection (
) are reported for FEL and MEME (in boldface) for each unique combination of negative selection (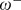 ), positive selection (
), positive selection ( ), and proportion of branches under positive selection (
), and proportion of branches under positive selection ( ) parameters.
) parameters.
